Abstract.
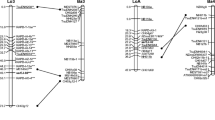
A genetic map of Pinus sylvestris was constructed using ESTP (expressed sequence tag polymorphism) markers and other gene-based markers, AFLP markers and microsatellites. Part of the ESTP markers (40) were developed and mapped earlier in Pinus taeda, and additional markers were generated based on P. sylvestris sequences or sequences from other pine species. The map** in P. sylvestris was based on 94 F1 progeny from a cross between plus-tree parents E635C and E1101. AFLP framework maps for the parent trees were first constructed. The ESTP and other gene sequence-based markers were added to the framework maps, as well as five published microsatellite loci. The separate maps were then integrated with the aid of AFLPs segregating in both trees (dominant segregation ratios 3:1) as well as gene markers and microsatellites segregating in both parent trees (segregation ratios 1:1:1:1 or 1:2:1). The integrated map consisted of 12 groups corresponding to the P. taeda linkage groups, and additionally three and six smaller groups for E1101 and E635C, respectively. The number of framework AFLP markers in the integrated map is altogether 194 and the number of gene markers 61. The total length of the integrated map was 1,314 cM. The set of markers developed for P. sylvestris was also added to existing maps of two P. taeda pedigrees. Starting with a mapped marker from one pedigree in the source species resulted in a mapped marker in a pedigree of the other species in more than 40% of the cases, with about equal success in both directions. The maps of the two species are largely colinear, even if the species have diverged more than 70 MYA. Most cases of different locations were probably due to problems in identifying the orthologous members of gene families. These data provide a first ESTP-containing map of P. sylvestris, which can also be used for comparing this species to additional species mapped with the same markers.


Similar content being viewed by others
References
Brown GR, Kadel EE, Bassoni DL, Kiehne KL, Temesgen B, Buijtenen JP, Sewell MM, Marshall KA, Neale DB (2001) Anchored reference loci in loblolly pine (Pinus taeda L.) for integrating pine genomics. Genetics 159:799–809
Campbell RB (1986) The interdependence of mating structure and inbreeding depression. Theor Pop Biol 30:232–244
Cato SA, Gardner RC, Kent J, Richardson TE (2001) A rapid PCR-based method for genetically map** ESTs. Theor Appl Genet 102:296–306
Chagné D, Brown G, Lalanne C, Madur D, Pot D, Neale D, Plomion C (2003) Comparative genome and QTL map** between maritime and loblolly pines. Mol Breed (in press)
Chang S, Puryear JD, Dias MADL, Funkhouser EA, Newton RJ, Cairney J (1996) Gene expression under water deficit in loblolly pine (Pinus taeda): isolation and characterization of cDNA clones. Physiol Plant 97:139–148
Chakravarti A, Lasher LK, Refer JE (1991) A maximum-likelihood method for estimating genome length using genetic linkage data. Genetics 128:175–182
Conkle MT (1981) Isozyme variation and linkage in six conifer species. Isozymes in North American Forest Trees and Forest Insects. Berkeley California, pp 11–17
Devey ME, Fiddler TA, Liu B-H, Knapp SJ, Neale DB (1994) An RFLP linkage map for loblolly pine based on a three-generation outbred pedigree. Theor Appl Genet 88:273–278
Devey ME, Bell JC, Smith DN, Neale DB, Moran GF (1996) A genetic linkage map for Pinus radiata based on RFLP, RAPD, and microsatellite markers. Theor Appl Genet 92:673–679
Devey ME, Sewell MM, Uren TL, Neale DB (1999) Comparative map** in loblolly and radiata pine using RFLP and microsatellite markers. Theor Appl Genet 99:656–662
Doudrick RL, Heslop-Harrison JS, Nelson CD, Schmidt T, Nance WL, Schwarzacher T (1995) Karyotype of slash pine (Pinus elliottii var. elliottii) using patterns of fluorescence in situ hybridization and fluorochrome banding. J Hered 86:289–296
