Abstract
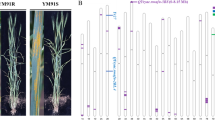
In cultivated barley (Hordeum vulgare ssp. vulgare), six-rowed spikes produce three times as many seeds per spike as do two-rowed spikes. The determinant of this trait is the Mendelian gene vrs1, located on chromosome 2H, which is syntenous with rice (Oryza sativa) chromosomes 4 and 7. We exploited barley–rice micro-synteny to increase marker density in the vrs1 region as a prelude to its map-based cloning. The rice genomic sequence, covering a 980 kb contig, identified barley ESTs linked to vrs1. A high level of conservation of gene sequence was obtained between barley chromosome 2H and rice chromosome 4. A total of 22 EST-based STS markers were placed within the target region, and the linear order of these markers in barley and rice was identical. The genetic window containing vrs1 was narrowed from 0.5 to 0.06 cM, which facilitated covering the vrs1 region by a 518 kb barley BAC contig. An analysis of the contig sequence revealed that a rice Vrs1 orthologue is present on chromosome 7, suggesting a transposition of the chromosomal segment containing Vrs1 within barley chromosome 2H. The breakdown of micro-collinearity illustrates the limitations of synteny cloning, and stresses the importance of implementing genomic studies directly in the target species.



Similar content being viewed by others
References
Akhunov ED, Goodyear AW, Geng S, Qi L-L, Echalier B, Gill BS, Miftahudin, Gustafson JP, Lazo G, Chao S, Anderson OD, Linkiewicz AM, Dubcovsky J, La Rota M, Sorrells ME, Zhang D, Nguyen HT, Kalavacharla V, Hossain K, Kianian SF, Peng J, Lapitan NLV, Gonzalez-Hernandez JL, Anderson JA, Choi D-W, Close TJ, Dilbirligi M, Gill KS, Walker-Simmons MK, Steber C, McGuire PE, Qualset CO, Dvorak J (2003) The organization and rate of evolution of wheat genomes are correlated with recombination rates along chromosome arms. Genome Res 13:753–763
Arumuganathan K, Earle ED (1991) Nuclear DNA content of some important plant species. Plant Mol Biol Rep 9:208–218
Bennetzen JL, Ma J (2003) The genetic colinearity of rice and other cereals on the basis of genomic sequence analysis. Curr Opin Plant Biol 6:128–133
Bothmer Rv, Jacobsen N (1985) Origin, taxonomy, and related species. In: Rasmusson D (ed) Barley-ASA agronomy monograph, vol 26. American Society of Agronomy, Crop Science Society of America, Soil Science Society of America, Madison, WI, pp 19–56
Bothmer Rv, Jacobsen N, Baden C, Jørgensen RB, Linde-Laursen I (1995) An ecogeographical study of the genus Hordeum, 2nd edn. Systematic and ecogeographic studies on crop genepools 7. International Plant Genetic Resources Institute, Rome
Brueggeman R, Rostoks N, Kudrna D, Kilian A, Han F, Chen J, Druka A, Steffenson B, Kleinhofs A (2002) The barley stem rust-resistance gene Rpg1 is a novel disease-resistance gene with homology to receptor kinases. Proc Natl Acad Sci USA 99:9328–9333
Brunner S, Keller B, Feuillet C (2003) A large rearrangement involving genes and low-copy DNA interrupts the microcollinearity between rice and barley at the Rph7 locus. Genetics 164:673–683
Caldwell KS, Langridge P, Powell W (2004) Comparative sequence analysis of the region harboring the hardness locus in barley and its colinear region in rice. Plant Physiol 136:3177–3190
Chantret N, Cenci A, Sabot F, Anderson O, Dubcovsky J (2004) Sequencing of the Triticum monococcum hardness locus reveals good microcolinearity with rice. Mol Gen Genomics 271:377–386
Collins NC, Thordal-Christensen H, Lipka V, Bau S, Kombrink E, Qiu J-L, Hückelhoven R, Stein M, Freialdenhoven A, Somerville SC, Schulze-Lefert P (2003) SNARE-protein-mediated disease resistance at the plant cell wall. Nature 425:973–977
Devos KM (2005) Updating the ‘Crop Circle’. Curr Opin Plant Biol 8:155–162
Doust AN, Devos KM, Gadberry MD, Gale MD, Kellogg EA (2004) Genetic control of branching in foxtail millet. Proc Natl Acad Sci USA 101:9045–9050
Dubcovsky J, Ramakrishna W, SanMiguel PJ, Busso CS, Yan L, Shiloff BA, Bennetzen JL (2001) Comparative sequence analysis of colinear barley and rice bacterial artificial chromosomes. Plant Physiol 125:1342–1353
