Abstract
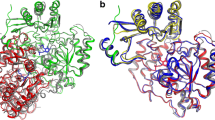
Understanding the metabolic potential of organisms or a bacterial community based on their (meta) genome requires the reliable prediction of an enzyme’s function from its amino acid sequence. Besides a remarkable development in prediction algorithms, the substrate scope of sequences with low identity to well-characterized enzymes remains often very elusive. From a recently conducted structure function analysis study of PLP-dependent enzymes, we identified a putative transaminase from Bacillus anthracis (Ban-TA) with the crystal structure 3N5M (deposited in the protein data bank in 2011, but not yet published). The active site residues of Ban-TA differ from those in related (class III) transaminases, which thereby have prevented function predictions. By investigating 50 substrate combinations its amine and ω-amino acid:pyruvate transaminase activity was revealed. Even though Ban-TA showed a relatively narrow amine substrate scope within the tested substrates, it accepts 2-propylamine, which is a prerequisite for industrial asymmetric amine synthesis. Structural information implied that the so-called dual substrate recognition of chemically different substrates (i.e. amines and amino acids) differs from that in formerly known enzymes. It lacks the normally conserved ‘flip**’ arginine, which enables dual substrate recognition by its side chain flexibility in other ω-amino acid:pyruvate transaminases. Molecular dynamics studies suggested that another arginine (R162) binds ω-amino acids in Ban-TA, but no side chain movements are required for amine and amino acid binding. These results, supported by mutagenesis studies, provide functional insights for the B. anthracis enzyme, enable function predictions of related proteins, and broadened the knowledge regarding ω-amino acid and amine converting transaminases.


Similar content being viewed by others
References
Bommer M, Ward JM (2013) A 1-step microplate method for assessing the substrate range of L-α-amino acid aminotransferase. Enzyme Microb Tech 52:218–225. doi:10.1016/j.enzmictec.2013.02.007
Cassimjee KE, Manta B, Himo F (2015) A quantum chemical study of the ω-transaminase reaction mechanism. Org Biomol Chem 13:8453–8464. doi:10.1039/c5ob00690b
Crismaru CG, Wybenga GG, Szymanski W, Wijma HJ, Wu B, Bartsch S, de Wildeman S, Poelarends GJ, Feringa BL, Dijkstra BW, Janssen DB (2013) Biochemical properties and crystal structure of a β-phenylalanine aminotransferase from Variovorax paradoxus. Appl Environ Microbiol 79:185–195. doi:10.1128/aem.02525-12
Davies MT (1959) A universal buffer solution for use in ultra-violet spectrophotometry. Analyst 84:248–251. doi:10.1039/an9598400248
Dey S, Lane JM, Lee RE, Rubin EJ, Sacchettini JC (2010) Structural characterization of the Mycobacterium tuberculosis biotin biosynthesis enzymes 7,8-diaminopelargonic acid synthase and dethiobiotin synthetase. Biochemistry 49:6746–6760. doi:10.1021/bi902097j
Duan Y, Wu C, Chowdhury S, Lee MC, **ong G, Zhang W, Yang R, Cieplak P, Luo R, Lee T, Caldwell J, Wang J, Kollman P (2003) A point-charge force field for molecular mechanics simulations of proteins based on condensed-phase quantum mechanical calculations. J Comput Chem 24:1999–2012. doi:10.1002/jcc.10349
Engelmark Cassimjee K, Branneby C, Abedi V, Wells A, Berglund P (2010) Transaminations with isopropyl amine: equilibrium displacement with yeast alcohol dehydrogenase coupled to in situ cofactor regeneration. Chem Comm 46:5569–5571. doi:10.1039/c0cc00050g
Essmann U, Perera L, Berkowitz ML, Darden T, Lee H, Pedersen LG (1995) A smooth particle mesh Ewald method. J Chem Phys 103:8577–8593. doi:10.1063/1.470117
Gand M, Müller H, Wardenga R, Höhne M (2014) Characterization of three novel enzymes with imine reductase activity. J Mol Catal B Enzym 110:126–132. doi:10.1016/j.molcatb.2014.09.017
Höhne M, Bornscheuer UT (2012) Application of transaminases in organic synthesis. In: May O, Gröger H, Drauz W (eds) Enzymes in organic synthesis. Wiley-VCH, Weinheim, pp. 779–820
