Abstract
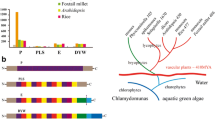
Prolamins are seed storage proteins in cereals and represent an important source of essential amino acids for feed and food. Genes encoding these proteins resulted from dispersed and tandem amplification. While previous studies have concentrated on protein sequences from different grass species, we now can add a new perspective to their relationships by asking how their genes are shared by ancestry and copied in different lineages of the same family of species. These differences are derived from alignment of chromosomal regions, where collinearity is used to identify prolamin genes in syntenic positions, also called orthologous gene copies. New or paralogous gene copies are inserted in tandem or new locations of the same genome. More importantly, one can detect the loss of older genes. We analyzed chromosomal intervals containing prolamin genes from rice, sorghum, wheat, barley, and Brachypodium, representing different subfamilies of the Poaceae. The Poaceae commonly known as the grasses includes three major subfamilies, the Ehrhartoideae (rice), Pooideae (wheat, barley, and Brachypodium), and Panicoideae (millets, maize, sorghum, and switchgrass). Based on chromosomal position and sequence divergence, it becomes possible to infer the order of gene amplification events. Furthermore, the loss of older genes in different subfamilies seems to permit a faster pace of divergence of paralogous genes. Change in protein structure affects their physical properties, subcellular location, and amino acid composition. On the other hand, regulatory sequence elements and corresponding transcriptional activators of new gene copies are more conserved than coding sequences, consistent with the tissue-specific expression of these genes.






Similar content being viewed by others
References
Albani D, Hammond-Kosack MC, Smith C, Conlan S, Colot V, Holdsworth M, Bevan MW (1997) The wheat transcriptional activator SPA: a seed-specific bZIP protein that recognizes the GCN4-like motif in the bifactorial endosperm box of prolamin genes. Plant Cell 9:171–184
Bagga S, Adams HP, Rodriguez FD, Kemp JD, Sengupta-Gopalan C (1997) Coexpression of the maize delta-zein and beta-zein genes results in stable accumulation of delta-zein in endoplasmic reticulum-derived protein bodies formed by beta-zein. Plant Cell 9:1683–1696
Baumgarten A, Cannon S, Spangler R, May G (2003) Genome-level evolution of resistance genes in Arabidopsis thaliana. Genetics 165:309–319
Boronat A, Martìnez MC, Reina M, Puigdomënech P, Palau J (1986) Isolation and sequencing of a 28 kD glutelin-2 gene from maize. Common elements in the 5′ flanking regions among zein and glutelin genes. Plant Science 47:95
Coleman CE, Herman EM, Takasaki K, Larkins BA (1996) The maize gamma-zein sequesters alpha-zein and stabilizes its accumulation in protein bodies of transgenic tobacco endosperm. Plant Cell 8:2335–2345
Conlan RS, Hammond-Kosack M, Bevan M (1999) Transcription activation mediated by the bZIP factor SPA on the endosperm box is modulated by ESBF-1 in vitro. Plant J 19:173–181
de Freitas FA, Yunes JA, da Silva MJ, Arruda P, Leite A (1994) Structural characterization and promoter activity analysis of the gamma-kafirin gene from sorghum. Mol Gen Genet 245:177–186
