Abstract
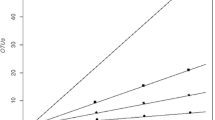
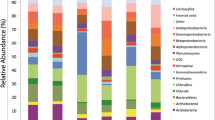
The bacterial diversity of two soil samples collected from the periphery of the Roopkund glacial lake and one soil sample from the surface of the Roopkund Glacier in the Himalayan ranges was determined by constructing three 16S rRNA gene clone libraries. The three clone libraries yielded a total of 798 clones belonging to 25 classes. Actinobacteria was the most predominant class (>10% of the clones) in the three libraries. In the library from the glacial soil, class Betaproteobacteria (24.2%) was the most predominant. The rarefaction analysis indicated coverage of 43.4 and 41.2% in the samples collected from the periphery of the lake thus indicating a limited bacterial diversity covered; at the same time, the coverage of 98.4% in the glacier sample indicated most of the diversity was covered. Further, the bacterial diversity in the Roopkund glacier soil was low, but was comparable with the bacterial diversity of a few other glaciers. The results of principal component analysis based on the 16S rRNA gene clone library data, percentages of OTUs and biogeochemical data revealed that the lake soil samples were different from the glacier soil sample and the biogeochemical properties affected the diversity of microbial communities in the soil samples.









Similar content being viewed by others
References
Abyzov SS, Mitskevich IN, Poglazova MN (1998) Microflora of the deep glacier horizons of central Antarctica. Microbiology 67:66–73
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Amato P, Hennebelle R, Magand O, Sancelme M, Delort AM, Barbante C, Boutron C, Ferrari C (2007) Bacterial characterization of the snow cover at Spitzberg, Svalbard. FEMS Microbiol Ecol 59:255–264
APHA (1995) Standard methods for the examination of water and wastewater, 19th edn. American Public Health Association, Washington, DC
Bill A (1994) The Nanda Devi affair, Penguin Books India, New Delhi. ISBN 0-14-024045-4
Blakemore LC, Searle PL, Daly BK (1987) Methods for chemical analysis of soils. NZ Soil Bureau Scientific Report 80
Bottos EM, Vincent CW, Greer WF, Whyte LG (2008) Prokaryotic diversity of arctic ice shelf microbial mats. Environ Microbiol 10:950–966
Bowman JP, McCuaig RD (2003) Biodiversity, community structural shifts, and biogeography of prokaryotes within Antarctic continental shelf sediment. Appl Environ Microbiol 69:2463–2483
Bowman JP, Rea SM, McCammon SA, McMeekin TA (2000) Diversity and community structure within anoxic sediment from marine salinity meromictic lakes and a coastal meromictic marine basin, Vestfold Hills, Eastern Antarctica. Environ Microbiol 2:227–237
Brambilla E, Hippe H, Hagelstein A, Tindall BJ, Stackebrandt E (2001) 16S rDNA diversity of cultured and uncultured prokaryotes of a mat sample from Lake Fryxell, McMurdo Dry Valleys, Antarctica. Extremophiles 5:23–33
Bray RH, Kurtz LT (1945) Determination of total organic and available forms of phosphorus in soils. Soil Sci 59:39–45
Burkert U, Warnecke F, Babenzien D, Zwirnmann E, Pernthaler J (2003) Members of a readily enriched beta-proteobacterial clade are common in surface waters of a humic lake. Appl Environ Microbiol 69:6550–6559
Carpenter EJ, Lin S, Capone DG (2000) Bacterial activity in South Pole snow. Appl Environ Microbiol 66:4514–4517
Certini G, Campbell CD, Edwards AC (2004) Rock fragments in soil support a different microbial community from the fine earth. Soil Biol Biochem 36:1119–1128
Cheng SM, Foght JM (2007) Cultivation-independent and -dependent characterization of bacteria resident beneath John Evans Glacier. FEMS Microbiol Ecol 59:318–330
Christner BC, Mosley-Thompson E, Thompson LG, Reeve JN (2001) Isolation of bacteria and 16S rDNAs from Lake Vostok accretion ice. Environ Microbiol 3:570–577
Christner BC, Mosley-Thompson E, Thompson LG, Reeve JN (2003) Bacterial recovery from ancient glacial ice. Environ Microbiol 5:433–436
Chun J, Lee J-H, Jung Y, Kim M, Kim S, Kim BK, Lim YW (2007) EzTaxon: a web-based tool for the identification of prokaryotes based on 16S ribosomal RNA gene sequences. Int J Syst Evol Microbiol 57:2259–2261
