Abstract
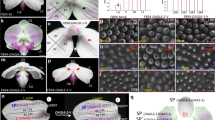
The AGAMOUS (AG) family of MADS-box genes plays important roles in controlling the development of the reproductive organs of flowering plants. To understand the molecular mechanisms behind the floral development in the orchid, we isolated and characterized two AG-like genes from Phalaenopsis that we denoted PhalAG1 and PhalAG2. Phylogenetic analysis indicated that PhalAG1 and PhalAG2 fall into different phylogenetic positions in the AG gene family as they belong to the C- and D-lineages, respectively. Reverse transcription-polymerase chair reaction (RT-PCR) analyses showed that PhalAG1 and PhalAG2 transcripts were detected in flower buds but not in vegetative organs. Moreover, in situ hybridization experiments revealed that PhalAG1 and PhalAG2 hybridization signals were observed in the lip, column, and ovule during the floral development of Phalaenopsis, with little difference between the expression patterns of the two genes. These results suggest that both AG-like genes in Phalaenopsis act redundantly with each other in floral development.








Similar content being viewed by others
References
Aharoni A, van Tunen AJ, Rosin FM, Hannapel DJ (1999) Isolation of AGAMOUS cDNA (STAG1) from Strawberry (Fragaria x ananassa cv. Elsanta). Plant Physiol 121:685–686
Ainsworth C, Crossley S, Buchanan-Wollaston V, Thangavelu M, Parker J (1995) Male and female flowers of the dioecious plant sorrel show different patterns of MADS box gene expression. Plant Cell 7:1583–1598
Angenent GC, Colombo L (1996) Molecular control of ovule development. Trends Plant Sci 1:228–232
Angenent GC, Franken J, Busscher M, van Dijken A, van Went JL, Dons HJM, van Tunen AJ (1995) A novel class of MADS box genes is involved in ovule development in Petunia. Plant Cell 7:1569–1582
Belarmino MM, Mii M (2000) Agrobacterium-mediated genetic transformation of a Phalaenopsis orchid. Plant Cell Rep 19:435–442
Benedito VA, Visser PB, van Tuy JM, Angenent GC, de Vries SC, Krens FA (2004) Ectopic expression of LLAG1, an AGAMOUS homologue from lily (Lilium longiflorum Thunb.) causes floral homeotic modifications in Arabidopsis. J Exp Bot 55:1391–1399
Boss PK, Vivier M, Matsumoto S, Dry IB, Thomas MR (2001) A cDNA from grapevine (Vitis vinifera L.), which shows homology to AGAMOUS and SHATTERPROOF, is not only expressed in flowers but also throughout berry development. Plant Mol Biol 45:541–553
Boss PK, Sensi E, Hua C, Davies C, Thomas MR (2002) Cloning and characterization of grapevine (Vitis vinifera L.) MADS-box genes expressed during inflorescence and berry development. Plant Sci 162:887–895
Bowman JL, Smyth DR, Meyerowitz EM (1989) Gene directing flower development in Arabidopsis. Plant Cell 1:37–52
Bradley D, Carpenter R, Sommer H, Hartley N, Coen E (1993) Complementary floral homeotic phenotypes result from opposite orientations of transposon at the plena locus of Antirrhinum. Cell 72:85–95
Brunner AM, Rottmann WH, Sheppard LA, Krutovskii K, DiFazio SP, Leonardi S, Strauss SH (2000) Structure and expression of duplicate AGAMOUS orthologues in poplar. Plant Mol Biol 44:619–634
Busi MV, Bustamante C, D’Angelo C, Hidalgo-Cuevas M, Boggio SB, Valle EM, Zabaleta E (2003) MADS-box genes expressed during tomato seed and fruit development. Plant Mol Biol 52:801–815
Cameron KM, Chase MW, Whitten WM, Kores PJ, Jarrell DC, Albert VC, Yukawa T, Hills HG, Goldman DH (1999) A phylogenetic analysis of the orchidaceae: evidence from RBCL nucleotide sequences. Am J Bot 86:208–224