Esen A, Stetler DA (1992) Immunocytochemical localization of delta-zein in the protein bodies of maize endosperm cells. Am J Bot 79:243–248
Forde J, Malpica JM, Halford NG, Shewry PR, Anderson OD, Greene FC, Miflin BJ (1985) The nucleotide sequence of a HMW glutenin subunit gene located on chromosome 1A of wheat (Triticum aestivum L.). Nucleic Acids Res 13:6817–6832
Gale MD, Devos KM (1998) Comparative genetics in the grasses. Proc Natl Acad Sci USA 95:1971–1974
Gao S, Gu YQ, Wu J, Coleman-Derr D, Huo N, Crossman C, Jia J, Zuo Q, Ren Z, Anderson OD, Kong X (2007) Rapid evolution and complex structural organization in genomic regions harboring multiple prolamin genes in the polyploid wheat genome. Plant Mol Biol 65:189–203
Gibbon BC, Larkins BA (2005) Molecular genetic approaches to develo** quality protein maize. Trends Genet 21:227–233
Griffiths S, Sharp R, Foote TN, Bertin I, Wanous M, Reader S, Colas I, Moore G (2006) Molecular characterization of Ph1 as a major chromosome pairing locus in polyploid wheat. Nature 439:749–752
Gu YQ, Salse J, Coleman-Derr D, Dupin A, Crossman C, Lazo GR, Huo N, Belcram H, Ravel C, Charmet G, Charles M, Anderson OD, Chalhoub B (2006) Types and rates of sequence evolution at the high-molecular-weight glutenin locus in hexaploid wheat and its ancestral genomes. Genetics 174:1493–1504
Herman EM, Larkins BA (1999) Protein storage bodies and vacuoles. Plant Cell 11:601–614
Higo K, Ugawa Y, Iwamoto M, Korenaga T (1999) Plant cis-acting regulatory DNA elements (PLACE) database: 1999. Nucleic Acids Res 27:297–300
Holdsworth MJ, Munoz-Blanco J, Hammond-Kosack M, Colot V, Schuch W, Bevan MW (1995) The maize transcription factor Opaque-2 activates a wheat glutenin promoter in plant and yeast cells. Plant Mol Biol 29:711–720
Huo N, Lazo GR, Vogel JP, You FM, Ma Y, Hayden DM, Coleman-Derr D, Hill TA, Dvorak J, Anderson OD, Luo MC, Gu YQ (2007) The nuclear genome of Brachypodium distachyon: analysis of BAC end sequences. Functional and Integrated Genomics 8:135–147
Huo N, Vogel JP, Lazo GR, You FM, Ma Y, McMahon S, Dvorak J, Anderson OD, Luo MC, Gu YQ (2009) Structural characterization of Brachypodium genome and its syntenic relationship with rice and wheat. Plant Mol Biol 70:47–61
Katoh K, Toh H (2008) Recent developments in the MAFFT multiple sequence alignment program. Brief Bioinform 9:286–298
Kawagoe Y, Suzuki K, Tasaki M, Yasuda H, Akagi K, Katoh E, Nishizawa NK, Ogawa M, Takaiwa F (2005) The critical role of disulfide bond formation in protein sorting in the endosperm of rice. Plant Cell 17:1141–1153
Kawaura K, Mochida K, Ogihara Y (2005) Expression profile of two storage-protein gene families in hexaploid wheat revealed by large-scale analysis of expressed sequence tags. Plant Physiol 139:1870–1880
Kellogg EA (2001) Evolutionary history of the grasses. Plant Physiol 125:1198–1205
Kreis M, Forde BG, Rahman S, Miflin BJ, Shewry PR (1985) Molecular evolution of the seed storage proteins of barley, rye and wheat. J Mol Biol 183:499–502
Leister D (2004) Tandem and segmental gene duplication and recombination in the evolution of plant disease resistance gene. Trends Genet 20:116–122
Lending CR, Larkins BA (1989) Changes in the zein composition of protein bodies during maize endosperm development. Plant Cell 1:1011–1023
Levanony H, Rubin R, Altschuler Y, Galili G (1992) Evidence for a novel route of wheat storage proteins to vacuoles. J Cell Biol 119:1117–1128