Dvornyk V, Sirviö A, Mikkonen M, Savolainen O (2002) Low nucleotide diversity at the pal1 locus in the widely distributed Pinus sylvestris. Mol Biol Evol 19:179–199
Echt CS, Vendramin GG, Nelson CD, Marquart P (1999) Microsatellite DNA as shared genetic markers among conifer species. Can J For Res 29:365–371
Elsik CG, Williams CG (2001) Families of clustered microsatellites in a conifer genome. Mol Genet Genomics 265:535–542
Elsik CG, Minihan VT, Scarpa AM, Hall SE, Williams CG (2000) Low-copy microsatellites for Pinus taeda L. Genome 43:550–555
Groover A, Devey M, Fiddler T, Lee J, Megraw R, Mitchell-Olds T, Sherman B, Vujcic S, Williams C, Neale D (1994) Identification of quantitative trait loci influencing wood specific gravity in an outbred pedigree of loblolly pine. Genetics 138:1293–1300
Harry DE, Temesgen B, Neale DB (1998) Codominant PCR-based markers for Pinus taeda developed from mapped cDNA clones. Theor Appl Genet 97:327–336
Hizume F, Shibata F, Matsusaki Y, Garajova Z (2002) Chromosome identification and comparative karyotypic analysis of four Pinus species. Theor Appl Genet 105:491–497
Hurme P, Savolainen O (1999) Comparison of homology and linkage of RAPD markers between individual trees of Scots pine (Pinus sylvestris L.). Mol Ecol 8:15–22
Hurme P, Repo T, Pääkkönen T, Savolainen O (1997) Climatic adaptation of bud set and frost hardiness in Scots pine (Pinus sylvestris L.). Can J For Res 27:716–523
Hurme P, Sillanpää M, Repo T, Arjas E, Savolainen O (2000) Genetic basis of climatic adaptation in Scots pine by Bayesian QTL analysis. Genetics 156:1309–1322
Jermstad KD, Bassoni DL, Wheeler NC, Anekonda TS, Aitken SN, Adams WT, Neale DB (2001) Map** of quantitative trait loci controlling adaptive traits in coastal Douglas-fir. II. Spring and fall cold-hardiness. Theor Appl Genet 102:1152–1158
Karhu A, Dieterich J-H, Savolainen O (2000) Long persistence times and rapid expansion of microsatellites in pines. Mol Biol Evol 17:259–265
Kinlaw CS, Neale DB (1997) Complex gene families in pine genomes. Trends Plant Sci 2:356–359
Kostia S, Varvio S-L, Vakkari P, Pulkkinen P (1995) Microsatellite sequences in a conifer, Pinus sylvestris. Genome 38:1244–1248
Kriebel HB (1985) DNA sequence components of the Pinus strobus nuclear genome. Can J For Res 15:1–4
Lagercrantz U (1998) Comparative map** between Arabidopsis thaliana and Brassica nigra indicates that Brassica genomes have evolved through extensive genome replication accompanied by chromosome fusions and frequent rearrangements. Genetics 150:1217–1228
Lander E, Green P, Abrahamson J, Barlow A, Daley M, Lincoln S, Newburg L (1987) MAPMAKER: an interactive computer package for constructing primary genetic linkage maps of experimental and natural populations. Genomics 1:174–181
Lerceteau E, Plomion C, Andresson B (2001) AFLP map** and detection of quantitative trait loci (QTLs) for economically important traits in Pinus sylvestris: a preliminary study. Mol Breed 6:451–458
Lu MZ, Szmidt AE, Wang X-R (1995) Inheritance of RAPD fragments in haploid and diploid tissue of Pinus sylvestris (L.). Heredity 74:582–589
Meyers RM, Fischer SG, Maniatis T, Lerman LS (1985) Modification of the melting properties of duplex DNA by attachment of a GC-rich DNA sequence as determined by denaturing gradient gel electrophoresis. Nucleic Acids Res 13:3131–3145
Mikola J (1982) Bud-set phenology as an indicator of climatic adaptation of Scots pine in Finland. Silva Fennica 16:178–184
Myburg AA, Remington DL, O'Malley DM, Sederoff RR, Whetten RW (2001) High-throughput AFLP analysis using infrared dye-labeled primers and an automated DNA sequencer. BioTechniques 30:348–357
Niebling CR, Johnson K, Gerhold HD (1987) Electrophoretic analysis of genetic linkage in Scots pine (Pinus sylvestris L). Biochem Genet 25:803–814
Orita M, Iwahana H, Kanazawa H, Hayashi K, Sekiya T (1989) Detection of polymorphisms of human DNA by gel electrophoresis as single-strand conformation polymorphisms. Proc Natl Acad Sci USA 86:2766–2770
