Abstract
Background
Histone methylation has been considered as one of the epigenetic mechanisms of carcinogenesis and progression. Researches on the correlation between histone lysine methylation and gastric cancer (GC) will help in finding novel epigenetic biomarkers for monitoring cancers.
Aims
The study detected the expression patterns of histone 3 lysine 9 dimethylation (H3K9me2), histone 3 lysine 9 trimethylation (H3K9me3), and histone 3 lysine 27 trimethylation (H3K27me3) in GC tissues and evaluated their clinical merit for GC patients.
Methods
One hundred thirty-three paraffin-embedded GC samples were examined by immunohistochemistry for the histone markers: H3K9me2, H3K9me3, and H3K27me3. The relationship and clinicopathological significance of the three lysine methylations on histone H3 with GC were assessed by Paired t test, Chi-square test, Kaplan–Meier analysis with log-rank test, and Cox proportional hazard analyses.
Results
Strong positive immunostaining of H3K9me2, H3K9me3, and H3K27me3 was observed in cancerous tissues than in their counterpart non-cancer tissues. Higher expression patterns of H3K9me2, H3K9me3, and H3K27me3 significantly related to differentiation degree, lymph nodes metastases, and pathological TNM staging in GC. The GC patients with low scoring of the three markers implied long survival period and best prognosis. In contrast, the patients’ survival time was significantly shorter if their cancerous tissues presented high expression of the three markers.
Conclusions
H3K9me2, H3K9me3, and H3K27me3 expression patterns closely relate to clinicopathological features and may be the independent risk factors for the survival of GC patients. The combined pattern of the three markers rather than an individual marker is considered to more accurately evaluate the outcome of GC patients.



Similar content being viewed by others
References
Aoyama T, Yoshikawa T. Adjuvant therapy for locally advanced gastric cancer. Surg Today. 2017;47:1295–1302.
Correa P. Gastric cancer: overview. Gastroenterol Clin N Am. 2013;42:211–217.
Li T, Mo X, Fu L, **ao B, Guo J. Molecular mechanisms of long noncoding RNAs on gastric cancer. Oncotarget. 2016;7:8601–8612.
Cheetham SW, Gruhl F, Mattick JS, Dinger ME. Long noncoding RNAs and the genetics of cancer. Br J Cancer. 2013;108:2419–2425.
Hirst M, Marra MA. Epigenetics and human disease. Int J Biochem Cell Biol. 2009;41:136–146.
Panani AD. Cytogenetic and molecular aspects of gastric cancer: clinical implications. Cancer Lett. 2008;266:99–115.
Jenuwein T, Allis CD. Translating the histone code. Science. 2001;293:1074–1080.
Jeltsch A, Jurkowska RZ. New concepts in DNA methylation. Trends Biochem Sci. 2014;39:310–318.
Yang WY, Gu JL, Zhen TM. Recent advances of histone modification in gastric cancer. J Cancer Res Therapeut. 2014;10 Suppl:240–245.
Mulero-Navarro S, Esteller M. Epigenetic biomarkers for human cancer: the time is now. Crit Rev Oncol/Hematol. 2008;68:1–11.
Noma K, Allis CD, Grewal SI. Transitions in distinct histone H3 methylation patterns at the heterochromatin domain boundaries. Science. 2001;293:1150–1155.
Peters AH, O’Carroll D, Scherthan H, et al. Loss of the Suv39h histone methyltransferases impairs mammalian heterochromatin and genome stability. Cell. 2001;107:323–337.
Kondo Y, Shen L, Issa JP. Critical role of histone methylation in tumor suppressor gene silencing in colorectal cancer. Mol Cell Biol. 2003;23:206–215.
Park YS, ** MY, Kim YJ, Yook JH, Kim BS, Jang SJ. The global histone modification pattern correlates with cancer recurrence and overall survival in gastric adenocarcinoma. Ann Surg Oncol. 2008;15:1968–1976.
Meng CF, Zhu XJ, Peng G, Dai DQ. Promoter histone H3 lysine 9 di-methylation is associated with DNA methylation and aberrant expression of p16 in gastric cancer cells. Oncol Rep. 2009;22:1221–1227.
Kuzmichev A, Nishioka K, Erdjument-Bromage H, Tempst P, Reinberg D. Histone methyltransferase activity associated with a human multiprotein complex containing the Enhancer of Zeste protein. Genes Dev. 2002;16:2893–2905.
Cao R, Wang L, Wang H, et al. Role of histone H3 lysine 27 methylation in Polycomb-group silencing. Science. 2002;298:1039–1043.
Cao R, Zhang Y. The functions of E(Z)/EZH2-mediated methylation of lysine 27 in histone H3. Curr opin Genet Dev. 2004;14:155–164.
Sparmann A, van Lohuizen M. Polycomb silencers control cell fate, development and cancer. Nat Rev Cancer. 2006;6:846–856.
Plath K, Fang J, Mlynarczyk-Evans SK, et al. Role of histone H3 lysine 27 methylation in X inactivation. Science. 2003;300:131–135.
Fujii S, Ochiai A. Enhancer of zeste homolog 2 downregulates E-cadherin by mediating histone H3 methylation in gastric cancer cells. Cancer Sci. 2008;99:738–746.
Matsukawa Y, Semba S, Kato H, Ito A, Yanagihara K, Yokozaki H. Expression of the enhancer of zeste homolog 2 is correlated with poor prognosis in human gastric cancer. Cancer Sci. 2006;97:484–491.