Li X, Franceschi VR, Okita TW (1993) Segregation of storage protein mRNAs on the rough endoplasmic reticulum membranes of rice endosperm cells. Cell 72:869–879
Llaca V, Messing J (1998) Amplicons of maize zein genes are conserved within genic but expanded and constricted in intergenic regions. Plant J 15:211–220
Lohmer S, Maddaloni M, Motto M, Di Fonzo N, Hartings H, Salamini F, Thompson RD (1991) The maize regulatory locus Opaque-2 encodes a DNA-binding protein which activates the transcription of the b-32 gene. EMBO J 10:617–624
Marzabal P, Busk PK, Ludevid MD, Torrent M (1998) The bifactorial endosperm box of gamma-zein gene: characterisation and function of the Pb3 and GZM cis-acting elements. Plant J 16:41–52
Matsumoto T, Wu JZ, Kanamori H, Katayose Y, Fujisawa M, Namiki N, Mizuno H, Yamamoto K, Antonio BA, Baba T, Sakata K, Nagamura Y, Aoki H, Arikawa K, Arita K, Bito T, Chiden Y, Fujitsuka N, Fukunaka R, Hamada M, Harada C, Hayashi A, Hijishita S, Honda M, Hosokawa S, Ichikawa Y, Idonuma A, Iijima M, Ikeda M, Ikeno M, Ito K, Ito S, Ito T, Ito Y, Iwabuchi A, Kamiya K, Karasawa W, Kurita K, Katagiri S, Kikuta A, Kobayashi H, Kobayashi N, Machita K, Maehara T, Masukawa M, Mizubayashi T, Mukai Y, Nagasaki H, Nagata Y, Naito S, Nakashima M, Nakama Y, Nakamichi Y, Nakamura M, Meguro A, Negishi M, Ohta I, Ohta T, Okamoto M, Ono N, Saji S, Sakaguchi M, Sakai K, Shibata M, Shimokawa T, Song JY, Takazaki Y, Terasawa K, Tsugane M, Tsuji K, Ueda S, Waki K, Yamagata H, Yamamoto M, Yamamoto S, Yamane H, Yoshiki S, Yoshihara R, Yukawa K, Zhong HS, Yano M, Sasaki T, Yuan QP, Shu OT, Liu J, Jones KM, Gansberger K, Moffat K, Hill J, Bera J, Fadrosh D, ** SH, Johri S, Kim M, Overton L, Reardon M, Tsitrin T, Vuong H, Weaver B, Ciecko A, Tallon L, Jackson J, Pai G, Van Aken S, Utterback T, Reidmuller S, Feldblyum T, Hsiao J, Zismann V, Iobst S, de Vazeille AR, Buell CR, Ying K, Li Y, Lu TT, Huang YC, Zhao Q, Feng Q, Zhang L, Zhu JJ, Weng QJ, Mu J, Lu YQ, Fan DL, Liu YL, Guan JP, Zhang YJ, Yu SL, Liu XH, Zhang Y, Hong GF, Han B, Choisne N, Demange N, Orjeda G, Samain S, Cattolico L, Pelletier E, Couloux A, Segurens B, Wincker P, D’Hont A, Scarpelli C, Weissenbach J, Salanoubat M, Quetier F, Yu Y, Kim HR, Rambo T, Currie J, Collura K, Luo MZ, Yang TJ, Ammiraju JSS, Engler F, Soderlund C, Wing RA, Palmer LE, de la Bastide M, Spiegel L, Nascimento L, Zutavern T, O’Shaughnessy A, Dike S, Dedhia N, Preston R, Balija V, McCombie WR, Chow TY, Chen HH, Chung MC, Chen CS, Shaw JF, Wu HP, Hsiao KJ, Chao YT, Chu MK, Cheng CH, Hour AL, Lee PF, Lin SJ, Lin YC, Liou JY, Liu SM, Hsing YI, Raghuvanshi S, Mohanty A, Bharti AK, Gaur A, Gupta V, Kumar D, Ravi V, Vij S, Kapur A, Khurana P, Khurana JP, Tyagi AK, Gaikwad K, Singh A, Dalal V, Srivastava S, Dixit A, Pal AK, Ghazi IA, Yadav M, Pandit A, Bhargava A, Sureshbabu K, Batra K, Sharma TR, Mohapatra T, Singh NK, Messing J, Nelson AB, Fuks G, Kavchok S, Keizer G, Llaca ELV, Song RT, Tanyolac B, Young S, Il KH, Hahn JH, Sangsakoo G, Vanavichit A, de Mattos LAT, Zimmer PD, Malone G, Dellagostin O, de Oliveira AC, Bevan M, Bancroft I, Minx P, Cordum H, Wilson R, Cheng ZK, ** WW, Jiang JM, Leong SA, Iwama H, Gojobori T, Itoh T, Niimura Y, Fujii Y, Habara T, Sakai H, Sato Y, Wilson G, Kumar K, McCouch S, Juretic N, Hoen D, Wright S, Bruskiewich R, Bureau T, Miyao A, Hirochika H, Nishikawa T, Kadowaki K, Sugiura M (2005) The map-based sequence of the rice genome. Nature 436:793–800