Costello EK, Schmidt SK (2006) Microbial diversity in alpine tundra wet meadow soil: novel Chloroflexi from a cold, water-saturated environment. Environ Microbiol 8:1471–1486
Dancer SJ, Shears P, Platt DJ (1997) Isolation and characterization of coliforms from glacial ice and water in Canada’s High Arctic. J Appl Microbiol 82:597–609
Eichorst SA, Breznak JA, Schmidt TM (2007) Isolation and characterization of soil bacteria that define Terriglobus gen. nov. in the phylum Acidobacteria. Appl Environ Microbiol 73:2708–2717
Fierer N, Schimel JP, Holden PA (2003) Controls on microbial CO2 production: a comparison of surface and subsurface soil horizons. Glob Chang Biol 9:1322–1332
Fierer N, Bradford MA, Jackson RB (2007) Toward an ecological classification of soil bacteria. Ecology 88:1354–1364
Foght J, Aislabie J, Turner S, Brown CE, Ryburn J, Saul DJ, Lawson W (2004) Culturable bacteria in subglacial sediments and ice from two southern hemisphere. Glaciers Microbiol Ecol 47:329–340
Gangwar P, Alam SI, Bansod S, Singh L (2009) Bacterial diversity of soil samples from the western Himalayas, India. Can J Microbiol 55:564–577
Glöckner FO, Zaichikov E, Belkova N, Denissova L, Pernthaler J, Pernthaler A, Amann R (2000) Comparative 16S rRNA analysis of lake bacterioplankton reveals globally distributed phylogenetic clusters including an abundant group of actinobacteria. Appl Environ Microbiol 66:5053–5065
Good IJ (1954) The population frequencies of species and the estimation of population parameters. Biometrica 40:237–264
Hur I, Chun J (2004) A method for comparing multiple bacterial community structures from 16S rDNA clone library sequences. J Microbiol 42:9–13
Jackson ML (1967) Soil chemical analysis. Prentice-Hall of India, New Delhi
Karr EA, Sattley WM, Jung DO, Madigan MT, Achenbach LA (2003) Remarkable diversity of phototrophic purple bacteria in a permanently frozen Antarctic lake. Appl Environ Microbiol 69:4910–4914
Karr EA, Ng JM, Belchik SM, Sattley WM, Madigan MT, Achenbach LA (2006) Biodiversity of methanogenic and other archaea in the permanently frozen Lake Fryxell, Antarctica. Appl Environ Microbiol 72:1663–1666
Kastovská K, Elster J, Stibal M, Santrůcková H (2005) Microbial assemblages in soil microbial succession after glacial retreat in Svalbard (High Arctic). Microbiol Ecol 50:396–407
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Kirchman DL, Yu L, Cottrell MT (2003) Diversity and abundance of uncultured cytophaga-like bacteria in the Delaware estuary. Appl Environ Microbiol 69:6587–6596
LaMontagne MG, Schimel JP, Holden PA (2003) Comparison of subsurface and surface soil bacterial communities in California grassland as assessed by terminal restriction fragment length polymorphisms of PCRamplified 16S rRNA genes. Microbiol Ecol 46:216–227
Lane DJ (1991) 16S/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M (eds) Nucleic acid techniques in bacterial systematics. Wiley, New York, pp 115–147
Lefauconnier B, Hagen JO, Rudant JP (1994) Flow speed and calving rate of Kongsbreen glacier, Svalbard, using SPOT images. Polar Res 10:56–65
Li S, **ao X, Yin X, Wang F (2006) Bacterial community along a historic lake sediment core of Ardley Island, west Antarctica. Extremophiles 10:461–467
Li Y, Li F, Zhang X, Qin S, Zeng Z, Dang H, Qin Y (2008) Vertical distribution of bacterial and archaeal communities along discrete layers of a deep-sea cold sediment sample at the East Pacific Rise (approximately 13 degrees N). Extremophiles 12:573–585
Liu Y, Yao T, Jiao N, Kang S, Zeng Y, Huang S (2006) Microbial community structure in moraine lakes and glacial meltwaters, Mount Everest. FEMS Microbiol Lett 265:98–105
Liu Y, Yao T, Jiao N, Kang S, Xu B, Zeng Y, Huang S, Liu X (2009) Bacterial diversity in the snow over Tibetan Plateau Glaciers. Extremophiles 13:411–423
López-García P, López-López A, Moreira D, Rodríguez-Valera F (2001) Diversity of free-living prokaryotes from a deep-sea site at the Antarctic Polar Front. FEMS Microbiol Ecol 36:193–202
