Abstract
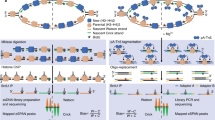
The replisome is a large protein machine containing multiple enzymatic activities needed to complete DNA replication. In addition to helicase and polymerases needed for copying the DNA, the replisome also contains proteins like DNA methyltransferases, histone chaperones, and chromatin modifying enzymes to couple DNA replication with chromatin deposition and establishment of the epigenetic code. In addition, since template DNA strands often contain DNA damage or other roadblocks to the replication machinery, replication stress response proteins associate with the replisome to stabilize, repair, and restart stalled replication forks. Hundreds of proteins are needed to accomplish these tasks. Identifying these proteins, monitoring their posttranslational modifications, and understanding how their activities are coordinated is essential to understand how the genome and epigenome are duplicated rapidly, completely, and accurately every cell division cycle. Here we describe an updated iPOND (isolation of proteins on nascent DNA) method to facilitate these analyses.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Sirbu BM, Couch FB, Cortez D (2012) Monitoring the spatiotemporal dynamics of proteins at replication forks and in assembled chromatin using isolation of proteins on nascent DNA. Nat Protoc 7:594–605
Sirbu BM, Couch FB, Feigerle JT et al (2011) Analysis of protein dynamics at active, stalled, and collapsed replication forks. Genes Dev 25: 1320–1327
Lopez-Contreras AJ, Ruppen I, Nieto-Soler M et al (2013) A proteomic characterization of factors enriched at nascent DNA molecules. Cell Rep 3:1105–1116
Sirbu BM, McDonald WH, Dungrawala H et al (2013) Identification of proteins at active, stalled, and collapsed replication forks using isolation of proteins on nascent DNA (iPOND) coupled with mass spectrometry. J Biol Chem 288:31458–31467
Salic A, Mitchison TJ (2008) A chemical method for fast and sensitive detection of DNA synthesis in vivo. Proc Natl Acad Sci U S A 105:2415–2420
Nagarajan P, Ge Z, Sirbu B et al (2013) Histone acetyl transferase 1 is essential for mammalian development, genome stability, and the processing of newly synthesized histones H3 and H4. PLoS Genet 9:e1003518
Acknowledgements
This work was supported by NCI grant R01CA136933 to D.C. We thank Bianca Sirbu and Frank Couch who contributed to the initial invention of iPOND.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer Science+Business Media New York
About this protocol
Cite this protocol
Dungrawala, H., Cortez, D. (2015). Purification of Proteins on Newly Synthesized DNA Using iPOND. In: Hancock, R. (eds) The Nucleus. Methods in Molecular Biology, vol 1228. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-1680-1_10
Download citation
DOI: https://doi.org/10.1007/978-1-4939-1680-1_10
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-1679-5
Online ISBN: 978-1-4939-1680-1
eBook Packages: Springer Protocols