Mena M, Vicente-Carbajosa J, Schmidt RJ, Carbonero P (1998) An endosperm-specific DOF protein from barley, highly conserved in wheat, binds to and activates transcription from the prolamin-box of a native B-hordein promoter in barley endosperm. Plant J 16:53–62
Messing J (2009) Synergy of two reference genomes for the grass family. Plant Physiol 149:117–124
Meyers BC, Kozik A, Griego A, Kuang H, Michelmore RW (2003) Genome-wide analysis of NBS-LRR-encoding genes in Arabidopsis. Plant cell 15:809–834
Michelmore RW, Meyers BC (1998) Clusters of resistance genes in plants evolve by divergent selection and a birth-and-death process. Genome Res 8:1113–1130
Moore G, Devos KM, Wang Z, Gale MD (1995) Cereal genome evolution, Grasses, line up and form a circle. Curr Biol 5:737–739
Muench DG, Ogawa M, Okita TW (1999) The prolamins of rice. In: Shewry PR, Casey R (eds) Seed proteins. Kluwer Academic Publishers, Dordrecht, pp 93–108
Muth JR, Muller M, Lohmer S, Salamini F, Thompson RD (1996) The role of multiple binding sites in the activation of zein gene expression by Opaque-2. Mol Gen Genet 252:723–732
Nakase M, Hotta H, Adachi T, Aoki N, Nakamura R, Masumura T, Tanaka K, Matsuda T (1996) Cloning of the rice seed alpha-globulin-encoding gene: sequence similarity of the 5′-flanking region to those of the genes encoding wheat high-molecular-weight glutenin and barley D hordein. Gene 170:223–226
Nakase M, Aoki N, Matsuda T, Adachi T (1997) Characterization of a novel rice bZIP protein which binds to the alpha-globulin promoter. Plant Mol Biol 33:513–522
Nei M, Gojobori T (1986) Simple methods for estimating the numbers of synonymous and nonsynonymous nucleotide substitutions. Mol Biol Evol 3:418–426
Okita TW, Krishnan HB, Kim WT (1988) Immunological relationships among the major seed proteins of cereals. Plant Sci 57:103–111
Onate L, Vicente-Carbajosa J, Lara P, Diaz I, Carbonero P (1999) Barley BLZ2, a seed-specific bZIP protein that interacts with BLZ1 in vivo and activates transcription from the GCN4-like motif of B-hordein promoters in barley endosperm. J Biol Chem 274:9175–9182
Onodera Y, Suzuki A, Wu CY, Washida H, Takaiwa F (2001) A rice functional transcriptional activator, RISBZ1, responsible for endosperm-specific expression of storage protein genes through GCN4 motif. J Biol Chem 276:14139–14152
Paterson AH, Bowers JE, Bruggmann R, Dubchak I, Grimwood J, Gundlach H, Haberer G, Hellsten U, Mitros T, Poliakov A, Schmutz J, Spannagl M, Tang H, Wang X, Wicker T, Bharti AK, Chapman J, Feltus FA, Gowik U, Grigoriev IV, Lyons E, Maher CA, Martis M, Narechania A, Otillar RP, Penning BW, Salamov AA, Wang Y, Zhang L, Carpita NC, Freeling M, Gingle AR, Hash CT, Keller B, Klein P, Kresovich S, McCann MC, Ming R, Peterson DG, Mehboobur R, Ware D, Westhoff P, Mayer KF, Messing J, Rokhsar DS (2009) The Sorghum bicolor genome and the diversification of grasses. Nature 457:551–556
Payne PI, Lancaster GJ (1983) Catalogue of alleles for the complex gene loci Glu-A1, Glu-B1 and Glu-D1, which code for the high-molecular-weight subunits of glutenin in hexaploid wheat. Cereal Res Commun 11:29–35
