Abstract
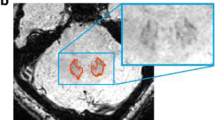
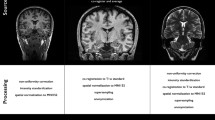
Dentate nuclei (DNs) segmentation is helpful for assessing their potential involvement in neurological diseases. Once DNs have been segmented, it becomes possible to investigate whether DNs are microstructurally affected, through analysis of quantitative MRI parameters, such as those derived from diffusion weighted imaging (DWI). This study developed a fully automated segmentation method using the non-DWI (b0) images from a DWI dataset to obtain DN masks inherently registered with parameter maps. Three different automatic methods were applied to healthy subjects: registration to SUIT (a spatially unbiased atlas template of the cerebellum and brainstem), OPAL (Optimized Patch Match for Label fusion) and CNN (Convolutional Neural Network). DNs manual segmentation was considered the gold standard. Results show that SUIT results have a Dice Similarity Coefficient (DSC) of 0.4907±0.0793 between automatic and gold standard masks. Comparing OPAL (DSC = 0.7624±0.1786) and CNN (DSC = 0.8658±0.0255), showed that a better performance was obtained with CNN. OPAL and CNN were optimised on high spatial resolution data from the Human Connectome Project. The three methods were then used to segment DNs of subjects with Temporal Lobe Epilepsy (TLE) from a 3T MRI research study with DWI data acquired with a coarser resolution. In TLE, SUIT performed similarly, with a DSC = 0.4145±0.1023. OPAL performed worse than using HCP data with a DSC of 0.4522±0.1178. CNN was able to extract the DNs without need for retraining and with a DSC = 0.7368±0.0799. Statistical comparison of quantitative parameters from DWI analysis, as well as volumes, revealed altered and lateralised changes in TLE patients compared to healthy controls. The proposed CNN is a viable option for accurate extraction of DNs from b0 images of DWI data with different resolutions and acquired at different sites.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Sure, D.R., Culicchia, F.: Duus’ Topical Diagnosis in Neurology. Thieme (2005)
Cattaneo, L.: Anatomia del sistema nervoso centrale e periferico dell’uomo. Monduzzi Editore (1989)
Habas, C.: Functional imaging of the deep cerebellar nuclei: A review. Cerebellum 9, 22–28 (2010). https://doi.org/10.1007/s12311-009-0119-3
Solbach, K., et al.: Cerebellar pathology in Friedreich’s ataxia: Atrophied dentate nuclei with normal iron content. NeuroImage Clin. 6, 93–99 (2014). https://doi.org/10.1016/j.nicl.2014.08.018
Fukutani, Y., et al.: Cerebellar dentate nucleus in Alzheimer’s disease with myoclonus. Dement. Geriatr. Cogn. Disord. 10, 81–88 (1999). https://doi.org/10.1159/000017106
Hermann, B.P., et al.: Cerebellar atrophy in temporal lobe epilepsy. Epilepsy Behav. 7, 279–287 (2005). https://doi.org/10.1016/j.yebeh.2005.05
Babb, T.L., et al.: Fastigiobulbar and dentatothalamic influences on hippocampal cobalt epilepsy in the cat. Electroencephalogr. Clin. Neurophysiol. 36, 141–154 (1974). https://doi.org/10.1016/0013-4694(74)90151-5
Krook-Magnuson, E., et al.: Cerebellar directed optogenetic intervention inhibits spontaneous hippocampal seizures in a mouse model of temporal lobe epilepsy. eNeuro. 1 (2014). https://doi.org/10.1523/ENEURO.0005-14.2014
Kros, L., et al.: Cerebellar output controls generalized spike-and-wave discharge occurrence. Ann. Neurol. 77, 1027–1049 (2015). https://doi.org/10.1002/ana.24399
Diedrichsen, J.: A spatially unbiased atlas template of the human cerebellum. Neuroimage. 33, 127–138 (2006). https://doi.org/10.1016/j.neuroimage.2006.05.056
Acosta-Cabronero, J., et al.: The whole-brain pattern of magnetic susceptibility perturbations in Parkinson’s disease. Brain. 140, 118–131 (2017). https://doi.org/10.1093/brain/aww278
Lindig, T., et al.: Pattern of Cerebellar Atrophy in Friedreich’s Ataxia: Using the SUIT Template. Cerebellum 18, 435–447 (2019). https://doi.org/10.1007/s12311-019-1008-z
Akram, H., et al.: Connectivity derived thalamic segmentation in deep brain stimulation for tremor. NeuroImage Clin. 18, 130–142 (2018). https://doi.org/10.1016/j.nicl.2018.01.008