Carpenter R, Coen ES (1990) Floral homeotic mutations produced by transposon-mutagenesis in Antirrhinum majus. Genes Dev 4:1483–1493
Coen ES, Meyerowitz EM (1991) The war of the whorls: genetic interactions controlling flower development. Nature 353:31–37
Colombo L, Franken J, Koetje E, Van Went J, Dons HJM, Angenent GC, van Tunen AJ (1995) The Petunia MADS-box gene FBP11 determines ovule identity. Plant Cell 7:1859–1868
Colombo L, Franken J, Van del Krol AR, Wittich PE, Dons HJM, Angenent GC (1997) Downregulation of ovule-specific MADS box genes from Petunia results in maternally controlled defects in seed development. Plant Cell 9:703–715
Cozzoline S, Widmer A (2005) Orchid diversity: an evolutionary consequence of deception? Trends Ecol Evol 20:487–494
Davies B, Schwarz-Sommer Z (1994) Control of floral organ identity by homeotic MADS box transcription factors. In: Nover L (ed) Results and problems in cell differentiation. Springer, Berlin Heidelberg New York, pp 235–258
Davies B, Motte P, Keck E, Saedler H, Sommer H, Schwarz-Sommer Z (1999) PLENA and FARINELLI: redundancy and regulatory interactions between two Antirrhinum MADS-box factors controlling flower development. EMBO J 18:4023–4034
Endress PK (1994) Diversity and evolutionary biology of tropical flowers. Cambridge University Press, Cambridge
Favaro R, Immink RGH, Ferioli V, Bernasconi B, Byzova M, Angenent GC, Kater M, Colombo L (2002) Ovule-specific MADS-box proteins have conserved protein–protein interactions in monocot and dicot plants. Mol Genet Genomics 268:152–159
Favaro R, Pinyopich A, Battaglia R, Kooiker M, Borghi L, Ditta G, Yanofsky MF, Kater MM, Colombo L (2003) MADS-box protein complexes control carpel and ovule development in Arabidopsis. Plant Cell 15:2603–2611
Felsenstein J (1985) Confidence limits on phylogenetics: an approach using the bootstrap. Evolution 39:783–791
Felsenstein J (2004) Inferring phylogenies. Sinauer Associates. Sunderland, Massachussetts
Force A, Lynch M, Pickett FB, Amores A, Yan Y-L, Postlethwait J (1999) Preservation of duplicate genes by complementary, degenerative mutations. Genetics 151:1531–1545
Frohman MA, Dush MK, Martin GR (1988) Rapid production of full-length cDNA from rare transcripts: amplification using a single gene-specific oligonucleotide primer. Proc Natl Acad Sci U S A 85:8998–9002
Galant R, Carroll SB (2002) Evolution of a transcriptional repression domain in an insect HOX protein. Nature 415:910–913
Gustafson-Brown C, Savidge B, Yanofsky MF (1994) Regulation of the Arabidopsis homeotic gene APETALA1. Cell 76:131–143
Hardenack S, Ye D, Saedler H, Grant S (1994) Comparison of MADS-box gene expression in develo** male and female flowers of the dioecious plant white campion. Plant Cell 6:1775–1787
Hughes AL (1999) Adaptive evolution of genes and genomes. Oxford University Press, Oxford
Jack T, Sieburth L, Meyerowitz E (1997) Targeted misexpression of AGAMOUS in whorl 2 of Arabidopsis flowers. Plant J 11:825–839
Jager M, Hassanin A, Manuel M, Le Guyader H, Deutsch J (2003) MADS-box genes in Ginkgo biloba and the evolution of the AGAMOUS family. Mol Biol Evol 20:842–854
Kang HG, Noh YS, Chung YY, Costa MA, An K, An G (1995) Phenotypic alterations of petal and sepal by ectopic expression of a rice MADS box gene in tobacco. Plant Mol Biol 29:1–10
Kang HG, Jeon JS, Lee S, An G (1998) Identification of class B and class C floral organ identity genes from rice plants. Plant Mol Biol 38:1021–1029
