Abstract
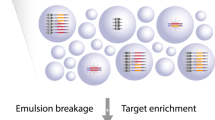
Complete comprehension of clinically relevant variation among human genomes is likely only to come from sequencing platforms that are cost-efficient, and which feature both accurate base calling and long-range DNA phasing capability. The NGS revolution has struggled to meet the latter of these needs. Here we describe a protocol to address this limitation by preserving the molecular origin of short sequencing reads with an insignificant increase to sequencing costs. Whole haplotype-resolved genomes with megabase-scale phase blocks can be obtained with this method; offering researchers a unique opportunity to tackle the hurdles of de novo sequencing without being limited by a lack of resources.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Chaisson MJP, Sanders AD, Zhao X et al (2019) Multi-platform discovery of haplotype-resolved structural variation in human genomes. Nat Commun 10(1784). https://doi.org/10.1038/s41467-018-08148-z
Church DM, Schneider VA, Steinberg KM et al (2015) Extending reference assembly models. Genome Biol 16:13
Schneider VA, Graves-Lindsey T, Howe K et al (2017) Evaluation of grch38 and de novo haploid genome assemblies demonstrates the enduring quality of the reference assembly. Genome Res 27(5):849–864. https://doi.org/10.1101/gr.213611.116
Clarke J, Wu HC, Jayasinghe L et al (2009) Continuous base identification for single-molecule nanopore DNA sequencing. Nat Nanotechnol 4:265–270
Eid J, Fehr A, Gray J et al (2009) Real-time DNA sequencing from single polymerase molecules. Science 323:133–138
Zheng GXY, Lau BT, Schnall-Levin M et al (2016) Haploty** germline and cancer genomes with high-throughput linked-read sequencing. Nat Biotechnol 34:303–311. https://doi.org/10.1038/nbt.3432
Amini S, Pushkarev D, Christiansen L et al (2014) Haplotype-resolved whole-genome sequencing by contiguity-preserving transposition and combinatorial indexing. Nat Genet 46:1343–1349. https://doi.org/10.1038/ng.3119
Peters BA, Kermani BG, Sparks AB et al (2012) Accurate whole-genome sequencing and haploty** from 10 to 20 human cells. Nature 487:190–195
Lan F, Haliburton JR, Yuan A, Abate AR (2016) Droplet barcoding for massively parallel single-molecule deep sequencing. Nat Commun 7(11784). https://doi.org/10.1038/ncomms11784
Redin D, Frick T, Aghelpasand H et al (2019) High throughput barcoding method for genome-scale phasing. Sci Rep 9(18116). https://doi.org/10.1038/s41598-019-54446-x
Redin D, Frick T, Aghelpasand H et al (2018) Efficient whole genome haploty** and high-throughput single molecule phasing with barcode-linked reads. bioRxiv:356121. https://doi.org/10.1101/356121
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Redin, D. (2023). Low-Cost Genome-Scale Phasing with Barcode-Linked Sequencing. In: Peters, B.A., Drmanac, R. (eds) Haploty**. Methods in Molecular Biology, vol 2590. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2819-5_6
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2819-5_6
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2818-8
Online ISBN: 978-1-0716-2819-5
eBook Packages: Springer Protocols