Abstract
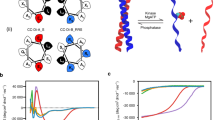
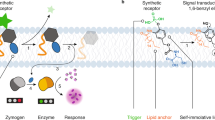
Intercellular membrane–membrane interfaces are compartments with specialized functions and unique biophysical properties that are essential in numerous cellular processes including cell signaling, development, and immunity. Using synthetic biology to engineer or to create novel cellular functions in the intercellular regions has led to an increasing need for a platform that allows generation of functionalized intercellular membrane–membrane interfaces. Here, we present a synthetic biology platform to engineer functional membrane–membrane interfaces using a pair of dimerizing proteins in both cell-free and cellular environments. We envisage this platform to be a helpful tool for synthetic biologists who wish to engineer novel intercellular signaling and communication systems.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Belardi B, Son S, Felce JH et al (2020) Cell–cell interfaces as specialized compartments directing cell function. Nat Rev Mol Cell Biol 21:750–764
Yang BA, Westerhof TM, Sabin K et al (2021) Engineered tools to study intercellular communication. Adv Sci (Weinh) 8:2002825
James JR, Vale RD (2012) Biophysical mechanism of T-cell receptor triggering in a reconstituted system. Nature 487:64–69
Otani T, Furuse M (2020) Tight junction structure and function revisited. Trends Cell Biol 30:805–817
Niessen CM (2007) Tight junctions/adherens junctions: basic structure and function. J Invest Dermatol 127:2525–2532
Südhof TC, Malenka RC (2008) Understanding synapses: past, present, and future. Neuron 60:469–476
Sjöqvist M, Andersson ER (2019) Do as I say, not(ch) as I do: lateral control of cell fate. Dev Biol 447:58–70
Prinz WA, Toulmay A, Balla T (2019) The functional universe of membrane contact sites. Nat Rev Mol Cell Biol 21:7–24
Feinberg EH, VanHoven MK, Bendesky A et al (2008) GFP Reconstitution Across Synaptic Partners (GRASP) defines cell contacts and synapses in living nervous systems. Neuron 57:353–363
Kanadome T, Hayashi K, Seto Y et al (2022) Development of intensiometric indicators for visualizing N-cadherin interaction across cells. Commun Biol 5:1–12
Schmid EM, Bakalar MH, Choudhuri K et al (2016) Size-dependent protein segregation at membrane interfaces. Nat Phys 12:704–711
Belardi B, Son S, Vahey MD et al (2019) Claudin-4 reconstituted in unilamellar vesicles is sufficient to form tight interfaces that partition membrane proteins. J Cell Sci 132:jcs221556
Freeman SA, Goyette J, Furuya W et al (2016) Integrins form an expanding diffusional barrier that coordinates phagocytosis. Cell 164:128–140
Zakeri B, Fierer JO, Celik E et al (2012) Peptide tag forming a rapid covalent bond to a protein, through engineering a bacterial adhesin. Proc Natl Acad Sci U S A 109:E690–E697
Feng S, Varshney A, Coto Villa D et al (2019) Bright split red fluorescent proteins for the visualization of endogenous proteins and synapses. Communications Biology 2:1–12
Moghimianavval H, Patel C, Mohapatra S et al (2022) Engineering functional membrane–membrane interfaces by InterSpy. Small 19:e2202104
Keeble AH, Turkki P, Stokes S et al (2019) Approaching infinite affinity through engineering of peptide-protein interaction. Proc Natl Acad Sci U S A 116:26523–26533
Moghimianavval H, Hsu YY, Groaz A et al (2022) In vitro reconstitution platforms of mammalian cell-free expressed membrane proteins. Methods Mol Biol 2433:105–120
Majumder S, Hsu YY, Moghimianavval H et al (2022) In vitro synthesis and reconstitution using mammalian cell-free lysates enables the systematic study of the regulation of LINC complex assembly. Biochemistry 61:1495–1507
Sharma B, Moghimianavval H, Hwang SW et al (2021) Synthetic cell as a platform for understanding membrane-membrane interactions. Membranes (Basel) 11:912
Groaz A, Moghimianavval H, Tavella F et al (2021) Engineering spatiotemporal organization and dynamics in synthetic cells. Wiley Interdiscip Rev Nanomed Nanobiotechnol 13. https://doi.org/10.1002/wnan.1685
Mátés L, Chuah MKL, Belay E et al (2009) Molecular evolution of a novel hyperactive slee** beauty transposase enables robust stable gene transfer in vertebrates. Nat Genet 41:753–761
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2024 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Moghimianavval, H., Mohapatra, S., Liu, A.P. (2024). A Mammalian-Based Synthetic Biology Toolbox to Engineer Membrane–Membrane Interfaces. In: Ceroni, F., Polizzi, K. (eds) Mammalian Synthetic Systems. Methods in Molecular Biology, vol 2774. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-3718-0_4
Download citation
DOI: https://doi.org/10.1007/978-1-0716-3718-0_4
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-3717-3
Online ISBN: 978-1-0716-3718-0
eBook Packages: Springer Protocols