Kempin SA, Mandel MA, Yanofsky MF (1993) Conversion of perianth into reproductive organs by ectopic expression of the tobacco floral homeotic gene NAGI. Plant Physiol 103:1041–1046
Kitahara K, Matsumoto S (2000) Rose MADS-box genes ‘MASAKO C1 and D1’ homologous to class C floral identity genes. Plant Sci 151:121–134
Kramer EM, Alejandra-Jaramillo M, Di Stilio VS (2004) Patterns of gene duplication and functional evolution during the diversification of the AGAMOUS subfamily of MADS box genes in angiosperms. Genetics 166:1011–1023
Lamb RS, Irish VF (2003) Functional divergence within the APETALA3 / PISTILLATA floral homeotic gene lineages. Proc Natl Acad Sci U S A 100:6558–6563
Lemmetyinen J, Hassinen M, Elo A, Porali I, Keinonen K, Makela H, Sopanen T (2004) Functional characterization of SEPALLATA 3 and AGAMOUS orthologues in silver birch. Physiol Plant 121:149–162
Levine M (2002) How insects lose their limbs. Nature 415:848–849
Li QZ, Li XG, Bai SN, Lu WL, Zhang XS (2002) Isolation of HAG1 and its regulation by plant hormones during in vitro floral organogenesis in Hyacinthus orientalis L. Planta 215:533–540
Liu J, Huang Y, Ding B, Tauer CG (1999) cDNA cloning and expression of a sweetgum gene that shows homology with Arabidopsis AGAMOUS. Plant Sci 142:73–82
Lopez-Dee ZP, Wittich P, Enrico Pe M, Rigola D, Del Buono I, Gorla MS, Kater MM, Colombo L (1999) OsMADS13, a novel rice MADS-box gene expressed during ovule development. Dev Genet 25:237–244
Lynch M, Force A (2000) The probability of duplicate gene preservation by subfunctionalization. Genetics 154:459–473
Ma H, Yanofsky MF, Meyerowitz EM (1991) AGL1-AGL6, an Arabidopsis gene family with similarity to floral homeotic and transcription factor genes. Genes Dev 5:484–495
Mena M, Ambrose BA, Meeley RB, Briggs SP, Yanofsky MF, Schmidt RJ (1996) Diversification of C-function activity in maize flower development. Science 274:1537–1540
Murray MG, Thompson WF (1980) Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res 8:4321–4325
Münster T, Pahnke J, Di Rosa A, Kim JT, Martin W, Saedler H, Theissen G (1997) Floral homeotic genes were recruited from homologous MADS-box genes preexisting in the common ancestor of ferns and seed plants. Proc Natl Acad Sci U S A 94:2415–2420
Münster T, Deleu W, Wingen LU, Ouzunova M, Cacharron J, Faigl W, Werth S, Kim JTT, Saedler H, Theissen G (2002) Maize MADS-box genes galore. Maydica 47:287–301
Nakamura T, Song IJ, Fukuda T, Yokoyama J, Maki M, Ochiai T, Kameya T, Kanno A (2005) Characterization of TrcMADS1 gene of Trillium camtschatcense (Trilliaceae) reveals functional evolution of the SOC1/TM3-like gene family. J Plant Res 118:229–234
Nurrish SJ, Treisman R (1995) DNA binding specificity determinants in MADS-box transcriptional factors. Mol Cell Biol 15:4076–4085
Ohno S (1970) Evolution by gene duplication. Springer, Berlin Heidelberg New York
Pelaz S, Ditta GS, Baumann E, Wisman E, Yanofsky MF (2000) B and C floral organ identity functions require SEPALLATA MADS-box genes. Nature 405:200–203
Perrière G, Gouy M (1996) WWW-Query: an on-line retrieval system for biological sequence banks. Biochemie 78:364–369
Perl-Treves R, Kahana A, Rosenman N, **ang Y, Silberstein L (1998) Expression of multiple AGAMOUS-like genes in male and female flowers of cucumber (Cucumis sativus L.). Plant Cell Physiol 39:701:710
Pinyopich A, Ditta GS, Savidge B, Liljegren SJ, Baumann E, Wisman E, Yanofsky MF (2003) Assessing the redundancy of MADS-box genes during carpel and ovule development. Nature 424:85–88