Dunford RP, Yano M, Kurata N, Sasaki T, Huestis G, Rocheford T, Laurie DA (2002) Comparative map** of the barley Ppd-H1 photoperiod response gene region, which lies close to a junction between two rice linkage segments. Genetics 161:825–834
Feuillet C, Keller B (1999) High gene density is conserved at syntenic loci of small and large grass genomes. Proc Natl Acad Sci USA 96:8265–8270
Gale MD, Devos KM (1998) Comparative genetics in the grasses. Proc Natl Acad Sci USA 95:1971–1974
Gill KS, Gill BS, Endo TR, Boyko EV (1996) Identification and high-density map** of gene-rich regions in chromosome group 5 of wheat. Genetics 143:1001–1012
Gottwald S, Stein N, Börner A, Sasaki T, Graner A (2004) The gibberellic-acid insensitive dwarfing gene sdw3 of barley is located on chromosome 2HS in a region that shows high colinearity with rice chromosome 7L. Mol Gen Genomics 271:426–436
Griffee F (1925) Correlated inheritance of botanical characters in barley, and manner of reaction to Helminthosporium sativum. J Agric Res 30:915–935
Griffiths S, Sharp R, Foote TN, Bertin I, Wanous M, Reader S, Colas I, Moore G (2006) Molecular characterization of Ph1 as a major chromosome pairing locus in polyploid wheat. Nature 439:749–752
Harlan JR (1995) Barley. In: Smartt J, Simmonds NW (eds) Evolution of crop plants, 2nd edn. Longman, New York, pp 140–147
He C, Sayed-Tabatabaei BE, Komatsuda T (2004) AFLP targeting of the 1-cM region conferring the vrs1 gene for six-rowed spike in barley, Hordeum vulgare L. Genome 47:1122–1129
International Rice Genome Sequencing Project (IRGSP) (2005) The map-based sequence of the rice genome. Nature 436:793–800
Kleinhofs A, Kilian A, Saghai Maroof MA, Biyashev RM, Hayes P, Chen FQ, Lapitan N, Fenwick A, Blake TK, Kanazin V, Ananiev E, Dahleen L, Kudrna D, Bollinger J, Knapp SJ, Liu B, Sorrells M, Heun M, Franckowiak JD, Hoffman D, Skadsen R, Steffenson BJ (1993) A molecular, isozyme and morphological map of the barley (Hordeum vulgare) genome. Theor Appl Genet 86:705–712
Komatsuda T, Nakamura I, Takaiwa F, Oka S (1998) Development of STS markers closely linked to the vrs1 locus in barley, Hordeum vulgare. Genome 41:680–685
Komatsuda T, Pourkheirandish M, He C, Azhaguvel P, Kanamori H, Perovic D, Stein N, Graner A, Wicker T, Tagiri A, Lundqvist U, Fujimura T, Matsuoka M, Matsumoto T, Yano M (2007) Six-rowed barley originated from a mutation in a homeodomain-leucine zipper I–class homeobox gene. Proc Natl Acad Sci USA 104:1424–1429
Komatsuda T, Tanno K (2004) Comparative high resolution map of the six-rowed spike locus 1 (vrs1) in several populations of barley, Hordeum vulgare L. Hereditas 141:68–73
Künzel G, Korzun L, Meister A (2000) Cytologically integrated physical restriction fragment length polymorphism maps for the barley genome based on translocation breakpoints. Genetics 154:397–412
Laurie DA (1997) Comparative genetics of flowering time. Plant Mol Biol 35:167–177
Li W, Gill BS (2002) The colinearity of the Sh2/A1 orthologous region in rice, sorghum and maize is interrupted and accompanied by genome expansion in the Triticeae. Genetics 160:1153–1162
Lichten M, Goldman ASH (1995) Meiotic recombination hotspots. Annu Rev Genet 29:423–444
Makino T, Furusho M, Fukuoka T (1995) A mutant having six-rowed gene allelic to v locus. Barley Genet Newsl 24:122
Moore G (1995) Cereal genome evolution: pastoral pursuits with ‘Lego’ genomes. Curr Opin Genet Dev 5:717–724
Perovic D, Stein N, Zhang H, Drescher A, Prasad M, Kota R, Kopahnke D, Graner A (2004) An integrated approach for comparative map** in rice and barley with special reference to the Rph16 resistance locus. Funct Integr Genomics 4:74–83
Ramakrishna W, Dubcovsky J, Park Y-J, Busso C, Emberton J, SanMiguel P, Bennetzen JL (2002) Different types and rates of genome evolution detected by comparative sequence analysis of orthologous segments from four cereal genomes. Genetics 162:1389–1400
Robertson DW, Wiebe GA, Shands RG, Hagberg A (1965) A summary of linkage studies in cultivated barley, Hordeum species: supplement III, 1954–1963. Crop Sci 5:33–43
