Abstract
Purpose
The metabolic etiology of breast cancer has been explored in the past several years using metabolomics. However, most of these studies only included non-Hispanic White individuals.
Methods
To fill this gap, we performed a two-step (discovery and validation) metabolomics profiling in plasma samples from 358 breast cancer patients and 138 healthy controls. All study subjects were either Hispanics or non-Hispanic African Americans.
Results
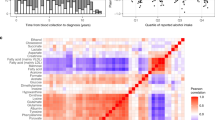
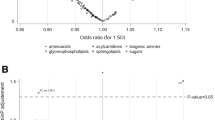
A panel of 14 identified metabolites significantly differed between breast cancer cases and healthy controls in both the discovery and validation sets. Most of these identified metabolites were lipids. In the pathway analysis, citrate cycle (TCA cycle), arginine and proline metabolism, and linoleic acid metabolism pathways were observed, and they significantly differed between breast cancer cases and healthy controls in both sets. From those 14 metabolites, we selected 9 non-correlated metabolites to generate a metabolic risk score. Increased metabolites risk score was associated with a 1.87- and 1.63-fold increased risk of breast cancer in the discovery and validation sets, respectively (Odds ratio (OR) 1.87, 95% Confidence interval (CI) 1.50, 2.32; OR 1.63, 95% CI 1.36, 1.95).
Conclusions
In summary, our study identified metabolic profiles and pathways that significantly differed between breast cancer cases and healthy controls in Hispanic or non-Hispanic African American women. The results from our study might provide new insights on the metabolic etiology of breast cancer.

Similar content being viewed by others
Change history
01 June 2019
In the original publication of the article, the sixth author name Krita A. Zanetti was mistakenly included as co-author. The corrected author group is given in the correction article. The original article has been corrected.
References
Siegel RL, Miller KD, Jemal A (2018) Cancer statistics, 2018. CA Cancer J Clin 68(1):7–30. https://doi.org/10.3322/caac.21442
Yedjou CG, Tchounwou PB, Payton M, Miele L, Fonseca DD, Lowe L, Alo RA (2017) Assessing the racial and ethnic disparities in breast cancer mortality in the United States. Int J Environ Res Public Health. https://doi.org/10.3390/ijerph14050486
Claudino WM, Goncalves PH, di Leo A, Philip PA, Sarkar FH (2012) Metabolomics in cancer: a bench-to-bedside intersection. Crit Rev Oncol Hematol 84(1):1–7. https://doi.org/10.1016/j.critrevonc.2012.02.009
Yu B, Zheng Y, Alexander D, Morrison AC, Coresh J, Boerwinkle E (2014) Genetic determinants influencing human serum metabolome among African Americans. PLoS Genet. https://doi.org/10.1371/journal.pgen.1004212
Playdon MC, Ziegler RG, Sampson JN, Stolzenberg-Solomon R, Thompson HJ, Irwin ML, Mayne ST, Hoover RN, Moore SC (2017) Nutritional metabolomics and breast cancer risk in a prospective study. Am J Clin Nutr 106(2):637–649. https://doi.org/10.3945/ajcn.116.150912
Moore SC, Playdon MC, Sampson JN, Hoover RN, Trabert B, Matthews CE, Ziegler RG (2018) A metabolomics analysis of body mass index and postmenopausal breast cancer risk. J Natl Cancer Inst 110(6):588–597. https://doi.org/10.1093/jnci/djx244