Höhne M, Schätzle S, Jochens H, Robins K, Bornscheuer UT (2010) Rational assignment of key motifs for function guides in silico enzyme identification. Nat Chem Biol 6:807–813. doi:10.1038/nchembio.447
Ivanova N, Sorokin A, Anderson I, Galleron N, Candelon B, Kapatral V, Bhattacharyya A, Reznik G, Mikhailova N, Lapidus A, Chu L, Mazur M, Goltsman E, Larsen N, D’Souza M, Walunas T, Grechkin Y, Pusch G, Haselkorn R, Fonstein M, Dusko Ehrlich S, Overbeek R, Kyrpides N (2003) Genome sequence of Bacillus cereus and comparative analysis with Bacillus anthracis. Nature 423:87–91. doi:10.1038/nature01582
Kohls H, Steffen-Munsberg F, Höhne M (2014) Recent achievements in develo** the biocatalytic toolbox for chiral amine synthesis. Curr Opin Chem Biol 19:180–192. doi:10.1016/j.cbpa.2014.02.021
Laue H, Cook AM (2000) Biochemical and molecular characterization of taurine:pyruvate aminotransferase from the anaerobe Bilophila wadsworthia. Eur J Biochem 267:6841–6848. doi:10.1046/j.1432-1033.2000.01782.x
Midelfort KS, Kumar R, Han S, Karmilowicz MJ, McConnell K, Gehlhaar DK, Mistry A, Chang JS, Anderson M, Villalobos A, Minshull J, Govindarajan S, Wong JW (2013) Redesigning and characterizing the substrate specificity and activity of Vibrio fluvialis aminotransferase for the synthesis of imagabalin. Protein Eng Des Sel 26:25–33. doi:10.1093/protein/gzs065
Nakano Y, Tokunaga H, Kitaoka S (1977) Two ω-amino acid transaminases from Bacillus cereus. J Biochem 81:1375–1381
Økstad O, Kolstø A-B (2011) Genomics of Bacillus species. In: Wiedmann M, Zhang W (eds) Genomics of foodborne bacterial pathogens. Food Microbiology and Food Safety. Springer, New York, pp. 29–53. doi:10.1007/978-1-4419-7686-4_2
Ono H, Sawada K, Khunajakr N, Tao T, Yamamoto M, Hiramoto M, Shinmyo A, Takano M, Murooka Y (1999) Characterization of biosynthetic enzymes for ectoine as a compatible solute in a moderately halophilic eubacterium, Halomonas elongata. J Bacteriol 181:91–99
Park E-S, Shin J-S (2013) ω-transaminase from Ochrobactrum anthropi is devoid of substrate and product inhibitions. Appl Environ Microbiol 79:4141–4144. doi:10.1128/aem.03811-12
Radivojac P, Clark WT, Oron TR, Schnoes AM, Wittkop T, Sokolov A, Graim K, Funk C, Verspoor K, Ben-Hur A, Pandey G, Yunes JM, Talwalkar AS, Repo S, Souza ML, Piovesan D, Casadio R, Wang Z, Cheng J, Fang H, Gough J, Koskinen P, Toronen P, Nokso-Koivisto J, Holm L, Cozzetto D, Buchan DWA, Bryson K, Jones DT, Limaye B, Inamdar H, Datta A, Manjari SK, Joshi R, Chitale M, Kihara D, Lisewski AM, Erdin S, Venner E, Lichtarge O, Rentzsch R, Yang H, Romero AE, Bhat P, Paccanaro A, Hamp T, Kaszner R, Seemayer S, Vicedo E, Schaefer C, Achten D, Auer F, Boehm A, Braun T, Hecht M, Heron M, Honigschmid P, Hopf TA, Kaufmann S, Kiening M, Krompass D, Landerer C, Mahlich Y, Roos M, Bjorne J, Salakoski T, Wong A, Shatkay H, Gatzmann F, Sommer I, Wass MN, Sternberg MJE, Skunca N, Supek F, Bosnjak M, Panov P, Dzeroski S, Smuc T, Kourmpetis YAI, van Dijk ADJ, Braak CJF, Zhou Y, Gong Q, Dong X, Tian W, Falda M, Fontana P, Lavezzo E, Di Camillo B, Toppo S, Lan L, Djuric N, Guo Y, Vucetic S, Bairoch A, Linial M, Babbitt PC, Brenner SE, Orengo C, Rost B, Mooney SD, Friedberg I (2013) A large-scale evaluation of computational protein function prediction. Nat Meth 10:221–227. doi:10.1038/nmeth.2340
Rausch C, Lerchner A, Schiefner A, Skerra A (2013) Crystal structure of the ω-aminotransferase from Paracoccus denitrificans and its phylogenetic relationship with other class III aminotransferases that have biotechnological potential. Proteins Struct Funct Bioinf 81:774–787. doi:10.1002/prot.24233
Read TD, Peterson SN, Tourasse N, Baillie LW, Paulsen IT, Nelson KE, Tettelin H, Fouts DE, Eisen JA, Gill SR, Holtzapple EK, Okstad OA, Helgason E, Rilstone J, Wu M, Kolonay JF, Beanan MJ, Dodson RJ, Brinkac LM, Gwinn M, DeBoy RT, Madpu R, Daugherty SC, Durkin AS, Haft DH, Nelson WC, Peterson JD, Pop M, Khouri HM, Radune D, Benton JL, Mahamoud Y, Jiang L, Hance IR, Weidman JF, Berry KJ, Plaut RD, Wolf AM, Watkins KL, Nierman WC, Hazen A, Cline R, Redmond C, Thwaite JE, White O, Salzberg SL, Thomason B, Friedlander AM, Koehler TM, Hanna PC, Kolsto A-B, Fraser CM (2003) The genome sequence of Bacillus anthracis Ames and comparison to closely related bacteria. Nature 423:81–86. doi:10.1038/nature01586
