Abstract
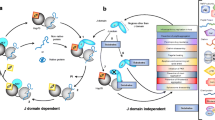
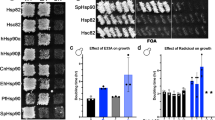
The development of mutant microorganisms lacking J domain proteins (JDPs; formerly called Hsp40s) has enabled the development of complementation assays for testing the co-chaperone function of JDPs. In these assays, an exogenously expressed novel JDP is tested for its ability to functionally substitute for a non-expressed or nonfunctional endogenous JDP(s) by reversing a stress phenotype. For example, the in vivo functionality of prokaryotic JDPs can be tested on the basis of their ability to reverse the thermosensitivity of a dnaJ cbpA mutant strain of the bacterium Escherichia coli (OD259). Similarly, the in vivo functionality of eukaryotic JDPs can be assessed in a thermosensitive ydj1 mutant strain of the yeast Saccharomyces cerevisiae (JJ160). Here we outline the use of these thermosensitive microorganisms in complementation assays to functionally characterize a JDP from the bacterium, Agrobacterium tumefaciens (AgtDnaJ), and a JDP from the trypanosomal parasite, Trypanosoma cruzi (TcJ2).
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Deloche O, Kelley WL, Georgopoulos C (1997) Structure-function analyses of the Ssc1p, Mdj1p, and Mge1p Saccharomyces cerevisiae mitochondrial proteins in Escherichia coli. J Bacteriol 179:6066–6075
Johnson JL, Craig EA (2001) An essential role for the substrate-binding region of Hsp40s in Saccharomyces cerevisiae. J Cell Biol 152:851–856
Shonhai A, Boshoff A, Blatch GL (2005) Plasmodium falciparum heat shock protein 70 is able to suppress the thermosensitivity of an Escherichia coli DnaK mutant strain. Mol Gen Genomics 274:70–78
Edkins AL, Ludewig MH, Blatch GL (2004) A Trypanosoma cruzi heat shock protein 40 is able to stimulate the adenosine triphosphate hydrolysis activity of heat shock protein 70 and can substitute for a yeast heat shock protein 40. Int J Biochem Cell Biol 36:1585–1598
Wang T, Bisson WH, Mäser P, Scapozza L, Picard D (2014) Differences in conformational dynamics between Plasmodium falciparum and human Hsp90 orthologues enable the structure-based discovery of pathogen-selective inhibitors. J Med Chem 57:2524–2535
Kam**a HH, Hageman J, Vos MJ, Kubota H, Tanguay RM, Bruford EA, Cheetham ME, Chen B, Hightower LE (2009) Guidelines for the nomenclature of the human heat shock proteins. Cell Stress Chaperones 14:105–111
Kelley WL, Georgopoulos C (1997) The T/t common exon of simian virus 40, JC, and BK polyomavirus T antigens can functionally replace the J-domain of the Escherichia coli DnaJ molecular chaperone. Proc Natl Acad Sci U S A 94:3679–3684
Yan W, Craig EA (1999) The glycine-phenylalanine-rich region determines the specificity of the yeast Hsp40 Sis1. Mol Cell Biol 19:7751–7758
Hennessy F, Boshoff A, Blatch GL (2005) Rational mutagenesis of a 40 kDa heat shock protein from Agrobacterium tumefaciens identifies amino acid residues critical to its in vivo function. Int J Biochem Cell Biol 37:177–191
Johnson JL, Craig EA (2000) A role for the Hsp40 Ydj1 in repression of basal steroid receptor activity in yeast. Mol Cell Biol 20:3027–3036
Boshoff A, Hennessy F, Blatch GL (2004) The in vivo and in vitro characterization of DnaK from Agrobacterium tumefaciens RUOR. Protein Expr Purif 38:161–169
Mbaba M, de la Mare JA, Sterrenberg JN, Kajewole D, Maharaj S, Edkins AL, Isaacs M, Hoppe HC, Khanye SD (2019) Novobiocin-ferrocene conjugates possessing anticancer and antiplasmodial activity independent of HSP90 inhibition. J Biol Inorg Chem 24:139–149
Acknowledgments
We gratefully acknowledge Dr. O Deloche (Geneva, Switzerland) for supplying the E. coli OD259 strain and plasmids pBAD22A and pWKG90. The S. cerevisiae JJ160 strain and PKG6 and pRS317-YDJ1 plasmids were a kind gift of Elizabeth Craig (University of Madison—Wisconsin). G.L.B. acknowledges the financial support of Higher Colleges of Technology, UAE (Interdisciplinary Research Grant, IRG; Grant No. 213471), and Rhodes University, South Africa (Rated Researcher Grant, RRG; project number IFRR100006). Research in the lab of A.L.E is supported by a Newton Advanced Fellowships from the Academy of Medical Sciences (UK) and grants from the Resilient Futures Challenge-Led Initiative from the Royal Society (UK) (Grant No. CHL\R1\180142), the South African Research Chairs Initiative of the Department of Science and Technology (DST) and the NRF (Grant No. 98566), Poliovirus Research Foundation (PRF, South Africa) (Grant No. 18/06), and Rhodes University and the Grand Challenges Africa Drug Discovery Programme (which is a partnership between the African Academy of Sciences [AAS], the Bill & Melinda Gates Foundation, Medicines for Malaria Venture [MMV], and the University of Cape Town Drug Discovery and Development Centre [H3D]) (Grant No. GCA/DD/rnd3/043).
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Edkins, A.L., Blatch, G.L. (2023). Complementation Assays for Co-chaperone Function. In: Calderwood, S.K., Prince, T.L. (eds) Chaperones. Methods in Molecular Biology, vol 2693. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-3342-7_9
Download citation
DOI: https://doi.org/10.1007/978-1-0716-3342-7_9
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-3341-0
Online ISBN: 978-1-0716-3342-7
eBook Packages: Springer Protocols