Zhang L, Zhong K, Dai Y, Zhou H. Genome-wide analysis of histone H3 lysine 27 trimethylation by ChIP-chip in gastric cancer patients. J Gastroenterol. 2009;44:305–312.
Zhang W, Li Y, Yang S, et al. Differential mitochondrial proteome analysis of human lung adenocarcinoma and normal bronchial epithelium cell lines using quantitative mass spectrometry. Thorac Cancer. 2013;4:373–379.
Varghese F, Bukhari AB, Malhotra R, De A. IHC Profiler: an open source plugin for the quantitative evaluation and automated scoring of immunohistochemistry images of human tissue samples. PloS ONE. 2014;9:e96801.
Zhang T, Cooper S, Brockdorff N. The interplay of histone modifications—writers that read. EMBO Rep. 2015;16:1467–1481.
Liu F, Wang L, Perna F, Nimer SD. Beyond transcription factors: how oncogenic signalling reshapes the epigenetic landscape. Nat Rev Cancer. 2016;16:359–372.
Alam H, Gu B, Lee MG. Histone methylation modifiers in cellular signaling pathways. Cell Mol Life Sci CMLS. 2015;72:4577–4592.
Benard A, Goossens-Beumer IJ, van Hoesel AQ, et al. Histone trimethylation at H3K4, H3K9 and H4K20 correlates with patient survival and tumor recurrence in early-stage colon cancer. BMC Cancer. 2014;14:531.
Healey MA, Hu R, Beck AH, et al. Association of H3K9me3 and H3K27me3 repressive histone marks with breast cancer subtypes in the Nurses’ Health Study. Breast Cancer Res Treat. 2014;147:639–651.
Yokoyama Y, Matsumoto A, Hieda M, et al. Loss of histone H4K20 trimethylation predicts poor prognosis in breast cancer and is associated with invasive activity. Breast Cancer Res BCR. 2014;16:R66.
Chen YW, Kao SY, Wang HJ, Yang MH. Histone modification patterns correlate with patient outcome in oral squamous cell carcinoma. Cancer. 2013;119:4259–4267.
Keung EZ, Akdemir KC, Al Sannaa GA, et al. Increased H3K9me3 drives dedifferentiated phenotype via KLF6 repression in liposarcoma. J Clin Investig. 2015;125:2965–2978.
Maia LL, Peterle GT, Dos Santos M, et al. JMJD1A, H3K9me1, H3K9me2 and ADM expression as prognostic markers in oral and oropharyngeal squamous cell carcinoma. PLoS ONE. 2018;13:e0194884.
Sasidharan Nair V, El Salhat H, Taha RZ, John A, Ali BR, Elkord E. DNA methylation and repressive H3K9 and H3K27 trimethylation in the promoter regions of PD-1, CTLA-4, TIM-3, LAG-3, TIGIT, and PD-L1 genes in human primary breast cancer. Clin Epigenet. 2018;10:78.
Li Q, Wang X, Lu Z, et al. Polycomb CBX7 directly controls trimethylation of histone H3 at lysine 9 at the p16 locus. PLoS ONE. 2010;5:e13732.
Cui, H, Hu, Y, Guo, D, Zhang, A, Gu, Y, Zhang, S, et al. DNA methyltransferase 3A isoform b contributes to repressing E-cadherin through cooperation of DNA methylation and H3K27/H3K9 methylation in EMT-related metastasis of gastric cancer. Oncogene 2018;37:4358–4371.
Cao Q, Yu J, Dhanasekaran SM, et al. Repression of E-cadherin by the polycomb group protein EZH2 in cancer. Oncogene. 2008;27:7274–7284.
Chase A, Cross NC. Aberrations of EZH2 in cancer. Cancer Res. 2011;17:2613–2618.
Chen MW, Hua KT, Kao HJ, et al. H3K9 histone methyltransferase G9a promotes lung cancer invasion and metastasis by silencing the cell adhesion molecule Ep-CAM. Cancer Res. 2010;70:7830–7840.
Acknowledgments
This work was supported by the grants from National Natural Science Foundation of China (81672414 and 81472548). This work was also supported by the Innovation Capability Development Project of Jiangsu Province (BM2015004).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
We declare that we do not have any commercial or associative interest that represents a conflict of interest in connection with the work submitted.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Fig. S1
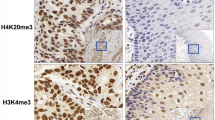
The expression patterns for the stained histone markers by IHC assay. The expression was scored according to the staining intensity and staining area. The staining intensity was marked as the following, 0 for no staining, 1 for weak staining, 2 for moderate staining, 3 for intensive staining. Based on the percentage of positive cells in the observed epithelial tissues, the staining area was identified as 5 different levels. 0 for 0–5% positive cells, 1, 2, 3, 4 for 6–25%, 26–50%, 51–75%, and 76–100%, respectively. The final score is the product of the staining intensity and staining area. Low indicates for the tissues with the final score ≤ 8, high for the final score > 8. Scale bar = 200 μm (TIFF 12090 kb)
Rights and permissions
About this article
Cite this article
Li, Y., Guo, D., Sun, R. et al. Methylation Patterns of Lys9 and Lys27 on Histone H3 Correlate with Patient Outcome in Gastric Cancer. Dig Dis Sci 64, 439–446 (2019). https://doi.org/10.1007/s10620-018-5341-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10620-018-5341-8