Shen J, Yan L, Liu S, Ambrosone CB, Zhao H (2013) Plasma metabolomic profiles in breast cancer patients and healthy controls: by race and tumor receptor subtypes. Transl Oncol 6(6):757–765. https://doi.org/10.1593/tlo.13619
Hart CD, Vignoli A, Tenori L, Uy GL, To TV, Adebamowo C, Hossain SM, Biganzoli L, Risi E, Love RR, Luchinat C, Di Leo A (2017) Serum metabolomic profiles identify ER-positive early breast cancer patients at increased risk of disease recurrence in a multicenter population. Clin Cancer Res 23(6):1422–1431. https://doi.org/10.1158/1078-0432.Ccr-16-1153
Tenori L, Oakman C, Morris PG, Gralka E, Turner N, Cappadona S, Fornier M, Hudis C, Norton L, Luchinat C, Di Leo A (2015) Serum metabolomic profiles evaluated after surgery may identify patients with oestrogen receptor negative early breast cancer at increased risk of disease recurrence. Results from a retrospective study. Mol Oncol 9(1):128–139. https://doi.org/10.1016/j.molonc.2014.07.012
Oakman C, Tenori L, Claudino WM, Cappadona S, Nepi S, Battaglia A, Bernini P, Zafarana E, Saccenti E, Fornier M, Morris PG, Biganzoli L, Luchinat C, Bertini I, Di Leo A (2011) Identification of a serum-detectable metabolomic fingerprint potentially correlated with the presence of micrometastatic disease in early breast cancer patients at varying risks of disease relapse by traditional prognostic methods. Ann Oncol 22(6):1295–1301. https://doi.org/10.1093/annonc/mdq606
Asiago VM, Alverado LZ, Shanaiah N, Gowda NG, Owusu-Sarfo K, Ballas R, Raftery D (2010) Early detection of recurrent breast cancer using metabolite profiling. Cancer Res. https://doi.org/10.1158/0008-5472.Sabcs10-P3-10-18
Zhang AH, Sun H, Qiu S, Wang XJ (2013) Metabolomics in noninvasive breast cancer. Clin Chim Acta 424:3–7. https://doi.org/10.1016/j.cca.2013.05.003
Hadi NI, Jamal Q, Iqbal A, Shaikh F, Somroo S, Musharraf SG (2017) Serum metabolomic profiles for breast cancer diagnosis, grading and staging by gas chromatography-mass spectrometry. Sci Rep. https://doi.org/10.1038/s41598-017-01924-9
Khodadadi-Jamayran A, Akgol-Oksuz B, Afanasyeva Y, Heguy A, Thompson M, Ray K, Giro-Perafita A, Sanchez I, Wu X, Tripathy D, Zeleniuch-Jacquotte A, Tsirigos A, Esteva FJ (2018) Prognostic role of elevated mir-24-3p in breast cancer and its association with the metastatic process. Oncotarget 9(16):12868–12878. https://doi.org/10.18632/oncotarget.24403
Evans AM, DeHaven CD, Barrett T, Mitchell M, Milgram E (2009) Integrated, nontargeted ultrahigh performance liquid chromatography/electrospray ionization tandem mass spectrometry platform for the identification and relative quantification of the small-molecule complement of biological systems. Anal Chem 81(16):6656–6667. https://doi.org/10.1021/ac901536h
DeHaven CD, Evans AM, Dai HP, Lawton KA (2010) Organization of GC/MS and LC/MS metabolomics data into chemical libraries. J Cheminform. https://doi.org/10.1186/1758-2946-2-9
Sampson JN, Boca SM, Shu XO, Stolzenberg-Solomon RZ, Matthews CE, Hsing AW, Tan YT, Ji BT, Chow WH, Cai QY, Liu DK, Yang G, **ang YB, Zheng W, Sinha R, Cross AJ, Moore SC (2013) Metabolomics in epidemiology: sources of variability in metabolite measurements and implications. Cancer Epidem Biomar 22(4):631–640. https://doi.org/10.1158/1055-9965.Epi-12-1109
