Abstract
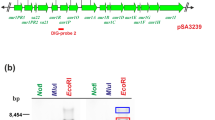
The pSKC2 cosmid, which has 32 kb and 28 open-reading frames, was isolated from Streptomyces kanamyceticus ATCC12853 as the gene cluster of kanamycin. This gene cluster includes the minimal biosynthetic genes of kanamycin with the resistance and regulatory genes. It was heterologously expressed in Streptomyces venezuelae YJ003, which has the advantage of fast growth, good efficiency of the transformation host, and rapid production of the aminoglycosides antibiotic. The isolated compound was analyzed by electrospray ionization–mass spectrometry, liquid chromatography–mass spectrometry, high-performance liquid chromatography, and tandem mass spectrometry and shows a molecular weight of 485 as kanamycin A.






Similar content being viewed by others
References
Baud H, Betencourt A, Peyre M, Penasse L (1977) Ribostamycin, as an intermediate in the biosynthesis of neomycin. J Antibiot 30:720–723
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Eustaquio AS, Gust B, Galm U, Li SM, Chater KF, Heide L (2005) Heterologous expression of novobiocin and clorobiocin biosynthetic gene clusters. Appl Environ Microbiol 71:2452–2459
Hong JS, Park SH, Choi CY, Sohng JK, Yoon YJ (2004) New olivosyl-derivatives of methymycin/pikromycin from an engineered strain of Streptomyces venezuelae. FEMS Microbiol Lett 238:391–399
Huang F, Haydock SF, Mironenko T, Spiteller D, Li Y, Spencer JB (2005) The neomycin biosynthetic gene cluster of Streptomyces fradiae NCIMB 8233: characterisation of an aminotransferase involved in the formation of 2-deoxystreptamine. Org Biomol Chem 3:1410–1418
Kharel MK, Basnet BD, Lee HC, Liou K, Woo JS, Sohng JK (2003a) Isolation and characterization of the tobramycin biosynthetic gene cluster from Streptomyces tenebrarius. FEMS Microbiol Lett 230:185–190
Kharel MK, Subba B, Lee HC, Liou K, Woo JS, Sohng JK (2003b) Identification of 2-deoxy-scyllo-inosose synthase in aminoglycoside producer Streptomyces. J Microbiol Biotechnol 13:828–831
Kharel MK, Subba B, Basnet BD, Woo JS, Lee HC, Sohng JK (2004) A gene cluster for biosynthesis of kanamycin from Streptomyces kanamyceticus: comparison with gentamicin biosynthetic gene cluster. Arch Biochem Biophys 429:204–214
Kharel MK, Subba B, Lee HC, Liou K, Sohng JK (2005) Characterization of L-glutamine: 2-deoxy-scyllo-inosose aminotransferase (tbmB) from Streptomyces tenebrarius. Bioorg Med Chem Lett 15:89–92
Kieser T, Bibb MJ, Buttner MJ, Chater KF, Hopwood DA (2000) Practical streptomyces genetics. John Innes Foundation, Norwich
Sambrook J, Russell DW (2001) Molecular cloning: a laboratory manual, 3rd edn. Cold Spring Harbor Laboratory, Cold Spring Harbor, New York
Stead DA, Richards RM (1996) Sensitive fluorimetric determination of gentamicin sulfate in biological matrices using solid-phase extraction, pre-column derivatization with 9-fluorenylmethyl chloroformate and reversed-phase high-performance liquid chromatography. J Chromatogr B Biomed Appl 675:295–302
Stead DA, Richards RM (1997) Sensitive high-performance liquid chromatographic assay for aminoglycosides in biological matrices enables the direct estimation of bacterial drug uptake. J Chromatogr, B, Biomed Sci Appl 693:415–421
Sztaricskai F (2001) The fight against bacteria: the antibiotics policy. Acta Pharm Hung 71:89–98
Xue Y, Zhao L, Liu HW, Sherman DH (1998) A gene cluster for macrolide antibiotic biosynthesis in Streptomyces venezuelae: Architecture of metabolic diversity. Proc Natl Acad Sci USA 95:12111–12116
Acknowledgments
This work was supported by Ministry of Commerce, Industry and Energy (grant no. 10023194) and by a grant (no. 20050401034682) from the BioGreen 21 Program, Rural Development Administration, Republic of Korea.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Thapa, L.P., Oh, TJ., Lee, H.C. et al. Heterologous expression of the kanamycin biosynthetic gene cluster (pSKC2) in Streptomyces venezuelae YJ003. Appl Microbiol Biotechnol 76, 1357–1364 (2007). https://doi.org/10.1007/s00253-007-1096-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-007-1096-4