Pnueli L, Hareven D, Rounsley SD, Yanofsky MF, Lifschitz E (1994) Isolation of tomato AGAMOUS gene TAGI and analysis of its role in transgenic plants. Plant Cell 6:163–173
Rasmussen FN (1985) Orchids. In: Dahlgren RMT, Clifford PF, Yeo PF (eds) The families of the monocotyledons. Springer, Berlin Heidelberg New York, pp 249–274
Rigola D, Pe ME, Fabrizio C, Me G, Sari-Gorla M (1998) CaMADS1, a MADS box gene expressed in the carpel of hazelnut. Plant Mol Biol 38:1147–1160
Ronshaugen M, McGinnis N, McGinnis W (2002) Hox protein mutation and macroevolution of the insect body plan. Nature 415:914–917
Rounsley SD, Ditta GS, Yanofsky MF (1995) Diverse roles for MADS box genes in Arabidopsis development. Plant Cell 7:1259–1269
Rutledge R, Regan S, Nicolas O, Fobert P, Côté C, Bosnich W, Kauffeldt C, Sunohara G, Séguin A, Stewart D (1998) Characterization of an AGAMOUS homologue from the conifer black spruce (Picea mariana) that produces floral homeotic conversions when expressed in Arabidopsis. Plant J 15:625–634
Sasaki T, Matsumoto T, Yamamoto K, Sakata K, Baba T, Katayose Y, Wu J, Niimura Y, Cheng Z, Nagamura Y, Antonio BA, Kanamori H, Hosokawa S, Masukawa M, Arikawa K, Chiden Y, Hayashi M, Okamoto M, Ando T, Aoki H, Arita K, Hamada M, Harada C, Hijishita S, Honda M, Ichikawa Y, Idonuma A, Iijima M, Ikeda M, Ikeno M, Ito S, Ito T, Ito Y, Ito Y, Iwabuchi A, Kamiya K, Karasawa W, Katagiri S, Kikuta A, Kobayashi N, Kono I, Machita K, Maehara T, Mizuno H, Mizubayashi T, Mukai Y, Nagasaki H, Nakashima M, Nakama Y, Nakamichi Y, Nakamura M, Namiki N, Negishi M, Ohta I, Ono N, Saji S, Sakai K, Shibata M, Shimokawa T, Shomura A, Song J, Takazaki Y, Terasawa K, Tsuji K, Waki K, Yamagata H, Yamane H, Yoshiki S, Yoshihara R, Yukawa K, Zhong H, Iwama H, Endo T, Ito H, Hahn JH, Kim HI, Eun MY, Yano M, Jiang J, Gojobori T (2002) The genome sequence and structure of rice chromosome 1. Nature 420:312–316
Schmidt RJ, Veit B, Mandel MA, Mena M, Hake S, Yanofsky MF (1993) Identification and molecular characterization of ZAG1, the maize homolog of the Arabidopsis floral homeotic gene AGAMOUS. Plant Cell 5:729–737
Schwarz-Sommer Z, Huijser P, Nacken W, Saedler H, Sommer H (1990) Genetic control of flower development by homeotic genes in Antirrhinum majus. Science 250:931–936
Shore P, Sharrocks AD (1995) The MADS-box family of transcription factors. Eur J Biochem 229:1–13
Tandre K, Albert VA, Sundas A, Engstrom P (1995) Conifer homologues to genes that control floral development in angiosperms. Plant Mol Biol 27:69–78
Theissen G (2001) Development of floral organ identity: stories from the MADS house. Curr Opin Plant Biol 4:75–85
Theissen G, Saedler H (2001) Floral quartets. Nature 409:469–471
Theissen G, Strater T, Fischer A, Saedler H (1995) Structural characterization, chromosomal localization and phylogenetic evaluation of two pairs of AGAMOUS-like MADS-box genes from maize. Genec156:155–166
Theissen G, Becker A, Di Rosa A, Kanno A, Kim JT, Münster T, Winter K-U, Saedler H (2000) A short history of MADS-box genes in plants. Plant Mol Biol 42:115–149
Theissen G, Becker A, Winter K-U, Münster T, Kirchner C, Saedler H (2002) How the land plants learned their floral ABCs: the role of MADS-box genes in the evolutionary origin of flowers. In: Cronk QCB, Bateman RM, Hawkins JA (eds) Developmental genetics and plant evolution. Taylor & Francis, London, pp 173–205
