Abstract
The small grass Brachypodium distachyon has attributes that make it an excellent model for the development and improvement of cereal crops and bioenergy feedstocks. To realize the potential of this system, many tools have been developed (e.g., the complete genome sequence, a large collection of natural accessions, a high density genetic map, BAC libraries, EST sequences, microarrays, etc.). In this chapter, we describe a high-efficiency transformation system, an essential tool for a modern model system. Our method utilizes the natural ability of Agrobacterium tumefaciens to transfer a well-defined region of DNA from its tumor-inducing (Ti) plasmid DNA into the genome of a host plant cell. Immature embryos dissected out of develo** B. distachyon seeds generate an embryogenic callus that serves as the source material for transformation and regeneration of transgenic plants. Embryogenic callus is cocultivated with A. tumefaciens carrying a recombinant plasmid containing the desired transformation sequence. Following cocultivation, callus is transferred to selective media to identify and amplify the transgenic tissue. After 2–5 weeks on selection media, transgenic callus is moved onto regeneration media for 2–4 weeks until plantlets emerge. Plantlets are grown in tissue culture until they develop roots and are transplanted into soil. Transgenic plants can be transferred to soil 6–10 weeks after cocultivation. Using this method with hygromycin selection, transformation efficiencies average 42 %, and it is routinely observed that 50–75 % of cocultivated calluses produce transgenic plants. The time from dissecting out embryos to having the first transgenic plants in soil is 14–18 weeks, and the time to harvesting transgenic seeds is 20–31 weeks.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Draper J, Mur LA, Jenkins G, Ghosh-Biswas GC, Bablak P, Hasterok R et al (2001) Brachypodium distachyon. A new model system for functional genomics in grasses. Plant Physiol 4:1539–1555
Garvin D, Gu Y, Hasterok R, Hazen S, Jenkins G, Mockler T et al (2008) Development of genetic and genomic research resources for Brachypodium distachyon, a new model system for grass crop research. Plant Genome 48:69–84
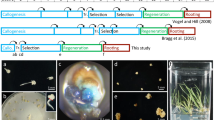
Vogel J, Bragg J (2009) Brachypodium distachyon, a new model for the Triticeae. In: Feuillet C, Muehlbauer GJ (eds) Genetics and genomics of the Triticeae. Springer, New York, pp 427–449
Bablak P, Draper J, Davey M, Lynch P (1995) Plant regeneration and micropropagation of Brachypodium distachyon. Tissue Organ Cult 42:97–107
Christiansen P, Andersen CH, Didion T, Folling M, Nielsen KK (2005) A rapid and efficient transformation protocol for the grass Brachypodium distachyon. Plant Cell Rep 23:751–758
Dai S, Zheng P, Marmey P, Zhang S, Tian W, Chen S et al (2001) Comparative analysis of transgenic rice plants obtained by Agrobacterium-mediated transformation and particle bombardment. Mol Breed 7:25–33
Kohli A, Twyman RM, Abranches R, Wegel E, Stoger E, Christou P (2003) Transgene integration, organization and interaction in plants. Plant Mol Biol 52:247–258
Svitashev S, Somers D (2002) Characterization of transgene loci in plants using FISH: a picture is worth a thousand words. Plant Cell Tissue Organ Cult 69:205–214
Tzfira T, Citovsky V (2006) Agrobacterium-mediated genetic transformation of plants: biology and biotechnology. Curr Opin Biotechnol 17:147–154
Feldmann K (1991) T-DNA insertion mutagenesis in Arabidopsis: mutational spectrum. Plant J 1:71–82
Alonso JM, Stepanova AN, Leisse TJ, Kim CJ, Chen H, Shinn P et al (2003) Genome-wide insertional mutagenesis of Arabidopsis thaliana. Science 301:653–657
Jeon J, Lee S, Jung K, Jun S, Jeong D, Lee J et al (2000) T-DNA insertional mutagenesis for functional genomics in rice. Plant J 22:561–570
Sallaud C, Meynard D, van Boxtel J, Gay C, Bès M, Brizard JP et al (2003) Highly efficient production and characterization of T-DNA plants for rice (Oryza sativa L.) functional genomics. Theor Appl Genet 106:1396–1408
Ma Y, Liu L, Zhu C, Sun C, Xu B, Fang J et al (2009) Molecular analysis of rice plants harboring a multi-functional T-DNA tagging system. J Genet Genomics 36:267–276
Vogel J, Garvin D, Leong O, Hayden D (2006) Agrobacterium-mediated transformation and inbred line development in the model grass Brachypodium distachyon. Plant Cell Tissue Organ Cult 85:199–211
Vogel J, Hill T (2008) High-efficiency Agrobacterium-mediated transformation of Brachypodium distachyon inbred line Bd21-3. Plant Cell Rep 27:471–478
Vain P, Worland B, Thole V, McKenzie N, Alves SC, Opanowicz M et al (2008) Agrobacterium-mediated transformation of the temperate grass Brachypodium distachyon (genotype Bd21) for T-DNA insertional mutagenesis. Plant Biotechnol J 6:236–245
Păcurar DI, Thordal-Christensen H, Nielsen KK, Lenk I (2008) A high-throughput Agrobacterium-mediated transformation system for the grass model species Brachypodium distachyon L. Transgenic Res 17:965–975
Komari T, Takakura Y, Ueki J, Kato N, Ishida Y, Hiei Y (2006) Binary vectors and super-binary vectors. Methods Mol Biol 343:15–41
Bragg JN, Wu J, Gordon SP, Guttman MA, Thilmony RL, Lazo GR, Gu YQ, Vogel JP (2012) Generation and characterization of the Western Regional Research Center Brachypodium T-DNA insertional mutant collection. PLoS One 7(9):e41916
Lazo GR, Stein PA, Ludwig RA (1991) A DNA transformation-competent Arabidopsis genomic library in Agrobacterium. Biotechnology 9:963–967
Acknowledgments
This work was supported by USDA CRIS project 5325-21000-017-00 and by the Office of Science (BER), US Department of Energy, Interagency Agreement No. DE-AI02-07ER64452.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer Science+Business Media New York
About this protocol
Cite this protocol
Bragg, J.N., Anderton, A., Nieu, R., Vogel, J.P. (2015). Brachypodium distachyon . In: Wang, K. (eds) Agrobacterium Protocols. Methods in Molecular Biology, vol 1223. Springer, New York, NY. https://doi.org/10.1007/978-1-4939-1695-5_2
Download citation
DOI: https://doi.org/10.1007/978-1-4939-1695-5_2
Published:
Publisher Name: Springer, New York, NY
Print ISBN: 978-1-4939-1694-8
Online ISBN: 978-1-4939-1695-5
eBook Packages: Springer Protocols