Abstract
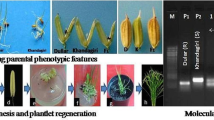
In order to analyse the genetic control of anther culture response in barley, a doubled-haploid (DH) population from the cross between a medium responsive cultivar ‘Dobla’ and the model cultivar ‘Igri’ was produced. A linkage map was constructed with 91 markers. A sub-population of 41 lines was characterised for different components of the anther culture response, and was used for quantitative trait loci (QTL) analysis. The vrs1 locus region on chromosome 2H, which determines inflorescence row type, was coincident with the largest putative QTL for number of embryos (nEMB) and albino plants. A region of chromosome 6H was associated with QTLs for nEMB and green plants. QTLs for number and percentage of green plants were located on the long arm of chromosome 5H. Therefore, new QTLs for main components of barley anther culture response were identified on chromosomes 2H, 5H and 6H, indicating that anther culture response in barley could be controlled by relative few genes of large effect. This work is a useful step towards the identification of new regions on the barley genome that could be associated with fundamental biological process implicated in the anther culture response.

Similar content being viewed by others
References
Beaumont VH, Rocheford TR, Widholm JM (1995) Map** the anther culture response genes in maize (Zea mays L.). Genome 38:968–975
Becker J, Heun M (1995) Barley microsatellites: allele variation and map**. Plant Mol Biol 27:835–845
Blake TK, Kadyrzhanova CL, Shepherd KW, Islam AK, Langridge PL, McDonald CL, Erpelding J, Larson S, Blake NK, Talbert LE (1996) STS-PCR markers appropriate for wheat -barley introgression. Theor Appl Genet 93:826–832
Bregitzer P, Campbell RD (2001) Genetic markers associated with green and albino plant regeneration from embryogenic barley callus. Crop Sci 41:173–179
Castillo AM, Cistué L, Romagosa I, Vallés MP (2001a) Low responsiveness of six-rowed genotypes to androgenesis in barley does not have a pleiotropic basis. Genome 44:936–940
Castillo AM, Cistué L, Vallés MP, Sanz JM, Romagosa I, Molina-Cano JL (2001b) Efficient production of androgenic doubled haploid mutants in barley by the application of sodium azide to anther and microspore cultures. Plant Cell Rep 20:105–111
Castillo AM, Vallés MP, Cistué L (2001c) Improvements of barley androgenesis for plant breeding. In: Bohanec B (ed.) Biotechnological approaches for utilization of gametic cells. Office for Official Publications of the European Communities, Luxembourg, pp 15–21
Cistué L, Ramos A, Castillo AM (1999) Influence of anther pretreatment and culture medium composition on the production of barley doubled haploids from model and low responding cultivars. Plant Cell Tiss Org Cult 55:159–166
Cistué L, Vallés MP, Echávarri B, Sanz JM, Castillo AM (2003) Barley anther culture. In: Malupszynski M, Kasha K, Foster B (eds) Doubled haploid production in crop plants. A Manual. FAO/IAEA Division, Wien, pp 29–35
Cowen NM, Johnson CD, Armstrong K (1992) Map** genes conditioning in vitro androgenesis in maize using RFLP analysis. Theor Appl Genet 84:720–724
Devaux P, Pickering R (2005) Haploids in the improvement of Poaceae. In: Palmer CE, Keller WA, Kasha KJ (eds) Haploids in crop improvement II. Biothechnology in Agriculture and Forestry 56. Springer, Berlin Heidelberg New York, pp 215–242
Foroughi-Wehr B, Friedt W, Wenzel G (1982) On the genetic improvement of androgenic haploid formation in Hordeum vulgare L. Theor Appl Genet 62:233–239
Forster BP, Thomas WTB (2003) Doubled haploids in genetic map** and genomics. In: Malupszynski M, Kasha K, Foster B (eds) Doubled haploid production in crop plants. A Manual. FAO/IAEA Division, Wien, pp 367–390
Forster BP, Thomas WTB (2005) Doubled haploids in genetics and plant breeding. Plant Breed Rev 25:57–88
Grobe BA, Deimling S, Geiger HH (1996) Map** of genes for anther culture ability in rye by molecular markers. Vortr Pflanzenzüchtg 35:282–283
He P, Shen L, Lu C, Chen Y, Zhu L (1998) Analysis of quantitative trait loci which contribute to anther culturability in rice (Oryza sativa L). Mol Breed 4:165–172
Horvath H, Huang J, Wong OT, von Wettstein D (2002) Experiencies with genetic transformation of barley and characteristics of transgenic plants. In: Slafer GA, Molina-Cano JL, Savin R, Araus JL, Romagosa I (eds) Barley science: recent advances from molecular biology to agronomy of yield and quality. Food Products Press, New York, pp 143–174
Hou L, Ullrich SE, Kleinhofs A (1994) Inheritance of anther culture traits in Barley. Crop Sci 34:1243–1247
Jacquard C, Asakaviciute R, Hamalian AM, Sangwan RS, Devaux P, Clément C (2006) Barley anther culture: effects of annual cycle and spike position on microspore embryogenesis and albinism. Plant Cell Rep 25:375–381
Komatsuda T, Annaka T, Oka S (1993) Genetic map** of a quantitative trait locus (QTL) that enhances the shoot differentiation rate in Hordeum vulgare L. Theor Appl Genet 86:713–720