Payne PI, Corfield KG, Blackman JA (1981) Correlation between the inheritance of certain high-molecular-weight subunits of glutenin and bread-making quality in progenies of six crosses of bread wheat. J Sci Food Agric 32:51–60
Richly E, Kurth J, Leister D (2002) Mode of amplification and reorganization of resistance genes during recent Arabidopsis thaliana evolution. Mol Biol Evol 19:76–84
Schmidt RJ, Ketudat M, Aukerman MJ, Hoschek G (1992) Opaque-2 is a transcriptional activator that recognizes a specific target site in 22-kD zein genes. Plant Cell 4:689–700
Shewry PR, Tatham AS (1990) The prolamin storage proteins of cereal seeds: structure and evolution. Biochem J 267:1–12
Shewry PR, Tatham AS (1999) The characteristics, structure and evolutionary relationships of prolamins. In: Shewry PR, Casey R (eds) Seed proteins. Kluwer Academic Publishers, Dordrecht, The Netherlands, pp 11–33
Shewry PR, Miflin BJ, Kasarda DD (1984) The structural and evolutionary relationships of the prolamin storage proteins of barley, rye and wheat. Philos Trans R Soc Lond 304:297–308
Shewry PR, Tatham AS, Halford NG (1999) The prolamins of the Triticeae. In: Shewry PR, Casey R (eds) Seed Proteins. Kluwer Academic Publishers, Dordrecht, The Netherlands, pp 35–78
Shyur LF, Wen TN, Chen CS (1994) Purification and characterization of rice prolamins. Bot Bull Acad Sinica 35:65–71
Song R, Messing J (2003) Gene expression of a gene family in maize based on noncollinear haplotypes. Proc Natl Acad Sci USA 100:9055–9060
Song R, Llaca V, Messing J (2002) Mosaic organization of orthologous sequences in grass genomes. Genome Res 12:1549–1555
Swigonova Z, Lai J, Ma J, Ramakrishna W, Llaca V, Bennetzen JL, Messing J (2004) Close split of sorghum and maize genome progenitors. Genome Res 14:1916–1923
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Tanaka T, Antonio BA, Kikuchi S, Matsumoto T, Nagamura Y, Numa H, Sakai H, Wu J, Itoh T, Sasaki T, Aono R, Fujii Y, Habara T, Harada E, Kanno M, Kawahara Y, Kawashima H, Kubooka H, Matsuya A, Nakaoka H, Saichi N, Sanbonmatsu R, Sato Y, Shinso Y, Suzuki M, Takeda J, Tanino M, Todokoro F, Yamaguchi K, Yamamoto N, Yamasaki C, Imanishi T, Okido T, Tada M, Ikeo K, Tateno Y, Gojobori T, Lin YC, Wei FJ, Hsing YI, Zhao Q, Han B, Kramer MR, McCombie RW, Lonsdale D, O’Donovan CC, Whitfield EJ, Apweiler R, Koyanagi KO, Khurana JP, Raghuvanshi S, Singh NK, Tyagi AK, Haberer G, Fujisawa M, Hosokawa S, Ito Y, Ikawa H, Shibata M, Yamamoto M, Bruskiewich RM, Hoen DR, Bureau TE, Namiki N, Ohyanagi H, Sakai Y, Nobushima S, Sakata K, Barrero RA, Sato Y, Souvorov A, Smith-White B, Tatusova T, An S, An G, Oota S, Fuks G, Messing J, Christie KR, Lieberherr D, Kim H, Zuccolo A, Wing RA, Nobuta K, Green PJ, Lu C, Meyers BC, Chaparro C, Piegu B, Panaud O, Echeverria M (2008) The rice annotation project database (RAP-DB): 2008 update. Nucleic Acids Res 36:D1028–D1033
Ueda T, Waverczak W, Ward K, Sher N, Ketudat M, Schmidt RJ, Messing J (1992) Mutations of the 22- and 27-kD zein promoters affect transactivation by the Opaque-2 protein. Plant cell 4:701–709
Ueda T, Wang Z, Pham N, Messing J (1994) Identification of a transcriptional activator-binding element in the 27-kilodalton zein promoter, the -300 element. Mol Cell Biol 14:4350–4359
Vicente-Carbajosa J, Moose SP, Parsons RL, Schmidt RJ (1997) A maize zinc-finger protein binds the prolamin box in zein gene promoters and interacts with the basic leucine zipper transcriptional activator Opaque2. Proc Natl Acad Sci USA 94:7685–7690
