Abstract
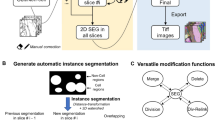
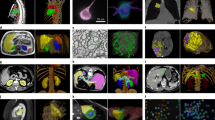
Zebrafish is a popular model system for biomedical analysis, especially for compound screening in drug research. In this paper, we present a comprehensive investigation aimed at enhancing the processing pipeline for segmenting zebrafish larvae images. The emphasis is on the application of an unsupervised segmentation method for segmenting zebrafish in Optical Projection Tomography (OPT) images. We propose a novel pipeline that integrates the Transformer and U-Net, a convolutional neural network for biomedical image segmentation, to achieve accurate segmentation of zebrafish larvae images. This accuracy is critical for precise 3D reconstruction. Leveraging transfer learning, we broaden the capabilities of our trained model to segment OPT images.
This approach is intended to enhance the robustness and versatility of our pipeline, allowing it to cater to a broad range of imaging modalities beyond traditional microscopic images. The developed processing pipeline is then used for 3D reconstruction of the segmented areas, demonstrating its potential for advanced biomedical analysis. Our findings confirm the efficiency and accuracy of the proposed pipeline providing robust tools for future Zebrafish-based research, particularly in the domains of drug screening and cancer treatment.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Mishra, Y.K., et al.: Zebrafish (danio rerio) as an ecotoxicological model for nanomaterial induced toxicity profiling. Precision Nanomed. 4(1), 750–782 (2021)
Wei, S., et al.: Perturbing tumor cell metabolism with a Ru (II) photo-redox catalyst to reverse the multidrug resistance of lung cancer. Sci. China Chem. 66(5), 1482–1488 (2023)
Nord, H., Dennhag, N., Muck, J., Hofsten, J.: Pax7 is required for establishment of the xanthophore lineage in zebrafish embryos. Mol. Biol. Cell 27(11), 1853–1862 (2016)
McCluskey, B.M., Postlethwait, J.H.: Phylogeny of zebrafish, a ‘model species’, within danio, a ‘model genus.’ Mol. Biol. Evol. 32(3), 635–652 (2015)
Barbazuk, W.B., et al.: The syntenic relationship of the zebrafish and human genomes. Genome Res. 10(9), 1351–1358 (2000)
Guo, Y., Veneman, W.J., Spaink, H.P., Verbeek, F.J.: Three-dimensional reconstruction and measurements of zebrafish larvae from high-throughput axial-view in vivo imaging. Biomed. Opt. Express 8(5), 2611–2634 (2017)
Phillips, J.B., Westerfield, M.: Zebrafish models in translational research: tip** the scales toward advancements in human health. Dis. Model. Mech. 7(7), 739–743 (2014)
Bradford, Y.M., et al.: Zebrafish models of human disease: gaining insight into human disease at ZFIN. ILAR J. 58(1), 4–16 (2017)
Liu, K., Petree, C., Requena, T., Varshney, P., Varshney, G.K.: Expanding the CRISPR toolbox in zebrafish for studying development and disease. Frontiers Cell Dev Biol. 7, 13 (2019)
Prykhozhij, S.V., Berman, J.N.: Zebrafish knock-ins swim into the mainstream, Disease Models and Mechanisms, vol. 11, no. 10, p. dmm037515 (2018)
Cagan, R.L., Zon, L.I., White, R.M.: Modeling cancer with flies and fish. Dev. Cell 49(3), 317–324 (2019)
Goessling, W., Sadler, K.C.: Zebrafish: an important tool for liver disease research. Gastroenterology 149(6), 1361–1377 (2015)
Rissone, A., Burgess, S.M.: Rare genetic blood disease modeling in zebrafish. Front. Genet. 9, 348 (2018)
Zhao, Y., Zhang, K., Sips, P., MacRae, C.A.: Screening drugs for myocardial disease in vivo with zebrafish: an expert update. Expert Opin. Drug Discov. 14(4), 343–353 (2019)
Griffin, A., Hamling, K.R., Hong, S., Anvar, M., Lee, L.P., Baraban, S.C.: Preclinical animal models for dravet syndrome: seizure phenotypes, comorbidities and drug screening. Front. Pharmacol. 9, 573 (2018)
Kalueff, A.V., et al.: Towards a comprehensive catalog of zebrafish behavior 1.0 and beyond. Zebrafish 10(1), 70–86 (2013)
Buda, M., Saha, A., Mazurowski, M.A.: Association of genomic subtypes of lower-grade gliomas with shape features automatically extracted by a deep learning algorithm. Comput. Biol. Med. 109, 218–225 (2019)
Chen, J., et al.: Transunet: transformers make strong encoders for medical image segmentation. ar**v preprint ar**v:2102.04306 (2021)
Guo, Y., **ong, Z., Verbeek, F.J.: An efficient and robust hybrid method for segmentation of zebrafish objects from bright-field microscope images. Mach. Vis. Appl. 29, 1211–1225 (2018)
Fukushima, H.C.S., et al.: Zebrafish toxicological screening could aid leishmaniosis drug discovery. Lab. Anim. Res. 37(1), 1–11 (2021)
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Nature Singapore Pte Ltd.
About this paper
Cite this paper
Javanmardi, S., Tang, X., Jahanbanifard, M., Verbeek, F.J. (2023). Unsupervised Segmentation of High-Throughput Zebrafish Images Using Deep Neural Networks and Transformers. In: Anutariya, C., Bonsangue, M.M. (eds) Data Science and Artificial Intelligence. DSAI 2023. Communications in Computer and Information Science, vol 1942. Springer, Singapore. https://doi.org/10.1007/978-981-99-7969-1_16
Download citation
DOI: https://doi.org/10.1007/978-981-99-7969-1_16
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-99-7968-4
Online ISBN: 978-981-99-7969-1
eBook Packages: Computer ScienceComputer Science (R0)