Thompson JD, Higgins DG, Gibson TJ (1994) Clustal W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Tsai WC, Kuoh CS, Chuang MH, Chen WH, Chen HH (2004) Four DEF-like MADS box genes displayed distinct floral morphogenetic roles in Phalaenopsis orchid. Plant Cell Physiol 45:831–844
Tsai WC, Lee PF, Chen HI, Hsiao YY, Wei WJ, Pan ZJ, Chuang MH, Kuoh CS, Chen WH, Chen HH (2005) PeMADS6, a GLOBOSA/PISTILLATA-like gene in Phalaenopsis equestris involved in petaloid formation, and correlated with flower longevity and ovary development. Plant Cell Physiol 46:1125–1139
Tsuchimoto S, van der Krol AR, Chua NH (1993) Ectopic expression of pMADS3 in transgenic petunia phenocopies the petunia blind mutant. Plant Cell 5:843–853
Tzeng TY, Chen HY, Yang CH (2002) Ectopic expression of carpel-specific MADS box genes from lily and lisianthus causes similar homeotic conversion of sepal and petal in Arabidopsis. Plant Physiol 130:1827–1836
Vandenbussche M, Theissen G, Van de Peer Y, Gerats T (2003) Structural diversification and neo-functionalization during floral MADS-box gene evolution by C-terminal frameshift mutations. Nucleic Acid Res 31:4401–4409
Wagner A (1999) Redundant gene functions and natural selection. J Evol Biol 12:1–16
Weigel D, Meyerowitz EM (1994) The ABCs of floral homeotic genes. Cell 78:203–209
Winter K-U, Becker A, Münster T, Kim JT, Saedler H, Theissen G (1999) MADS-box genes reveal that gnetophytes are more closely related to conifers than to flowering plants. Proc Natl Acad Sci U S A 96:7342–7347
Xu HY, Li XG, Li QZ, Bai SN, Lu WL, Zhang XS (2004) Characterization of HoMADS1 and its induction by plant hormones during in vitro ovule development in Hyacinthus orientalis L. Plant Mol Biol 55:209–220
Yanofsky MF, Ma H, Bowman JL, Drews GN, Feldmann KA, Meyerowitz EM (1990) The protein encoded by the Arabidopsis homeotic gene agamous resembles transcription factors. Nature 346:35–39
Yu D, Kotilainen M, Pollanen E, Mehto M, Elomaa P, Helariutta Y, Albert VA, Teeri TH (1999) Organ identity genes and modified patterns of flower development in Gerbera hybrida (Asteraceae). Plant J 17:51–62
Yun P-Y, Ito T, Kim S-Y, Kanno A, Kameya T (2004a) AVAG1 gene is involved in the development of reproductive organs in ornamental asparagus, Asparagus virgatus. Sex Plant Reprod 17:1–8
Yun P-Y, Kim S-Y, Ochiai T, Fukuda T, Ito T, Kanno A, Kameya T (2004b) AVAG2 is a putative D-class gene from an ornamental asparagus. Sex Plant Reprod 17:107–116
Acknowledgements
We deeply thank Professor H. Takahashi and S. Saito for technical assistance and also H. Tokairin for his collaboration in culturing the plants. We are also grateful to professor H. -Y. Lee, J. -H. Park, S. -S. Lee, Y. Ishikawa, T. Ochiai, P. -Y. Yun, B. -J. Park, Y. Mashiko, H. Ashizawa, A. Sato, M. Nakada, N. Kuroiwa, T. Shishido, R. Shinohara, M. Komatsu, Y. Akita, S. -Y. Kim, M. Hirai, T. Kamimura and H. Nakayama for providing much help and advice. This study was partly supported by a Grant-in-Aid for Scientific Research from the Ministry of Education, Culture, Sports, Science and Technology of Japan.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by G. Jürgens
Rights and permissions
About this article
Cite this article
Song, IJ., Nakamura, T., Fukuda, T. et al. Spatiotemporal expression of duplicate AGAMOUS orthologues during floral development in Phalaenopsis . Dev Genes Evol 216, 301–313 (2006). https://doi.org/10.1007/s00427-005-0057-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00427-005-0057-0