Wang Z, Messing J (1998) Modulation of gene expression by DNA-protein and protein-protein interactions in the promoter region of the zein multigene family. Gene 223:333–345
Wang Z, Ueda T, Messing J (1998) Characterization of the maize prolamin box-binding factor-1 (PBF-1) and its role in the developmental regulation of the zein multigene family. Gene 223:321–332
Wicker T, Yahiaoui N, Guyot R, Schlagenhauf E, Liu ZD, Dubcovsky J, Keller B (2003) Rapid genome divergence at orthologous low molecular weight glutenin loci of the A and Am genomes of wheat. Plant Cell 15:1186–1197
Woo YM, Hu DW, Larkins BA, Jung R (2001) Genomics analysis of genes expressed in maize endosperm identifies novel seed proteins and clarifies patterns of zein gene expression. Plant Cell 13:2297–2317
Wu CY, Suzuki A, Washida H, Takaiwa F (1998) The GCN4 motif in a rice glutelin gene is essential for endosperm-specific gene expression and is activated by Opaque-2 in transgenic rice plants. Plant J 14:673–683
Xu JH, Messing J (2006) Maize haplotype with a helitron-amplified cytidine deaminase gene copy. BMC Genet 7:52
Xu JH, Messing J (2008a) Diverged copies of the seed regulatory Opaque-2 gene by a segmental duplication in the progenitor genome of rice, sorghum, and maize. Mol Plant 1:760–769
Xu JH, Messing J (2008b) Organization of the prolamin gene family provides insight into the evolution of the maize genome and gene duplications in grass species. Proc Natl Acad Sci USA 105:14330–14335
Yunes JA, Cord Neto G, da Silva MJ, Leite A, Ottoboni LM, Arruda P (1994) The transcriptional activator Opaque2 recognizes two different target sequences in the 22-kD-like alpha-prolamin genes. Plant Cell 6:237–249
Acknowledgments
The research described in this manuscript was supported by the Selman A. Waksman Chair in Molecular Genetics and a grant from the DOE (# DE-FG05-95ER20194) to JM. The Brachypodium sequence data was produced by the US Department of Energy Joint Genome Institute http://www.jgi.doe.gov/.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by J. Snape.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplemental Figure 1.
Sequence alignments of orthologous regions of prolamins Ory10 (A, B), Ory13a (C), and Ory16 (D) between rice and sorghum. Prolamins are showed in red colors, and conserved genes in black colors (PDF 198 kb)
Supplemental Figure 2.
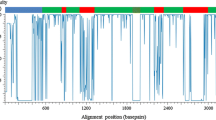
Amino acid sequence alignment of all prolamins from rice, sorghum, maize, wheat, barley, rye, Brachypodium with MAFFT program (Katoh and Toh 2008) with manually modified, repetitive domains of all prolamins are deleted, and S-poor prolamins (omega-gliadin, omega-secalin and B-hordein) are removed for analysis because of their most totally repetitive DNA. See Fig. 4 for details (PDF 169 kb)
Supplemental Figure 3.
Amino acid sequence alignment of HMW-like prolamin genes and related protein genes with MAFFT program (Katoh and Toh 2008) (PDF 55 kb)
Rights and permissions
About this article
Cite this article
Xu, JH., Messing, J. Amplification of prolamin storage protein genes in different subfamilies of the Poaceae . Theor Appl Genet 119, 1397–1412 (2009). https://doi.org/10.1007/s00122-009-1143-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-009-1143-x