Abstract
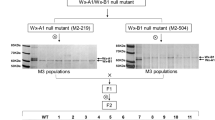
Japanese barnyard millet is an important food source in East Asian countries. However its crumbly texture limits desirability and consumption. Controlling amylose level in the endosperm is important to improve the eating quality of the millet. Because it is well known that the waxy gene determines the amylose level in the endosperm, we conducted a molecular analysis of the gene. Segregation analysis revealed that wild-type cultivars had three functional genes while low-amylose cultivars had one. We determined complete sequences of the three homoeologous waxy structural genes, EeWx1, EeWx2 and EeWx3, in a wild-type cultivar. These sequences showed high homology in the exon regions (97 %), and lower homology in the introns (82 %). Two spontaneous mutations were characterized in the low-amylose cultivars. In addition, one induced mutation was found in the fully waxy cultivar, Chojuromochi. Spontaneous mutations are deletions of whole and terminal regions in the EeWx2 and EeWx3 alleles, respectively. The induced mutation is a single-base deletion that led to a premature termination codon in EeWx1. These findings led us to develop useful markers for selecting low-amylose and waxy lines in millet.



Similar content being viewed by others
References
Bouchenak-Khelladi Y, Salamin N, Savolainen V, Forest F, van der Bank M, Chase MW, Hodkinson TR (2008) Large multi-gene phylogenetic trees of the grasses (Poaceae): progress towards complete tribal and generic level sampling. Mol Phylogenet Evol 47:488–505
Britt AB (1999) Molecular genetics of DNA repair in higher plants. Trends Plant Sci 4:20–25
Burge CB, Karlin S (1998) Finding the genes in genomic DNA. Curr Opin Struct Biol 8:346–354
Domon E, Saito A, Takeda K (2002) Comparison of the waxy locus sequence from a non-waxy strain and two waxy mutants of spontaneous and artificial origins in barley. Genes Genet Syst 77:351–359
Feldman M, Levy AA (2009) Genome evolution in allopolyploid wheat—a revolutionary reprogramming followed by gradual changes. J Genet Genomics 36:511–518
Frazer KA, Pachter L, Poliakov A, Rubin EM, Dubchak I (2004) VISTA: computational tools for comparative genomics. Nucleic Acids Res 32(Web Server issue):W273–W279
Garner TW (2002) Genome size and microsatellites: the effect of nuclear size on amplification potential. Genome 45:212–215
Gorbunova V, Levy AA (1999) How plants make ends meet: DNA double-strand break repair. Trends Plant Sci 4:263–269
Hasegawa S, Katsuta M (2005) Characteristics of barnyard millet landrace with lower amylose content in Iwate prefecture. Bull Iwate Agric Res Cent 5:53–62 (in Japanese)
Hashimoto H, Nishioka M, Fujiwara S, Takagi M, Imanaka T, Inoue T, Kai Y (2001) Crystal structure of DNA polymerase from hyperthermophilic archaeon Pyrococcus kodakaraensis KOD.1. J Mol Biol 306:469–477
Hoshino T, Nakamura T, Seimiya Y, Kamada T, Ishikawa G, Ogasawara A, Sagawa S, Saito M, Shimizu H, Nishi M, Watanabe M, Takeda J, Takahata Y (2010) Production of a fully waxy line and analysis of waxy genes in the allohexaploid crop, Japanese barnyard millet. Plant Breed 129:349–355
Ishikawa G, Takada Y, Nakamura T (2006) A PCR-based method to test for the presence or absence of β-conglycinin α′- and α-subunits in soybean. Mol Breed 17:365–374
Kamada T, Kiuchi R, Ogasawara A, Sagawa S, Shimizu H, Hoshino T (2009) Variation of amylose content and crude protein content among local varieties in Japanese barnyard millet. Breed Res 11:23–27 (in Japanese)
Kao F-T, Caldecott RS (1966) Genetic effects of recurrent irradiation in diploid and polyploidy Triticum species. Genetics 54:845–858
Kim JY, Jang KC, Park B-R, Han S-I, Choi K-J, Kim S-Y, Oh S-H, Ra J-E, Ha TJ, Lee JH, Hwang J, Kang HW, Seo WD (2011) Physicochemical and antioxidative properties of selected barnyard millet (Echinochloa utilis) species in Korea. Food Sci Biotechnol 20:461–469
Li W, Wu J, Weng S, Zhang Y, Zhang D, Shi C (2010) Identification and characterization of dwarf 62, a loss-of-function mutation in DLT/OsGRAS-32 affecting gibberellin metabolism in rice. Planta 232:1383–1396
Liu YG, Whttier RF (1995) Thermal asymmetric interlaced PCR: automatable amplification and sequncing of insert and fragments from P1 and YAC clones for chromosome walking. Genomics 25:74–681
Mayor C, Brudno M, Schwartz JR, Poliakov A, Rubin EM, Frazer KA, Pachter LS, Dubchak I (2000) VISTA: visualizing global DNA sequence alignments of arbitrary length. Bioinformatics 16:1046