Kuenzel G, Korzum L, Meister A (2000) Cytologically integrated physical restriction fragment length polymorphism maps for the barley genome based on translocation breakpoints. Genetics 154:397–412
Kwon YS, Kim KM, Eun MY, Sohn JK (2002) QTL map** and associated marker selection for the efficacy of green plant regeneration in anther culture of rice. Plant Breed 121:10–16
Lander ES, Green P, Abrahamson J, Barlow A, Daly MJ, Lincon SE, Newburg L (1987) Mapmaker: an interactive computer pachkage for constructing primary genetic linkage maps of experimental and natural populations. Genomics 1:174, 181
Larsen ET, Tuvesson IKD, Andersen SB (1991) Nuclear genes affecting percentage of green plants in barley (Hordeum vulgare L.) anther culture. Theor Appl Genet 82:417–420
Liu ZW, Biyashev RM, Saghai-Maroof MA (1996) Development of simple sequence repeat DNA markers and their integration into a barley linkage map. Theor Appl Genet 93:869–876
Magnard JL, Le Deunff E, Doménech J, Rogowsky PM, Testillano PS, Rougier M, Risueño MC, Vergne P, Dumas C (2000) Genes normally expressed in the endosperm are expressed at early stages of microspore embryogenesis in maize. Plant Mol Biol 44:559–574
Manninen OM (2000) Associations between anther-culture response and molecular markers on chromosome 2H, 3H and 4H of barley (Hordeum vulgare L.). Theor Appl Genet 100:57–62
Mano Y, Takahashi H, Sato K, Takeda K (1996) Map** genes for callus growth and shoot regeneration in barley (Hordeum vulgare L.). Breed Sci 46:137–142
Maraschin SF, Caspers M, Potokina E, Wülfert F, Graner A, Spaink HP, Wang M (2006) cDNA array analysis of stress-induced gene expression in barley androgenesis. Physiol Plant 127:535–550
Muñoz-Amatriaín M, Svensson JT, Castillo AM, Cistué L, Close TJ, Vallés MP (2006) Transcriptome analysis of barley anthers: effect of mannitol treatment on microspore embryogenesis. Physiol Plant 127:551–560
Murigneux A, Bentolila S, Hardy T, Baud S, Guitton C, Jullien H, Ben Tahar S, Freyssinet G, Beckert M (1994) Genotypic variation of quantitative trait loci controlling in vitro androgenesis in maize. Genome 37:970–976
Powell W (1988) Diallel analysis of barley anther culture response. Genome 30:152–157
Ramsay L, Macaulay M, degli Ivanissevich S, MacLean K, Cardle L, Fuller J, Edwards KJ, Tuvesson S, Morgante M, Massari A, Maestri E, Marmiroli N, Sjakste T, Ganal M, Powell W, Waugh R (2000) A simple sequence repeat-based linkage map of barley. Genetics 156:1997–2005
Reynolds TL, Crawford RL (1996) Changes in abundance of an abscisic acid-responsive, early cysteine-labeled metallothionein transcript during pollen embryogenesis in bread wheat (Triticum aestivum). Plant Mol Biol 32:823–829
SAS Institute (1989) SAS/STAT User’s guide, Version 6.03. SAS Institute, Cary, NC, USA
Sayed-Tabatabaei BE, Komatsuda T, Takaiwa F, Graner A (1999) DNA sequencing and primer designing for RFLP clones evenly distributed in the barley genome. Barley Genet Newsl 28:15–18
Szarejko I (2003) Doubled haploid mutation production. In: Maluszynski M, Kasha KJ, Forster BP, Szarejko I (eds) Doubled haploid production in crop plants, a manual. Kluwer Academic Publishers, Dordrecht, pp 351–362
Tinker NA, Mather DE (1995) MQTL: software for simplified composite interval map** of QTL in multiple environments. J Quant Trait Loci, http:/nalusda.gov:8000/otherdocs/jqtl/2
Torp AM, Hansen AL, Anderson SB (2001) Chromosomal regions associated with green plant regeneration in wheat (Triticum aestivum L.) anther culture. Euphytica 119:377–387
Torp AM, Bekesiova I, Holme IB, Hansen AL, Andersen SB (2004) Genetics related to doubled haploid induction in vitro. In: Mujib A (ed) In vitro applications in crop improvement. Science Publishers, Playmouth, pp 34–52
Vergne P, Riccardi F, Beckert M, Dumas C (1993) Identification of a 32-kDa anther marker protein for androgenic response in maize, Zea mays L. Theor Appl Genet 86:843–850
Vrinten PL, Nakamura T, Kasha KJ (1999) Characterization of cDNAs expressed in the early stages of microspore embryogenesis in barley (Hordeum vulgare) L. Plant Mol Biol 41:455–463
Yamagishi M, Otani M, Higashi M, Fukuta Y, Fukui K, Yano M, Shimada T (1998) Chromosomal regions controlling anther culturability in rice (Oryza sativa L). Euphytica 103:227–234
Acknowledgements
X-W. Chen was recipient of a fellowship form the Ministerio de Educación y Cultura from the Spanish government. The research was supported by Projects PB97-1159, and AGL2001-1631 from Plan Comisión Interministerial de Ciencia y Tecnología of Spain.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Chen, XW., Cistué, L., Muñoz-Amatriaín, M. et al. Genetic markers for doubled haploid response in barley. Euphytica 158, 287–294 (2007). https://doi.org/10.1007/s10681-006-9310-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10681-006-9310-5