Perry DJ, Bousquet J (1998) Sequence-tagged-site markers of arbitrary genes: development, characterization and analysis of linkage in black spruce. Genetics 149:1089–1098
Perry DJ, Furnier GR (1996) Pinus banksiana has at least seven expressed alcohol dehydrogenase genes in two linked groups. Proc Natl Acad Sci USA 93:13,020–13,023
Plomion C, O'Malley DM, Durel CE (1995) Genomic analysis in maritime pine (Pinus pinaster). Comparison of two RAPD maps using selfed and open-pollinated seeds of the same individual. Theor Appl Genet 90:1028–1034
Plomion C, Hurme P, Frigerio J-M, Ridolphi M, Pot D, Pionneau C, Avila C, Gallardo F, David H, Neuttelings G, Campbell M, Canova FM, Savolainen O, Kremer A (1999) Develo** SSCP markers for comparative map** and QTL characterization in pines. Mol Breed 5:21–31
Prager EM, Fowler DP, Wilson AC (1976) Rates of evolution of conifers. Evolution 30:637–649
Price RA, Liston A, Strauss SH (1998) Phylogeny and systematics of Pinus. In: Richardson DM (ed) Ecology and biogeography of Pinus. Cambrdige University Press, Cambridge, pp 49–68
Remington DL, Whetten RW, Liu BH, O'Malley DM (1999) Construction of an AFLP genetic map with nearly complete genome coverage in Pinus taeda. Theor Appl Genet 98:1279–1292
Rozas J, Rozas R (1999) DnaSP version 3: an integrated program for molecular population genetics and molecular evolution analysis. Bioinformatics 15:174–175
Rudin D, Ekberg I (1978) Linkage studies in Pinus sylvestris L. using macrogametophyte allozymes. Silvae Genet 27:1–12
Sax K, Sax HJ (1933) Chromosome number and morphology in the conifers. J Arnold Arbor 14:356–375
Sewell M, Sherman BK, Neale DB (1999) A consensus map for loblolly pine (Pinus taeda L.). I. Construction and integration of individual linkage maps from two outbred three-generation pedigrees. Genetics 151:321–330
Soranzo N, Provan J, Powell W (1998) Characterization of microsatellite loci in Pinus sylvestris L. Mol Ecol 7:1260–1261
Stam P (1993) Construction of integrated genetic linkage maps by means of a new computer package: JoinMap. Plant J 3:739–744
Szmidt AE, Muona O (1989) Linkage relationships of allozyme loci in Pinus sylvestris. Hereditas 111:91–97
Temesgen B, Brown GR, Harry DE, Kinlaw CS, Sewell MM (2001) Genetic map** of expressed sequence tag polymorphism (ESTP) markers in loblolly pine (Pinus taeda L.). Theor Appl Genet 102:664–675
Voorrips RE (2001) MapChart v. 2.0: windows software for the graphical representation of linkage maps and QTLs. Plant Research International, Wageningen, The Netherlands
Vos P, Hogers R, Bleeker M, Rijans M, Van de Lee T, Hornes M, Frijters A, Pot J, Peleman J, Kuiper M, Zabeau M (1995) AFLP: a new technique for DNA fingerprinting. Nucleic Acids Res 23:4407–4414
Wakamiya I, Newton RJ, Johnston JS, Price HJ (1993) Genome size and environmental factors in the genus Pinus. Am J Bot 80:1235–1241
Yazdani R, Muona O, Rudin D, Szmidt AE (1985) Genetic structure of a Pinus sylvestris L. seed tree stand and a naturally regenerated understory. For Sci 31:430–436
Yazdani R, Yeh FC, Rimsha J (1995) Genomic map** of Pinus sylvestris (L.) using random amplified polymorphic DNA markers. For Genet 2:109–116
Acknowledgements.
This work was supported by a grant from the Finnish Technology Development Center (TEKES) and from the Environment and Natural Resources Research Council to OS. D.B.N. and G.R.B. are supported by USDA/NRI Plant Genome grants 95-37300 and 00-35300-9316. M.M. was supported by Academy of Finland grant no. 45692. We thank R.R. Sederoff for facilities and support during AFLP analyses at the NCSU, Raleigh, USA.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by C. Möllers
Rights and permissions
About this article
Cite this article
Komulainen, P., Brown, G.R., Mikkonen, M. et al. Comparing EST-based genetic maps between Pinus sylvestris and Pinus taeda . Theor Appl Genet 107, 667–678 (2003). https://doi.org/10.1007/s00122-003-1312-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-003-1312-2