Mondal S, Badigannavar AM, D’Souza SF (2011) Induced variability for fatty acid profile and molecular characterization of high oleate mutant in cultivated groundnut (Arachis hypogaea L.). Plant Breed 130:242–247
Morita R, Kusaba M, Iida S, Yamaguchi H, Nishio T, Nishimura M (2009) Molecular characterization of mutations induced by gamma irradiation in rice. Genes Genet Syst 84:361–370
Murai J, Taira T, Ohta D (1999) Isolation and characterization of the three Waxy genes encoding the granule-bound starch synthase in hexaploid wheat. Gene 234:71–79
Naito K, Kusaba M, Shikazono N, Takano T, Tanaka A, Tanisaka T, Nishimura M (2005) Transmissible and nontransmissible mutations induced by irradiating Arabidopsis thaliana pollen with gamma-rays and carbon ions. Genetics 169:881–889
Nakamura T, Yamamori M, Hirano H, Hidaka S (1993) Identification of three Wx protein in wheat (Triticum aestivum L.). Biochem Genet 31:75–86
Nakamura T, Vrinten P, Saito M, Konda M (2002) Rapid classification of partial waxy wheats using PCR-based markers. Genome 45:1150–1156
Nelson OE, Renes HW (1962) The enzymatic deficiency in the waxy mutant of maize. Biochem Biophys Res Commun 9:297–300
Nishizawa N (2003) Health benefits of millets and application to the foods. Tohoku J Crop Sci 46:95–98 (in Japanese)
Nishizawa N, Fudamoto Y (1995) The elevation of plasma concentration of high-density lipoprotein cholesterol in mice fed with protein from proso millet. Biosci Biotechnol Biochem 59:333–335
Nishizawa N, Oikawa M, Hareyama S (1990) Effect of dietary protein from proso millet on the plasma cholesterol metabolism in rats. Agric Biol Chem 54:229–230
Sakamoto S (1996) Glutinous-endosperm starch food culture specific to Eastern and Southeastern Asia. In: Ellen R, Fukui K (eds) Redefining nature: ecology, culture and domestication. Berg Publishers, Oxford, pp 215–231
Sato Y, Nishio T (2003) Mutation detection in rice waxy mutants by PCR-RF-SSCP. Theor Appl Genet 107:560–567
Tabone T, Mather DE, Hayden MJ (2009) Temperature Switch PCR (TSP): robust assay design for reliable amplification and genoty** of SNPs. BMC Genomics 10:580
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Tsuchiya T, Kameya N, Nakamura I (2009) Strait walk: a modified method of ligation-mediated genome walking for plant species with large genomes. Anal Biochem 388:158–160
Vrinten P, Nakamura T, Yamamori M (1999) Molecular characterization of waxy mutations in wheat. Mol Gen Genet 261:463–471
Watanabe M (1999) Antioxidative phenolic compounds from Japanese barnyard millet (Echinochloa utilis) grains. J Agric Food Chem 47:4500–4505
Weber SA, Fuller DQ (2006) Millets and their role in early agriculture first farmers in global perspective. Uttar Pradesh State Department of Archaeology, Lucknow
Yabuno T (1966) Biosystematic study of the genus Echinochloa. Jpn J Bot 19:277–323
Yabuno T (1987) Japanese barnyard millet (Echinochloa utilis, Poaceae) in Japan. Econ Bot 41:484–493
Yamaguchi H, Utano A, Yasuda K, Yano A, Soejima A (2005) A molecular phylogeny of wild and cultivated Echinochloa in East Asia inferred from non-coding region sequences of trnT-L-F. Weed Biol Manag 5:210–218
Yamamori M, Nakamura T, Kuroda A (1992) Variations in the content of starch granule-bound protein among several Japanese cultivars of common wheat (Triticum aestivum L.). Euphytica 64:215–219
Yasui T, Sasaki T, Matsuki K, Yamamori M (1997) Waxy endosperm mutants of bread wheat (Triticum aestivum L.) and their starch properties. Breed Sci 47:161–163
Acknowledgments
The authors thank Prof. Dr. Mark E. Sorrells, Dr. Long-** Yu and Dr. Pawan Kulwal for their useful suggestions on improving the manuscript. Thanks are also due to Ms. Haruka Yoshida and members at the Field Science Center of the Iwate University for their technical help. This work was partly supported by Germplasm Enhancement Program of the National Institute of Agrobiological Science.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ishikawa, G., Seimiya, Y., Saito, M. et al. Molecular characterization of spontaneous and induced mutations in the three homoeologous waxy genes of Japanese barnyard millet [Echinochloa esculenta (A. Braun) H. Scholz]. Mol Breeding 31, 69–78 (2013). https://doi.org/10.1007/s11032-012-9769-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11032-012-9769-9