Ye, C., et al.: Fully automatic segmentation of the dentate nucleus using diffusion weighted images. 1128–1131 (2012)
Bermudez Noguera, C., et al.: Using deep learning for a diffusion-based segmentation of the dentate nucleus and its benefits over atlas-based methods. J. Med. Imaging. 6, 1 (2019). https://doi.org/10.1117/1.jmi.6.4.044007
Bazin, P.-L., et al.: Automated Segmentation of Cerebellar Nuclei from Ultra-High-Field Quantitative Susceptibility maps with multi-atlas shape fusion. Proc. Jt. Annu. Meet. ISMRM-ESMRMB, Paris, Fr. 695 (2018)
Li, X., et al.: Multi-atlas tool for automated segmentation of brain gray matter nuclei and quantification of their magnetic susceptibility. Neuroimage 191, 337–349 (2019). https://doi.org/10.1016/j.neuroimage.2019.02.016
Jensen, J.H., Helpern, J.A.: MRI quantification of non-Gaussian water diffusion by kurtosis analysis. NMR Biomed. 23, 698–710 (2010). https://doi.org/10.1002/nbm.1518
Zhang, H., et al.: NODDI: Practical in vivo neurite orientation dispersion and density imaging of the human brain. Neuroimage 61, 1000–1016 (2012). https://doi.org/10.1016/j.neuroimage.2012.03.072
Van Essen, D.C., Smith, S.M., Barch, D.M., Behrens, T.E.J., Yacoub, E., Ugurbil, K.: The WU-Minn Human Connectome Project: An overview. Neuroimage 80, 62–79 (2013). https://doi.org/10.1016/j.neuroimage.2013.05.041
WU - Minn Consortium Human Connectome Project: WU-Minn HCP 1200 Subjects Data Release: Reference Manual. 2017, 1-169 (2017). www.humanconnectome.org/documentation/S900/
Alexander, A.L., et al.: Diffusion Tensor Imaging of the Brain. Neurotherapeutics 4, 316–329 (2007). https://doi.org/10.1021/jf505777p
Giraud, R., et al.: An Optimized PatchMatch for multi-scale and multi-feature label fusion. Neuroimage 124, 770–782 (2016). https://doi.org/10.1016/j.neuroimage.2015.07.076
Barnes, C., et al.: PatchMatch: A randomized correspondence algorithm for structural image editing. ACM Trans. Graph. 28 (2009). https://doi.org/10.1145/1576246.1531330
Perone, C.S., et al.: Spinal cord gray matter segmentation using deep dilated convolutions. Sci. Rep. (2018). https://doi.org/10.1038/s41598-018-24304-3
Khan, S., et al.: A Guide to Convolutional Neural Networks for Computer Vision. Morga Claypool (2018)
Aylward, et al.: Deep Learning for Medical Image Analysis. Elsevier (2017)
Ioffe, S., Szegedy, C.: Batch Normalization: Accelerating Deep Network Training by Reducing Internal Covariate Shift (2015). ar**v:1502.03167
Fidon, L., et al.: Generalised wasserstein dice score for imbalanced multi-class segmentation using holistic convolutional networks (2018). ar**v:1707.00478v4
Kingma, D.P., et al.: Adam: A Method for Stochastic Optimization (2017). ar**v:1412.6980
Prados, F., et al.: Spinal cord grey matter segmentation challenge. Neuroimage (2017). https://doi.org/10.1016/j.neuroimage.2017.03.010
Bonekamp, D., et al.: Diffusion tensor imaging in children and adolescents: Reproducibility, hemispheric, and age-related differences. Neuroimage 34, 733–742 (2007). https://doi.org/10.1016/j.neuroimage.2006.09.020
Mavroudis, I.A., et al.: Dendritic, axonal, and spinal pathology of the purkinje cells and the neurons of the dentate nucleus after long-term phenytoin administration: A case report. J. Child Neurol. 28, 1299–1304 (2013). https://doi.org/10.1177/0883073812455694
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this paper
Cite this paper
Gaviraghi, M. et al. (2021). Automatic Segmentation of Dentate Nuclei for Microstructure Assessment: Example of Application to Temporal Lobe Epilepsy Patients. In: Gyori, N., Hutter, J., Nath, V., Palombo, M., Pizzolato, M., Zhang, F. (eds) Computational Diffusion MRI. Mathematics and Visualization. Springer, Cham. https://doi.org/10.1007/978-3-030-73018-5_21
Download citation
DOI: https://doi.org/10.1007/978-3-030-73018-5_21
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-73017-8
Online ISBN: 978-3-030-73018-5
eBook Packages: Mathematics and StatisticsMathematics and Statistics (R0)