Chong EY, Huang Y, Wu H, Ghasemzadeh N, Uppal K, Quyyumi AA, Jones DP, Yu T (2015) Local false discovery rate estimation using feature reliability in LC/MS metabolomics data. Sci Rep 5:17221. https://doi.org/10.1038/srep17221
Wishart DS, Jewison T, Guo AC, Wilson M, Knox C, Liu Y, Djoumbou Y, Mandal R, Aziat F, Dong E, Bouatra S, Sinelnikov I, Arndt D, **a J, Liu P, Yallou F, Bjorndahl T, Perez-Pineiro R, Eisner R, Allen F, Neveu V, Greiner R, Scalbert A (2013) HMDB 3.0—the human metabolome database in 2013. Nucleic Acids Res 41(Database issue):D801–D807. https://doi.org/10.1093/nar/gks1065
Chong J, **a JG (2018) MetaboAnalystR: an R package for flexible and reproducible analysis of metabolomics data. Bioinformatics 34(24):4313–4314. https://doi.org/10.1093/bioinformatics/bty528
Moore SC, Matthews CE, Sampson JN, Stolzenberg-Solomon RZ, Zheng W, Cai Q, Tan YT, Chow WH, Ji BT, Liu DK, **ao Q, Boca SM, Leitzmann MF, Yang G, **ang YB, Sinha R, Shu XO, Cross AJ (2014) Human metabolic correlates of body mass index. Metabolomics 10(2):259–269. https://doi.org/10.1007/s11306-013-0574-1
Zhao H, Shen J, Djukovic D, Daniel-MacDougall C, Gu H, Wu X, Chow WH (2016) Metabolomics-identified metabolites associated with body mass index and prospective weight gain among Mexican American women. Obes Sci Pract 2(3):309–317. https://doi.org/10.1002/osp4.63
Rodriguez-Enriquez S, Hernandez-Esquivel L, Marin-Hernandez A, El Hafidi M, Gallardo-Perez JC, Hernandez-Resendiz I, Rodriguez-Zavala JS, Pacheco-Velazquez SC, Moreno-Sanchez R (2015) Mitochondrial free fatty acid beta-oxidation supports oxidative phosphorylation and proliferation in cancer cells. Int J Biochem Cell B 65:209–221. https://doi.org/10.1016/j.biocel.2015.06.010
Nieman KM, Romero IL, Van Houten B, Lengyel E (2013) Adipose tissue and adipocytes support tumorigenesis and metastasis. BBA Mol Cell Biol Lipids 1831(10):1533–1541. https://doi.org/10.1016/j.bbalip.2013.02.010
Wang T, Fahrmann JF, Lee H, Li YJ, Tripathi SC, Yue C, Zhang C, Lifshitz V, Song J, Yuan Y, Somlo G, Jandial R, Ann D, Hanash S, Jove R, Yu H (2018) JAK/STAT3-regulated fatty acid beta-oxidation is critical for breast cancer stem cell self-renewal and chemoresistance. Cell Metab 27(1):136–150 e135. https://doi.org/10.1016/j.cmet.2017.11.001
Kinlaw WB, Baures PW, Lupien LE, Davis WL, Kuemmerle NB (2016) Fatty acids and breast cancer: make them on site or have them delivered. J Cell Physiol 231(10):2128–2141. https://doi.org/10.1002/jcp.25332
Cui M, Wang QL, Chen G (2016) Serum metabolomics analysis reveals changes in signaling lipids in breast cancer patients. Biomed Chromatogr 30(1):42–47. https://doi.org/10.1002/bmc.3556
O’Flaherty JT, Wooten RE, Samuel MP, Thomas MJ, Levine EA, Case LD, Akman SA, Edwards IJ (2013) Fatty acid metabolites in rapidly proliferating breast cancer. PLoS ONE. https://doi.org/10.1371/journal.pone.0063076
Zhang H, Zhou L, Shi W, Song N, Yu K, Gu Y (2012) A mechanism underlying the effects of polyunsaturated fatty acids on breast cancer. Int J Mol Med 30(3):487–494. https://doi.org/10.3892/ijmm.2012.1022
Li J, Humphreys K, Heikkinen T, Aittomaki K, Blomqvist C, Pharoah PD, Dunning AM, Ahmed S, Hooning MJ, Martens JW, van den Ouweland AM, Alfredsson L, Palotie A, Peltonen-Palotie L, Irwanto A, Low HQ, Teoh GH, Thalamuthu A, Easton DF, Nevanlinna H, Liu J, Czene K, Hall P (2011) A combined analysis of genome-wide association studies in breast cancer. Breast Cancer Res Treat 126(3):717–727. https://doi.org/10.1007/s10549-010-1172-9
