Abstract
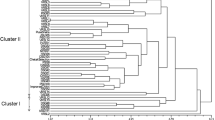
Rough lemon (Citrus × jambhiri Lush.) is one of the important species largely used as a rootstock for commercial plantations of Citrus across the world. In the present study, thirty-eight accessions of C. jambhiri were characterized using morphological and SSR markers for diversity analysis and population structure studies. Morphological characterization of 27 qualitative and 14 quantitative characters indicated the existence of moderate to sufficiently high amount of variability as revealed from the pair-wise similarity analysis value of 0.36. Molecular diversity analysis using 17 SSR primers detected 85.29% polymorphism indicating existence of moderately high amount of variability between the accessions in terms of studied loci. A total of 60 bands were generated, of which all the 60 were polymorphic (100%). The total number of alleles produced varied from 1 to 5 alleles with an average of 3.52 alleles per locus. Although the correlation between the morphological and molecular data was low in the analysed accessions of C. jambhiri, both methods allowed the clustering of accessions based on the analysed traits. Population genetic analysis by SSR markers revealed that accessions collected from North Eastern India were most diverse in terms of genetic diversity parameters and genetic distance analysis of populations showed that the accessions from North East and Himachal Pradesh were most similar genetically while population collected from Himachal Pradesh and Karnataka were the most distinct genetically. Structure analysis of different populations revealed that there is no genetic differentiation happening between the populations.







Similar content being viewed by others
References
Akhter S, Ferdous MJ, Hossain MR, Rabbani G (2009) Molecular characterization of Jamir (Citrus jambhiri) accessions of Bangladesh through PCR based RAPD markers. J Agrfor Environ 3(1):21–24
Barbhuiya AR, Khan ML, Dayanandan S (2016) Genetic structure and diversity of natural and domesticated populations of Citrus medica L. in the Eastern Himalayan region of Northeast India. Ecol Evol 6(12):3898–3911
Barkley NA, Roose ML, Krueger RR, Federici CT (2006) Assessing genetic diversity and population structure in a Citrus germplasm collection utilizing simple sequence repeat markers (SSRs). Theor Appl Genet 112:1519–1531
Chen H, Hong C, Liangliang H, Lixia W, Suhua W, Ming LW, Cheng X (2017) Genetic diversity and a population structure analysis of accessions in the Chinese cowpea [Vigna unguiculata (L.) Walp.] germplasm collection. Crop J 5(5):363–372
Doyle JJ, Doyle JL (1990) A rapid total DNA preparation procedure for fresh plant tissue. Focus 12:13–15
Gaikwad KA, Patil SR, Nagre PK, Potdukhe NR (2018) Morphological characterization of citrus rootstock genotypes. Int J Chem Sci 6(2):516–529
Jena SS, Kumar S, Nair NK (2009) Molecular phylogeny in Indian Citrus L. (Rutaceae) inferred through PCR-RFLP and trnL-trnF sequence data of chloroplast DNA. Sci Hortic 119:403–416
Lamine M, Mliki A (2015) Elucidating genetic diversity among sour orange rootstocks: a comparative study of the efficiency of RAPD and SSR markers. Appl Biochem Biotechnol 175(6):2996–3013
Liu K, Muse SV (2005) PowerMarker: an integrated analysis environment for genetic marker analysis. Bioinformatics 21:2128–2129
Maya MA, Rabbani MG, Mahboob MG, Matsubara Y (2012) Assessment of genetic relationship among 15 citrus fruits using RAPD. Asian J Biotechnol 4(1):30–37
Mouei RE, Choumane W, Dway F (2011) Characterization and estimation of genetic diversity in citrus rootstocks. Int J Agric Biol 13(4):571–575
Nei M (1972) Genetic distance between populations. Am Nat 106(949):283–292
Nei M (1978) Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics 89:583–590
NHB (2017–18) Fruit production database. National Horticulture Board. New Delhi, India. http://www.nhb.in
Paudyal KP, Haq N (2008) Variation of pomelo (Citrus grandis (L.) Osbeck) in Nepal and participatory selection of strains for further improvement. Agrofor Syst 72:195–204
Peakall R, Smouse PE (2012) GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research—an update. Bioinformatics 28(19):2537–2539
Perrier X, Jacquemoud-Collet J (2006) DARwin software; [cited November 15, 2010]. http://darwin.cirad.fr/
Rohlf FJ (2000) NTSYS-pc: numerical taxonomy and multivariate analysis system, ver.2.10e. Exter Ltd, Setauket
Scora RW (1975) On the history and Origin of Citrus. Bull Torrey Bot Club 102:369–375
Singh IP, Singh S (2006) Citrus monograph. National Research Centre for Citrus, Nagpur, pp 1–96
Singh J, Dhaliwal HS, Thakur A, Chhuneja P, Sidhu G, Singh R (2017) Morphological and genetic diversity in citrus genotypes to substantiate rootstock breeding for root rot resistance. Ind J Hortic 74(3):326
Smith JSC, Chin ECL, Shu H, Smith OS, Wall SJ, Senior ML, Mitchel SE, Kresorich S, Tiegle J (1997) An evaluation of the utility of SSR loci as molecular markers in maize (Zea mays L.): comparisons with data from RFLPs and pedigree. Theor Appl Genet 95:163–173
Tanaka T (1954) Species problem in Citrus (RevisioAurantiacearum IX). Jpn Soc Promot Sci, Tokyo
Uchoi A, Malik SK, Chaudhary R, Kumar S, Rohini MR, Pal D, Ercisli S, Chaudhury R (2016) Inferring phylogenetic relationships of Indian citron (Citrus medica L.) based on rbcL and matK sequences of chloroplast DNA. Biochem Genet 54(3):249–269
Webber HJ, Reuther W, Lawton HW (1967) History and development of the citrus industry. In: Reuther W, Webber HJ, Batchelor LD (eds) The citrus industry, history, world distribution, botany, and varieties, vol I. University of California, Division of Agricultural Sciences, Berkeley, pp 1–39
Yao D, Myriam H, Mohamed A, Claude L, Xavier V (2007) Evaluation de la diversité morphologique des variétés traditionnelles de sorgho du Nord- ouest du Maroc. Biotechnol Agron Soc Environ 11(1):39–46
Zerihun DU, Vashist U, Boora KS (2009) Molecular characterization of citrus cultivars using DNA markers. Int J Biotechnol Biochem 5(3):271–280
Zhu YF, Qin GC, Hu J, Wang J, Wang JC, Zhu S (2012) Fingerprinting and variety identification of rice (Oryza sativa L.) based on simple sequence repeat markers. Plant Omics J 5:421–426
Acknowledgements
Authors are thankful to Indian Council of Agricultural Research, New Delhi, Director, ICAR-National Bureau of Plant Genetic Resources, New Delhi, Director, ICAR-Indian Institute of Horticultural Research, Banglore for providing the fund and facilities for this research work.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Rohini, M.R., Sankaran, M., Rajkumar, S. et al. Morphological characterization and analysis of genetic diversity and population structure in Citrus × jambhiri Lush. using SSR markers. Genet Resour Crop Evol 67, 1259–1275 (2020). https://doi.org/10.1007/s10722-020-00909-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-020-00909-4