Lysnes K, Thorseth IH, Steinsbu BO, Øvreås L, Torsvik T, Pedersen RB (2004) Microbial community diversity in seafloor basalt from the Arctic spreading ridges. FEMS Microbiol Ecol 50:213–230
Magurran AE (1996) Ecological diversity and its measurement. Chapman & Hall, London, pp 274–286
Marcy Y, Ouverney C, Bik EM, Losekann T, Ivanova N, Martin HG, Szeto E, Platt D, Hugenholtz P, Relman DA, Quake SR (2007) Dissecting biological ‘dark matter’ with single-cell genetic analysis of rare and uncultivated TM7 microbes from the human mouth. Proc Natl Acad Sci USA 104:11889–11894
Mikucki JA, Priscu JC (2007) Bacterial diversity associated with Blood Falls, a subglacial outflow from the Taylor Glacier, Antarctica. Appl Environ Microbiol 73:4029–4039
Mindl B, Anesio AM, Meirer K, Hodson AJ, Laybourn-Parry J, Sommaruga R, Sattler B (2007) Factors influencing bacterial dynamics along a transect from supraglacial runoff to proglacial lakes of a high Arctic glacier. FEMS Microbiol Ecol 59:307–317
Mosier AC, Murray AE, Fritsen CH (2007) Microbiota within the perennial ice cover of Lake Vida, Antarctica. FEMS Microbiol Ecol 59:274–288
Nealson KH (1999) Post-Viking microbiology: new approaches, new data, new insights. Orig Life Evol Biosph 29:73–93
Neufeld JD, Yu Z, Lam W, Mohn WW (2004) Serial analysis of ribosomal sequence tags (SARST): a high-throughput method for profiling complex microbial communities. Environ Microbiol 6:131–144
Niederberger TD, McDonald IR, Hacker AL, Soo RM, Barrett JE, Wall DH, Cary SC (2008) Microbial community composition in soils of Northern Victoria Land, Antarctica. Environ Microbiol 10:1713–1724
Perreault NN, Andersen DT, Pollard WH, Greer CW, Whyte LG (2007) Characterization of the prokaryotic diversity in cold saline perennial springs of the Canadian high Arctic. Appl Environ Microbiol 73:1532–1543
Prabagaran SR, Manorama R, Delille D, Shivaji S (2007) Predominance of Roseobacter, Sulfitobacter, Glaciecola and Psychrobacter in seawater collected off Ushuaia, Argentina, Sub-Antarctica. FEMS Microbiol Ecol 59:342–355
Priscu JC, Christner BC (2003) Microbial diversity and bioprocessing. In: Bull AT (ed) Earth’s icy biosphere. ASM press, Washington DC, pp 130–145
Reddy GSN, Aggarwal RK, Matsumotto GI, Shivaji S (2000) Arthrobacter flavus sp. nov., a psychrophilic bacterium isolate from a pond in McMurdo Dry Valley, Antarctica. Int J Syst Evol Microbiol 50:1553–1561
Reddy GSN, Uttam A, Shivaji S (2008) Bacillus cecembensis sp. nov., isolated from the Pindari glacier of the Indian Himalayas. Int J Syst Evol Microbiol 58:2330–2335
Reddy GSN, Pradhan S, Manorama R, Shivaji S (2009) Cryobacterium roopkundense sp. nov., a psychrophilic bacterium from a Himalayan glacier. Int J Syst Evol Microbiol 60:866–870
Reed AJ, Lutz RA, Vetriani C (2006) Vertical distribution and diversity of bacteria and archaea in sulfide and methane-rich cold seep sediments located at the base of the Florida Escarpment. Extremophiles 10:199–211
Rhoades JD (1982) Soluble salts. In: Page AL (ed) Methods of soil analysis. Chemical and microbiological properties, 2nd edn. Soil Science Society of America, Madison, pp 167–179
Segawa T, Miyamoto K, Ushida K, Agata K, Okada N, Kohshima S (2005) Seasonal change in bacterial flora and biomass in mountain snow from the Tateyama Mountains, Japan, analyzed by 16S rRNA gene sequencing and real-time PCR. Appl Environ Microbiol 71:123–130
Shivaji S, Reddy GSN, Aduri RP, Kutty R, Ravenschlag K (2004) Bacterial diversity of a soil sample from Schirmacher Oasis, Antarctica. Cell Mol Biol 50:525–536
Sjöling S, Cowan DA (2003) High 16S rDNA bacterial diversity in glacial meltwater lake sediment, Bratina Island, Antarctica. Extremophiles 7:275–282
Skidmore M, Anderson SP, Sharp M, Foght J, Lanoil BD (2005) Comparison of microbial community composition in two subglacial environments reveals a possible role for microbes in chemical weathering processes. Appl Environ Microbiol 71:6986–6997
Smith JJ, Tow LA, Stafford W, Cary C, Cowan DA (2006) Bacterial diversity in three different Antarctic Cold Desert mineral soils. Microbiol Ecol 51:413–421