Maddocks ODK, Athineos D, Cheung EC, Lee P, Zhang T, van den Broek NJF, Mackay GM, Labuschagne CF, Gay D, Kruiswijk F, Blagih J, Vincent DF, Campbell KJ, Ceteci F, Sansom OJ, Blyth K, Vousden KH (2017) Modulating the therapeutic response of tumours to dietary serine and glycine starvation. Nature 544(7650):372–376. https://doi.org/10.1038/nature22056
Heber D, Byerly LO, Chlebowski RT (1985) Metabolic abnormalities in the cancer patient. Cancer 55 (1):225–229
Bi X, Henry CJ (2017) Plasma-free amino acid profiles are predictors of cancer and diabetes development. Nutr Diabetes 7(3):e249. https://doi.org/10.1038/nutd.2016.55
Miyagi Y, Higashiyama M, Gochi A, Akaike M, Ishikawa T, Miura T, Saruki N, Bando E, Kimura H, Imamura F, Moriyama M, Ikeda I, Chiba A, Oshita F, Imaizumi A, Yamamoto H, Miyano H, Horimoto K, Tochikubo O, Mitsushima T, Yamakado M, Okamoto N (2011) Plasma free amino acid profiling of five types of cancer patients and its application for early detection. PLoS ONE 6(9):e24143. https://doi.org/10.1371/journal.pone.0024143
Quevedo-Coli S, Crespi C, Benito E, Palou A, Roca P (1997) Alterations in circulating fatty acids and the compartmentation of selected metabolites in women with breast cancer. Biochem Mol Biol Int 41(1):1–10
Kuhn T, Sookthai D, Graf ME, Schubel R, Freisling H, Johnson T, Katzke V, Kaaks R (2017) Albumin, bilirubin, uric acid and cancer risk: results from a prospective population-based study. Br J Cancer 117(10):1572–1579. https://doi.org/10.1038/bjc.2017.313
Liu XA, Meng QH, Ye YQ, Hildebrandt MAT, Gu J, Wu XF (2015) Prognostic significance of pretreatment serum levels of albumin, LDH and total bilirubin in patients with non-metastatic breast cancer. Carcinogenesis 36(2):243–248. https://doi.org/10.1093/carcin/bgu247
Srivastava P, Hira SK, Srivastava DN, Gupta U, Sen P, Singh RA, Manna PP (2017) Protease-responsive targeted delivery of doxorubicin from bilirubin-BSA-capped mesoporous silica nanoparticles against colon cancer. ACS Biomater Sci Eng 3(12):3376–3385. https://doi.org/10.1021/acsbiomaterials.7b00635
Horsfall LJ, Rait G, Walters K, Swallow DM, Pereira SP, Nazareth I, Petersen I (2011) Serum Bilirubin and Risk of Respiratory Disease and Death. Jama-J Am Med Assoc 305(7):691–697. doi:https://doi.org/10.1001/jama.2011.124
Wen CP, Zhang FM, Liang D, Wen C, Gu J, Skinner H, Chow WH, Ye YQ, Pu X, Hildebrandt MAT, Huang MS, Chen CH, Hsiung CA, Tsai MK, Tsao CK, Lippman SM, Wu XF (2015) The ability of bilirubin in identifying smokers with higher risk of lung cancer: a large cohort study in conjunction with global metabolomic profiling. Clin Cancer Res 21(1):193–200. https://doi.org/10.1158/1078-0432.Ccr-14-0748
Temme EHM, Zhang JJ, Schouten EG, Kesteloot H (2001) Serum bilirubin and 10-year mortality risk in a Belgian population. Cancer Cause Control 12(10):887–894. doi:Doi 10.1023/A:1013794407325
Yadava N, Schneider SS, Jerry DJ, Kim C (2013) Impaired mitochondrial metabolism and mammary carcinogenesis. J Mammary Gland Biol 18(1):75–87. https://doi.org/10.1007/s10911-012-9271-3
Ahn CS, Metallo CM (2015) Mitochondria as biosynthetic factories for cancer proliferation. Cancer Metab. https://doi.org/10.1186/s40170-015-0128-2
