AbstractAbstract
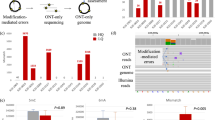
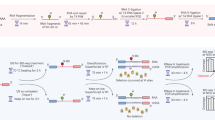
Map** DNA modifications at the base resolution is now possible at the genome level thanks to advances in sequencing technologies. Long-read sequencing data can be used to identify modified base patterns. However, the downstream analysis of Pacific Biosciences (PacBio) or Oxford Nanopore Technologies (ONT) data requires the integration of genomic annotation and comprehensive filtering to prevent the accumulation of artifact signals. We present in this chapter, a linear workflow to fully analyze modified base patterns using the DNA Modification Annotation (DNAModAnnot) package. This workflow includes a thorough filtering based on sequencing quality and false discovery rate estimation and provides tools for a global analysis of DNA modifications. Here, we provide an application example of this workflow with PacBio data and guide the user by explaining expected outputs via a fully integrated Rmarkdown script. This protocol is presented with tips showing how to adapt the provided code for annotating epigenomes of any organism according to the user needs.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Gouil Q, Keniry A (2019) Latest techniques to study DNA methylation. Essays Biochem 63:639–648. https://doi.org/10.1042/EBC20190027
Liu Y, Rosikiewicz W, Pan Z, Jillette N, Wang P, Taghbalout A, Foox J, Mason C, Carroll M, Cheng A, Li S (2021) DNA methylation-calling tools for Oxford Nanopore sequencing: a survey and human epigenome-wide evaluation. Genome Biol 22:295. https://doi.org/10.1186/s13059-021-02510-z
Methylome analysis technical note Pacificbiosciences/bioinformatics-training Wiki. In: GitHub. https://github.com/PacificBiosciences/Bioinformatics-Training/wiki/Methylome-Analysis-Technical-Note. Accessed 28 Jan 2022
Detecting DNA base modifications using single molecule, real-time sequencing. White Pap Base Modif. 2015. https://www.pacb.com/wp-content/uploads/2015/09/WP_Detecting_DNA_Base_Modifications_Using_SMRT_Sequencing.pdf. Accessed 27 Jan 2022
O’Brown ZK, Boulias K, Wang J, Wang SY, O’Brown NM, Hao Z, Shibuya H, Fady P-E, Shi Y, He C, Megason SG, Liu T, Greer EL (2019) Sources of artifact in measurements of 6mA and 4mC abundance in eukaryotic genomic DNA. BMC Genomics 20:445. https://doi.org/10.1186/s12864-019-5754-6
Hardy A, Matelot M, Touzeau A, Klopp C, Lopez-Roques C, Duharcourt S, Defrance M (2021) DNAModAnnot: a R toolbox for DNA modification filtering and annotation. Bioinformatics 37:2738–2740. https://doi.org/10.1093/bioinformatics/btab032
Lawrence M, Huber W, Pagès H, Aboyoun P, Carlson M, Gentleman R, Morgan MT, Carey VJ (2013) Software for computing and annotating genomic ranges. PLoS Comput Biol 9:e1003118. https://doi.org/10.1371/journal.pcbi.1003118
Pagès H, Aboyoun P, Gentleman R, DebRoy S (2022) Biostrings: efficient manipulation of biological strings. https://bioconductor.org/packages/Biostrings/. Accessed 26 Jan 2022
Wang Y, Chen X, Sheng Y, Liu Y, Gao S (2017) N6-adenine DNA methylation is associated with the linker DNA of H2A.Z-containing well-positioned nucleosomes in Pol II-transcribed genes in Tetrahymena. Nucleic Acids Res 45:11594–11606. https://doi.org/10.1093/nar/gkx883
**ong J, Lu X, Zhou Z, Chang Y, Yuan D, Tian M, Zhou Z, Wang L, Fu C, Orias E, Miao W (2012) Transcriptome analysis of the model protozoan, tetrahymena thermophila, using deep RNA sequencing. PLoS One 7:e30630. https://doi.org/10.1371/journal.pone.0030630
PacBio – software downloads. In: PacBio. https://www.pacb.com/support/software-downloads/. Accessed 26 Jan 2022
Ni P, Huang N, Zhang Z, Wang D-P, Liang F, Miao Y, **ao C-L, Luo F, Wang J (2019) DeepSignal: detecting DNA methylation state from Nanopore sequencing reads using deep-learning. Bioinformatics 35:4586–4595. https://doi.org/10.1093/bioinformatics/btz276
Morgan M, Ramos M (2021) BiocManager: access the bioconductor project package repository. https://CRAN.R-project.org/package=BiocManager. Accessed 26 Jan 2022
Wickham H, Hester J, Chang W, Bryan J (2021) devtools: tools to make develo** R packages easier. https://CRAN.R-project.org/package=devtools. Accessed 26 Jan 2022
Gurevich A, Saveliev V, Vyahhi N, Tesler G (2013) QUAST: quality assessment tool for genome assemblies. Bioinformatics 29:1072–1075. https://doi.org/10.1093/bioinformatics/btt086
Zhu S, Beaulaurier J, Deikus G, Wu TP, Strahl M, Hao Z, Luo G, Gregory JA, Chess A, He C, ** and characterizing N6-methyladenine in eukaryotic genomes using single-molecule real-time sequencing. Genome Res 28:1067–1078. https://doi.org/10.1101/gr.231068.117
Lawrence M, Gentleman R, Carey V (2009) rtracklayer: an R package for interfacing with genome browsers. Bioinformatics 25:1841–1842. https://doi.org/10.1093/bioinformatics/btp328
Hahne F, Ivanek R (2016) Visualizing genomic data using Gviz and bioconductor. In: Mathé E, Davis S (eds) Statistical genomics: methods and protocols. Springer, New York, pp 335–351. https://doi.org/10.1007/978-1-4939-3578-9_16
Ni P, Huang N, bioinformaticsCSU (2021) DeepSignal. https://github.com/bioinfomaticsCSU/deepsignal. Accessed 26 Jan 2022
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Hardy, A., Duharcourt, S., Defrance, M. (2023). DNA Modification Patterns Filtering and Analysis Using DNAModAnnot. In: Oliveira, P.H. (eds) Computational Epigenomics and Epitranscriptomics. Methods in Molecular Biology, vol 2624. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2962-8_7
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2962-8_7
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2961-1
Online ISBN: 978-1-0716-2962-8
eBook Packages: Springer Protocols