Abstract
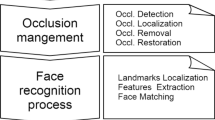
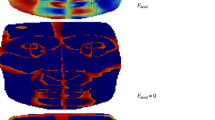
This study proposes a novel automatic method for facial landmark localization relying on geometrical properties of 3D facial surface working both on complete faces displaying different emotions and in presence of occlusions. In particular, 12 descriptors coming from Differential Geometry including the coefficients of the fundamental forms, Gaussian, mean, principal curvatures, shape index and curvedness are extracted as facial features and their local geometric properties are exploited to localize 13 soft-tissue landmarks from eye and nose areas. The method is deterministic and is backboned by a thresholding technique designed by studying the behaviour of each geometrical descriptor in correspondence to the locus of each landmark. Occlusions are managed by a detection algorithm based on geometrical properties which allows to proceed with the landmark localization avoiding the covered areas. Experimentations were carried out on 3132 faces of the Bosphorus database and of a 230-sized internal database, including expressive and occluded ones (mouth, eye, and eyeglasses occlusions), obtaining 4.75 mm mean localization error.























Similar content being viewed by others
References
Abate AF, Nappi M, Riccio D, Sabatino G (2007) 2D and 3D face recognition: a survey. Pattern Recogn Lett 28(14):1885–1906
Bagchi P, Bhattacharjee D, Nasipuri M, Basu DK (2012) A novel approach to nose-tip and eye corners detection using HK curvature analysis in case of 3D images. Third International Conference on Emerging Applications of Information Technology (EAIT), pp. 311–315
Boukamcha H, Elhallek M, Atri M, Smach F (2015) 3D face landmark auto detection. World Symposium on Computer Networks and Information Security (WSCNIS), pp. 1–6
Canavan S, Yin L (2015) Feature detection and tracking on geometric mesh data using a combined global and local shape model for face analysis. In Biometrics Theory, Applications and Systems (BTAS), 2015 I.E. 7th International Conference on. IEEE, pp. 1–6
Canavan S, Liu P, Zhang X, Yin L (2015) Landmark localization on 3D/4D range data using a shape index-based statistical shape model with global and local constraints. Comput Vis Image Underst 139:136–148
Creusot C, Pears N, Austin J (2012) 3D landmark model discovery from a registered set of organic shapes. In Computer Vision and Pattern Recognition Workshops (CVPRW), 2012 I.E. Computer Society Conference on. IEEE, pp. 57–64
Creusot C, Pears N, Austin J (2013) A machine-learning approach to keypoint detection and landmarking on 3D meshes. Int J Comput Vis 102(1–3):146–179
De Giorgis N, Rocca L, Puppo E (2015) Scale-space techniques for fiducial points extraction from 3D faces. Image Analysis and Processing—ICIAP, pp. 421–431
Dibeklioglu H, Salah AA, Gevers T (2010) Robust landmark localization and tracking for improved facial expression analysis
Ding H, Huang H, Wang D, Zhao Y, Li X, Morvan JM, Chen L (2015) An efficient multimodal 2D+ 3D feature-based approach to automatic facial expression recognition. Comput Vis Image Underst 140:83–92
Dorai C, Jain AK (1997) COSMOS-A representation scheme for 3D free-form objects. IEEE Trans Pattern Anal Mach Intell 19(10):1115–1130
Kakadiaris IA (2012) A third dimension in face recognition. 20 June 2012, SPIE Newsroom. doi:10.1117/2.1201205.004214. Link: http://spie.org/newsroom/4214-a-third-dimension-in-face-recognition
Koenderink JJ, van Doorn AJ (1992) Surface shape and curvature scales. Image Vis Comput 10(8):557–564
Lei J, You X, Abdel-Mottaleb M (2016) Automatic ear landmark localization, segmentation, and pose classification in range images. IEEE Transactions on Systems, Man, and Cybernetics: Systems 46(2):165–176
Leslie G (1981) Farkas, Anthropometry of the Head and Face in Medicine. Elsevier North Holland, Inc, New York
Leslie G (1994) Farkas, Anthropometry of the head and face (ed 2). Raven Press, New York
Li Y, Wang Y, Wang B, Sui L (2014) Nose tip detection on three-dimensional faces using pose-invariant differential surface features. Computer Vision IET 9(1):75–84
Perakis P, Passalis G, Theoharis T, Kakadiaris IA (2013) 3D facial landmark detection under large yaw and expression variations. IEEE Trans Pattern Anal Mach Intell 35(7):1552–1564
Perakis P, Theoharis T, Kakadiaris IA (2014) Feature fusion for facial landmark detection. Pattern Recogn 47(9):2783–2793
Ren S, Cao X, Wei Y, Sun J (2014) Face alignment at 3000 fps via regressing local binary features. Proc IEEE Conf Comput Vis Pattern Recognit 1685–1692
Sukno FM, Waddington JL, Whelan PF (2015) 3-d facial landmark localization with asymmetry patterns and shape regression from incomplete local features. IEEE Transactions on Cybernetics 45(9):1717–1730
Swennen GR, Schutyser FA, Hausamen JE (2006) Three-dimensional cephalometry: a color atlas and manual. Springer-Verlag Berlin Heidelberg
Szeptycki P, Ardabilian M, Chen L (2012) Nose tip localization on 2.5 D facial models using differential geometry based point signatures and SVM classifier. BIOSIG-Proceedings of the International Conference of the Biometrics Special Interest Group (BIOSIG), pp. 1–12
Vezzetti E, Marcolin F (2014) Geometry-based 3D face morphology analysis: soft-tissue landmark formalization. Multimed Tools Appl 68(3):895–929
Vezzetti E, Marcolin F (2012) Geometrical descriptors for human face morphological analysis and recognition. Robot Auton Syst 60(6):928–939
Vezzetti E, Marcolin F (2014) 3D Landmarking in multiexpression face analysis: a preliminary study on eyebrows and mouth. Aesthet Plast Surg 38:796–811
Vezzetti E, Marcolin F (2016) Novel descriptors for geometrical 3D face analysis. Multimedia Tools and Applications 10:1–30
Vezzetti E, Moos S, Marcolin F, Stola V (2012) A pose-independent method for 3D face landmark formalization. Comput Methods Prog Biomed 198(3):1078–1096
Vezzetti E, Speranza D, Marcolin F, Fracastoro G (2014) Exploiting 3D ultrasound for fetal diagnosis purpose through facial landmarking. Image Analysis & Stereology 33(3):167–188
Vezzetti E, Speranza D, Marcolin F, Fracastoro G (2015) Diagnosing cleft lip pathology in 3D ultrasound: a landmarking-based approach. Image Analysis & Stereology 35(1):53–65
Vezzetti E, Marcolin F, Fracastoro G (2014) 3D face recognition: an automatic strategy based on geometrical descriptors and landmarks. Robot Auton Syst 62(12):1768–1776
Vezzetti E, Marcolin F, Tornincasa S, Maroso P (2016) Application of geometry to RGB images for facial landmark localisation - a preliminary approach. Int J Biom 8(3–4):216–236
Zulqarnain Gilani S, Shafait F, Mian A (2015) Shape-based automatic detection of a large number of 3D facial landmarks. Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, pp. 4639–4648
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Vezzetti, E., Marcolin, F., Tornincasa, S. et al. 3D geometry-based automatic landmark localization in presence of facial occlusions. Multimed Tools Appl 77, 14177–14205 (2018). https://doi.org/10.1007/s11042-017-5025-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11042-017-5025-y