Abstract
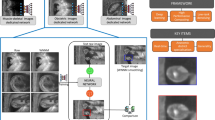
Medical Ultrasound (US), despite its wide use, is characterized by artefacts and operator dependency. Those attributes hinder the gathering and utilization of US datasets for the training of deep neural networks used for computer-assisted intervention systems. Data augmentation is commonly used to enhance model generalization and performance. However, common data augmentation techniques, such as affine transformations do not align with the physics of US and, when used carelessly can lead to unrealistic US images. To this end, we propose a set of physics-inspired transformations, including deformation, reverb and signal-to-noise ratio, that we apply on US B-mode images for data augmentation. We evaluate our method on a new spine US dataset for the tasks of bone segmentation and classification.
M. Tirindelli and C. Eilers—The authors contributed equally.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Van Sloun, R.J.G., Cohen, R., Eldar, Y.C.: Deep learning in ultrasound imaging. Proc. IEEE 108(1), 11–29 (2019)
Shorten, C., Khoshgoftaar, T.M.: A survey on image data augmentation for deep learning. J. Big Data 6(1), 1–48 (2019)
Goodfellow, I.J., et al.: Generative adversarial networks. ar**v preprint ar**v:1406.2661 (2014)
Zaman, A., Park, S.H., Bang, H., Park, C., Park, I., Joung, S.: Generative approach for data augmentation for deep learning-based bone surface segmentation from ultrasound images. Int J. Cars 15, 931–941 (2020). https://doi.org/10.1007/s11548-020-02192-1
Baka, N., Leenstra, S., van Walsum, T.: Ultrasound aided vertebral level localization for lumbar surgery. IEEE Trans. Med. Imaging 36(10), 2138–2147 (2017). https://doi.org/10.1109/TMI.2017.2738612
Duong, D.Q., et al.: Fully automated segmentation of alveolar bone using deep convolutional neural networks from intraoral ultrasound images. In: 41st Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), Berlin, Germany, pp. 6632–6635 (2019). https://doi.org/10.1109/EMBC.2019.8857060
Hohlmann, B., Glanz, J., Radermacher, K.: Segmentation of the distal femur in ultrasound images. Curr. Dir. Biomed. Eng. 6(1), 20200034 (2020)
Qi, X., Voar, N., Riera, L., Sarangi, A., Youssef, G., Vives, M., Hacihaliloglu, I.: Automatic Scan Plane Identification from 2D Ultrasound for Pedicle Screw Guidance. In: CAOS 2018 (EPiC Series in Health Sciences, vol. 2), pp. 168–174 (2018)
Benjdira, B., Ouni, K., Al Rahhal, M.M., Albakr, A., Al-Habib, A., Mahrous, E.: Spinal cord segmentation in ultrasound medical imagery. Appl. Sci. 10(4), 1370 (2020)
Alsinan, A. Z., Vives, M., Patel, V., Hacihaliloglu, I.: Spine surface segmentation from ultrasound using multi-feature guided CNN. In: CAOS 2019 (EPiC Series in Health Sciences), vol. 3, pp. 6–10 (2019)
Nguyen, K.C.T., et al.: Alveolar bone segmentation in intraoral ultrasonographs with machine learning. J. Dental Res. 99(9), 1054–1061 (2020). https://doi.org/10.1177/0022034520920593
Patel, H., Hacihaliloglu, I.: Improved automatic bone segmentation using large-scale simulated ultrasound data to segment real ultrasound bone surface data. In: IEEE 20th International Conference on Bioinformatics and Bioengineering (BIBE), Cincinnati, OH, USA, pp. 288–294 (2020). https://doi.org/10.1109/BIBE50027.2020.00054
Luan, K., Li, Z., Li, J.: An efficient end-to-end CNN for segmentation of bone surfaces from ultrasound. In: Computerized Medical Imaging and Graphics, vol. 84, p. 101766 (2020), ISSN 0895–6111. https://doi.org/10.1016/j.compmedimag.2020.101766
Ungi, T., et al.: Automatic spine ultrasound segmentation for scoliosis visualization and measurement. IEEE Trans. Biomed. Eng. 67(11), 3234–3241 (2020). https://doi.org/10.1109/TBME.2020.2980540
Bridge, C.P., Noble, J.A.: Object localisation in fetal ultrasound images using invariant features. In: 2015 IEEE 12th International Symposium on Biomedical Imaging (ISBI), pp. 156–159. IEEE (2015)
Ronneberger, O., Fischer, P., Brox, T.: U-Net: convolutional networks for biomedical image segmentation. In: Navab, N., Hornegger, J., Wells, W.M., Frangi, A.F. (eds.) MICCAI 2015. LNCS, vol. 9351, pp. 234–241. Springer, Cham (2015). https://doi.org/10.1007/978-3-319-24574-4_28
Huang, G., Liu, Z., Van Der Maaten, L., Weinberger, K.Q.: Densely connected convolutional networks. In: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, pp. 4700–4708 (2017)
Tirindelli, M., et al.: Force-ultrasound fusion: bringing spine robotic-us to the next “level.” IEEE Robot. Autom. Lett. 5(4), 5661–5668 (2020)
Esteban, J., et al.: Robotic ultrasound-guided facet joint insertion. Int. J. Comput. Assist. Radiol. Surg. 13(6), 895–904 (2018)
Hase, H., et al.: Ultrasound-guided robotic navigation with deep reinforcement learning. In: IEEE/RSJ International Conference on Intelligent Robots and Systems (IROS) (2020)
Wang, P., Vives, M., Patel, V.M., Hacihaliloglu, I.: Robust real-time bone surfaces segmentation from ultrasound using a local phase tensor-guided CNN. Int. J. Comput. Assist. Radiol. Surg. 15, 1127–1135 (2020)
Wang, P., Patel, V.M., Hacihaliloglu, I.: Simultaneous segmentation and classification of bone surfaces from ultrasound using a multi-feature guided CNN. In: Frangi, A.F., Schnabel, J.A., Davatzikos, C., Alberola-López, C., Fichtinger, G. (eds.) MICCAI 2018. LNCS, vol. 11073, pp. 134–142. Springer, Cham (2018). https://doi.org/10.1007/978-3-030-00937-3_16
Hetherington, J., Lessoway, V., Gunka, V., Abolmaesumi, P., Rohling, R.: SLIDE: automatic spine level identification system using a deep convolutional neural network. Int. J. Comput. Assist. Radiol. Surg. 12(7), 1189–1198 (2017)
Acknowledgements
This paper was partially funded by the Bayerische Forschungsstiftung, under Grant DOK-180-19, as well as the H2020 EU grant 688279 (EDEN2020) and the German Central Innovation Program for Small and Medium-sized Enterprises under grant agreement ZF4190502CR8 (PUMBA). We would also like to thank NVIDIA for the GPU donation.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 Springer Nature Switzerland AG
About this paper
Cite this paper
Tirindelli, M., Eilers, C., Simson, W., Paschali, M., Azampour, M.F., Navab, N. (2021). Rethinking Ultrasound Augmentation: A Physics-Inspired Approach. In: de Bruijne, M., et al. Medical Image Computing and Computer Assisted Intervention – MICCAI 2021. MICCAI 2021. Lecture Notes in Computer Science(), vol 12908. Springer, Cham. https://doi.org/10.1007/978-3-030-87237-3_66
Download citation
DOI: https://doi.org/10.1007/978-3-030-87237-3_66
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-87236-6
Online ISBN: 978-3-030-87237-3
eBook Packages: Computer ScienceComputer Science (R0)