Grzybowska-Szatkowska L, Rzymowska J, Slaska B (2016) Oxidative phosphorylation genes in breast cancer cells. Int J Mol Med 38:S78–S78
Marini C, Bianchi G, Buschiazzo A, Ravera S, Martella R, Bottoni G, Petretto A, Emionite L, Monteverde E, Capitanio S, Inglese E, Fabbi M, Bongioanni F, Garaboldi L, Bruzzi P, Orengo AM, Raffaghello L, Sambuceti G (2016) Divergent targets of glycolysis and oxidative phosphorylation result in additive effects of metformin and starvation in colon and breast cancer. Sci Rep. https://doi.org/10.1038/srep19569
Meric-Bernstam F, Evans K, Zheng XF, Su XP, Yuca E, Scott S, Akcakanat A, Ueno N, Lim B, Litton J, Valero V, Symmans F, Hortobagyi G, Perou C, Tripathy D, Draetta G, Marszalek J, Gonzalez-Angulo AM, Moulder S (2017) Oxidative phosphorylation as a target in triple negative breast cancer therapy. Cancer Res. https://doi.org/10.1158/1538-7445.Am2017-4970
Putignani L, Raffa S, Pescosolido R, Aimati L, Signore F, Torrisi MR, Grammatico P (2008) Alteration of expression levels of the oxidative phosphorylation system (OXPHOS) in breast cancer cell mitochondria. Breast Cancer Res Treat 110(3):439–452. https://doi.org/10.1007/s10549-007-9738-x
Phang JM, Liu W, Hancock CN, Fischer JW (2015) Proline metabolism and cancer: emerging links to glutamine and collagen. Curr Opin Clin Nutr 18(1):71–77. https://doi.org/10.1097/Mco.0000000000000121
Elia I, Broekaert D, Christen S, Boon R, Radaelli E, Orth MF, Verfaillie C, Grunewald TGP, Fendt SM (2017) Proline metabolism supports metastasis formation and could be inhibited to selectively target metastasizing cancer cells. Nat Commun. https://doi.org/10.1038/ncomms15267
Mouradian M, Kikawa KD, Dranka BP, Komas SM, Kalyanaraman B, Pardini RS (2015) Docosahexaenoic acid attenuates breast cancer cell metabolism and the Warburg phenotype by targeting bioenergetic function. Mol Carcinogen 54(9):810–820. https://doi.org/10.1002/mc.22151
Lewis GD, Farrell L, Wood MJ, Martinovic M, Arany Z, Rowe GC, Souza A, Cheng S, McCabe EL, Yang E, Shi X, Deo R, Roth FP, Asnani A, Rhee EP, Systrom DM, Semigran MJ, Vasan RS, Carr SA, Wang TJ, Sabatine MS, Clish CB, Gerszten RE (2010) Metabolic signatures of exercise in human plasma. Sci Transl Med 2(33):33ra37. https://doi.org/10.1126/scitranslmed.3001006
Acknowledgements
The study was supported by U01 CA179655 from NCI/NIH.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethics approval
All procedures performed in this study were approved by the Institutional Review Board at M D Anderson Cancer Center and in accordance with the ethical standards of 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Informed consent
Written informed consent was obtained from all participants.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original version of this article was revised: The sixth author name Krita A. Zanetti was mistakenly included as co-author. The author group was corrected.
Rights and permissions
About this article
Cite this article
Zhao, H., Shen, J., Moore, S.C. et al. Breast cancer risk in relation to plasma metabolites among Hispanic and African American women. Breast Cancer Res Treat 176, 687–696 (2019). https://doi.org/10.1007/s10549-019-05165-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-019-05165-4