Sayer C, Bommer M, Isupov M, Ward J, Littlechild J (2012) Crystal structure and substrate specificity of the thermophilic serine: pyruvate aminotransferase from Sulfolobus solfataricus. Acta Crystallogr D Biol Crystallogr 68:763–772. doi:10.1107/s0907444912011274
Schätzle S, Höhne M, Redestad E, Robins K, Bornscheuer UT (2009) Rapid and sensitive kinetic assay for characterization of ω-transaminases. Anal Chem 81:8244–8248. doi:10.1021/ac901640q
Steffen-Munsberg F, Vickers C, Kohls H, Land H, Mallin H, Nobili A, Skalden L, van den Bergh T, Joosten HJ, Berglund P, Hohne M, Bornscheuer UT (2015) Bioinformatic analysis of a PLP-dependent enzyme superfamily suitable for biocatalytic applications. Biotechnol Adv 33:566–604. doi:10.1016/j.biotechadv.2014.12.012
Steffen-Munsberg F, Vickers C, Thontowi A, Schätzle S, Meinhardt T, Svedendahl Humble M, Land H, Berglund P, Bornscheuer UT, Höhne M (2013a) Revealing the structural basis of promiscuous amine transaminase activity. ChemCatChem 5:154–157. doi:10.1002/cctc.201200545
Steffen-Munsberg F, Vickers C, Thontowi A, Schätzle S, Tumlirsch T, Svedendahl Humble M, Land H, Berglund P, Bornscheuer UT, Höhne M (2013b) Connecting unexplored protein crystal structures to enzymatic function. ChemCatChem 5:150–153. doi:10.1002/cctc.201200544
Strecker HJ (1953) Glutamic dehydrogenase. Arch Biochem Biophys 46:128–140. doi:10.1016/0003-9861(53)90176-3
Tamaki N, Kaneko M, Mizota C, Kikugawa M, Fujimoto S (1990) Purification, characterization and inhibition of D-3-aminoisobutyrate aminotransferase from the rat liver. Eur J Biochem 189:39–45. doi:10.1111/j.1432-1033.1990.tb15457.x
The PyMOL Molecular Graphics System, Version 1.6.0.0, Schrödinger, LCC, 2013
Toney MD (2011) Controlling reaction specificity in pyridoxal phosphate enzymes. Biochim Biophys Acta, Proteins Proteomics 1814:1407–1418. doi:10.1016/j.bbapap.2011.05.019
Tsai CS (1967) Spontaneous decarboxylation of oxalacetic acid. Can J Chem 45:873–880. doi:10.1139/v67-145
Váli Z, Kilár F, Lakatos S, Venyaminov SA, Závodszky P (1980) L-Alanine dehydrogenase from Thermus thermophilus. Biochim Biophys Acta, Enzymol 615:34–47. doi:10.1016/0005-2744(80)90006-6
Voellym R, Leisinger T (1976) Role of 4-aminobutyrate aminotransferase in the arginine metabolism of Pseudomonas aeruginosa. J Bacteriol 128:722–729
Wybenga GG, Crismaru CG, Janssen DB, Dijkstra BW (2012) Structural determinants of the β-selectivity of a bacterial aminotransferase. J Biol Chem 287:28495–28502. doi:10.1074/jbc.M112.375238
YASARA Structure, Version 14.7.17, YASARA Biosciences, 2014
Acknowledgments
FS thanks the Fonds der Chemischen Industrie (Chemiefonds-Stipendium) and PM thanks the Deutsche Forschungsgemeinschaft for financial support. We thank the European Union (KBBE-2011-5, grant no. 289350) and the COST Action Systems Biocatalysis (CM1303) for financial support within the European Union Seventh Framework Programme.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Funding
This study was funded by Deutsche Forschungsgemeinschaft (DFG grant number BO1862/16-1 and HO4754/4-1), Fonds der Chemischen Industrie (Chemiefonds-Stipendium), the European Union (KBBE-2011-5, grant no. 289350) and the COST Action Systems Biocatalysis (CM1303) within the European Union Seventh Framework Programme.
Conflict of interest
The authors declare that they have no conflict of interest.
Human and animal rights
This study did not contain research involving human participants or animals
Electronic supplementary material
ESM 1
(PDF 10.7 mb)
Rights and permissions
About this article
Cite this article
Steffen-Munsberg, F., Matzel, P., Sowa, M.A. et al. Bacillus anthracis ω-amino acid:pyruvate transaminase employs a different mechanism for dual substrate recognition than other amine transaminases. Appl Microbiol Biotechnol 100, 4511–4521 (2016). https://doi.org/10.1007/s00253-015-7275-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-015-7275-9