Stackebrandt E, Brambilla E, Cousin S, Dirks W, Pukall R (2004) Culture-independent analysis of bacterial species from an anaerobic mat from Lake Fryxell, Antarctica: prokaryotic diversity revisited. Cell Mol Biol 50:517–524
Steven B, Briggs G, McKay CP, Pollard WH, Greer CW, Whyte LG (2007) Characterization of the microbial diversity in a permafrost sample from the Canadian high Arctic using culturedependent and culture-independent methods. FEMS Microbiol Ecol 59:513–523
Subbiah BH, Asija GL (1956) A rapid procedure for determination of available nitrogen in soils. Curr Sci 25:259–260
Suzuki K, Sasaki J, Uramoto M, Nakase T, Komagata K (1997) Cryobacterium psychrophilum gen. nov., sp. nov., nom. rev., comb. nov., an obligately psychrophilic actinomycete to accommodate ‘‘Curtobacterium psychrophilum’’ Inoue and Komagata 1976. Int J Syst Bacteriol 47:474–478
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Thompson JD, Higgins DG, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Tindall BJ (2004) Prokaryotic diversity in the Antarctic: the tip of the iceberg. Microb Ecol 47:271–283
Treves DS, **a B, Tiedje JM (2003) A two-species test of the hypothesis that spatial isolation influences microbial diversity in soil. Microbiol Ecol 45:20–28
Tsai YL, Olson BH (1991) Rapid method for direct extraction of DNA from soil and sediments. Appl Environ Microbiol 57:1070–1074
Tsai YL, Olson BH (1992) Rapid method for seperation of bacterial DNA from humic substances in sediments for polymerase chain reaction. Appl Environ Microbiol 58:2292–2295
Wagner D, Kobabe S, Liebner S (2009) Bacterial community structure and carbon turnover in permafrost-affected soils of the Lena Delta, northeastern Siberia. Can J Microbiol 55:77–83
Walkley A, Black CA (1934) An examination of Degtjareff methods for determining soil organic matter and a proposed modification of chromic acid titration method. Soil Sci 37:29–38
**ang SR, Yao TD, An LZ, Xu BQ, Li Z, Wu GJ, Wang YQ, Ma S, Chen XR (2004) Bacterial diversity in Malan ice core from the Tibetan Plateau. Folia Microbiol 49:269–275
**ang S, Yao T, An L, Xu B, Wang J (2005) 16S rRNA sequences and differences in bacteria isolated from the Muztag Ata glacier at increasing depths. Appl Environ Microbiol 71:4619–4627
Yergeau E, Newsham KK, Pearce DA, Kowalchuk GA (2007) Patterns of bacterial diversity across a range of Antarctic terrestrial habitats. Environ Microbiol 9:2670–2682
Zhang X, Tandong Y, Lizhe A, Lide T, Shijian X (2006) A study on the vertical profile of bacterial DNA structure in the Puruogangri (Tibetan Plateau) ice core using denaturing gradient gel electrophoresis. Ann Glaciol 43:160–166
Zhang X, Yao T, Tian L, Xu S, An L (2008) Phylogenetic and physiological diversity of bacteria isolated from Puruogangri ice core. Microbiol Ecol 55:476–488
Zhou J, Davey ME, Figueras JB, Rivkina E, Gilichinsky D, Tiedje JM (1997) Phylogenetic diversity of a bacterial community determined from Siberian tundra soil DNA. Microbiology 143:3913–3919
Zhou J, **a B, Treves DS, Wu L-Y, Marsh TL, O’Neill RV, Palumbo AV, Tiedje JM (2002) Spatial and resource factors influencing high microbial diversity in soil. Appl Environ Microbiol 68:326–334
Acknowledgments
We would like to thank the Department of Biotechnology and Council of Scientific and Industrial Research, Government of India for financial support to Dr. S. Shivaji. TNRS acknowledges the CSIR, Government of India, for the award of Research Associateship. We would like to thank Dr. M. K. Reddy, Scientist and Head and Dr. M. Kalita, Scientist from National Environmental Engineering Research Institute, Zonal Laboratory, Hyderabad, and Dr. M. K. Singh, Scientist from National Centre for Antarctic Ocean Research, Goa, for providing the data of the soil chemical characteristics.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by A. Oren.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Pradhan, S., Srinivas, T.N.R., Pindi, P.K. et al. Bacterial biodiversity from Roopkund Glacier, Himalayan mountain ranges, India. Extremophiles 14, 377–395 (2010). https://doi.org/10.1007/s00792-010-0318-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00792-010-0318-3