Rostoks N, Mudie S, Cardle L, Russell J, Ramsay L, Booth A, Svensson JT, Wanamaker SI, Walia H, Rodriguez EM, Hedley PE, Liu H, Morris J, Close TJ, Marshall DF, Waugh R (2005) Genome-wide SNP discovery and linkage analysis in barley based on genes responsive to abiotic stress. Mol Gen Genomics 274:515–527
Sambrook J, Russell DW (2001) Molecular cloning: a laboratory manual, 3rd edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY
SanMiguel P, Tikhonov A, ** Y-K, Motchoulskaia N, Zakharov D, Melake-Berhan A, Springer PS, Edwards KJ, Lee M, Avramova Z, Bennetzen JL (1996) Nested retrotransposons in the intergenic regions of the maize genome. Science 274:765–768
Schnable PS, Hsia A-P, Nikolau BJ (1998) Genetic recombination in plants. Curr Opin Plant Biol 1:123–129
Shepherd K, Islam AKMR (1981) Wheat: barley hybrids-the first eighty years. In: Evans RT, Peacock KW (eds) Wheat science today and tomorrow. Cambridge University Press, Cambridge, New York, pp 107–128
Stein N, Perovic D, Kumlehn J, Pellio B, Stracke S, Streng S, Ordon F, Graner A (2005) The eukaryotic translation initiation factor 4E confers multiallelic recessive Bymovirus resistance in Hordeum vulgare (L.). Plant J 42:912–922
Tarchini R, Biddle P, Wineland R, Tingey S, Rafalski A (2000) The complete sequence of 340 kb of DNA around the rice Adh1–Adh2 region reveals interrupted colinearity with maize chromosome 4. Plant Cell 12:381–391
Tikhonov AP, SanMiguel PJ, Nakajima Y, Gorenstein NM, Bennetzen JL, Avramova Z (1999) Colinearity and its exceptions in orthologous adh regions of maize and sorghum. Proc Natl Acad Sci USA 96:7409–7414
Turner A, Beales J, Faure S, Dunford RP, Laurie DA (2005) The pseudo-response regulator Ppd-H1 provides adaptation to photoperiod in barley. Science 310:1031–1034
Wei F, Gobelman-Werner K, Morroll SM, Kurth J, Mao L, Wing R, Leister D, Schulze-Lefert P, Wise RP (1999) The Mla (Powdery Mildew) resistance cluster is associated with three NBS-LRR gene families and suppressed recombination within a 240-kb DNA interval on chromosome 5S (1HS) of barley. Genetics 153:1929–1948
Wicker T, Stein N, Albar L, Feuillet C, Schlagenhauf E, Keller B (2001) Analysis of a contiguous 211 kb sequence in diploid wheat (Triticum monococcum L.) reveals multiple mechanisms of genome evolution. Plant J 26:307–316
Yan L, Loukoianov A, Tranquilli G, Helguera M, Fahima T, Dubcovsky J (2003) Positional cloning of the wheat vernalization gene VRN1. Proc Natl Acad Sci USA 100:6263–6268
Yu Y, Tomkins JP, Waugh R, Frisch DA, Kudrna D, Kleinhofs A, Brueggeman RS, Muehlbauer GJ, Wise RP, Wing RA (2000) A bacterial artificial chromosome library for barley (Hordeum vulgare L.) and the identification of clones containing putative resistance genes. Theor Appl Genet 101:1093–1099
Zohary D (1963) Spontaneous brittle six-rowed barley, their nature and origin. Barley Genetics I. In: Proceedings of the 1st International Barley Genet Symposium, Wageningen 1963, Pudoc, Wageningen, The Netherlands, pp 27–31
Acknowledgements
We thank M. Sameri, S. Okamura, M. Chono, J. Song and Y. Nagamura for their help and advice. We also thank M. Matsuoka, Y. Turuspekov and P. Azhaguvel for useful discussions. We thank R. Koebner for linguistic assistance in the preparation of this manuscript. The financial support of the Ministry of Agriculture, Forestry and Fisheries of Japan is acknowledged (Rice Genome Project MP1113b and Green Techno Project GD3006). M. Pourkheirandish was a research fellow supported by Japan International Research Center for Agricultural Sciences (JIRCAS).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by B. Keller.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Pourkheirandish, M., Wicker, T., Stein, N. et al. Analysis of the barley chromosome 2 region containing the six-rowed spike gene vrs1 reveals a breakdown of rice–barley micro collinearity by a transposition. Theor Appl Genet 114, 1357–1365 (2007). https://doi.org/10.1007/s00122-007-0522-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-007-0522-4