Abstract
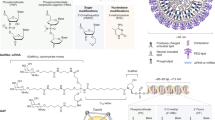
Nucleic acids are paving the way for advanced therapeutics. Unveiling the genome enabled a better understanding of unique genotype–phenotype profiling. Methods for engineering and analysis of nucleic acids, from polymerase chain reaction to Cre-Lox recombination, are contributing greatly to biomarkers’ discovery, map** of cellular signaling cascades, and smart design of therapeutics in precision medicine. Investigating the different subtypes of DNA and RNA via sequencing and profiling is empowering the scientific community with valuable information, to be used in advanced therapeutics, tracking epigenetics linked to disease. Recent results from the application of nucleic acids in novel therapeutics and precision medicine are very encouraging, demonstrating great potential to treat cancer, viral infections via inoculation (e.g., SAR-COV-2 mRNA vaccines), along with metabolic and genetic disorders. Limitations posed by challenges in delivery mode are being addressed to enable efficient guided-gene-programmed precision therapies. With the focus on genetic engineering and novel therapeutics, more precisely, in precision medicine, this chapter discusses the advance enabled by knowledge derived from these innovative branches of biotechnology.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Imelfort M, Batley J, Grimmond S, Edwards D (2009) Genome sequencing approaches and successes. Methods Mol Biol 513:345–358. https://doi.org/10.1007/978-1-59745-427-8_18
Zheng Z, Zhang N, Huang Z et al (2022) Genome survey sequencing and characterization of simple sequence repeat (SSR) markers in Platostoma palustre (Blume) A.J.Paton (Chinese mesona). Sci Rep 121(12):1–8. https://doi.org/10.1038/s41598-021-04264-x
Shamsi S, Sibraa L, Zhu X, Barton DP (2022) Characterisation of Temnocephalidae flatworms in common Australian freshwater prawn, Macrobrachium australiense. Sci Rep 121(12):1–8. https://doi.org/10.1038/s41598-022-05123-z
Rens W, Fu B, O’Brien PCM, Ferguson-Smith M (2006) Cross-species chromosome painting. Nat Protoc 1:783–790. https://doi.org/10.1038/NPROT.2006.91
Gates AJ, Gysi DM, Kellis M, Barabási AL (2021) A wealth of discovery built on the Human Genome Project — by the numbers. Nature 5907845(590):212–215. https://doi.org/10.1038/d41586-021-00314-6
Sun BB, Kurki MI, Foley CN et al (2022) Genetic associations of protein-coding variants in human disease. Nature 2022:1–8. https://doi.org/10.1038/s41586-022-04394-w
(2021) The next 20 years of human genomics must be more equitable and more open. Nature 590:183–184. https://doi.org/10.1038/D41586-021-00328-0
Dhingra DM, Ooi AT, Ruff DW (2022) Simultaneous DNA, RNA, and protein analysis from single cells using a high-throughput microfluidic workflow for resolution of genotype-to-phenotype modalities. Methods Mol Biol 2386:289–307. https://doi.org/10.1007/978-1-0716-1771-7_17
Pereira GC (2017) Genomics and artificial intelligence working together in drug discovery and repositioning: the advent of adaptive pharmacogenomics in glioblastoma and chronic arterial inflammation therapies. In: Biotechnology and production of anti-cancer compounds. Springer International Publishing, Cham, pp 253–281
Fagan D, O’Neill M, Galván-López E et al (2010) An analysis of genotype-phenotype maps in grammatical evolution. Lect Notes Comput Sci (including Subser Lect Notes Artif Intell Lect Notes Bioinformatics) 6021 LNCS:62–73. https://doi.org/10.1007/978-3-642-12148-7_6
Wang G, Li X, Gao X et al (2021) Analysis of genotype–phenotype relationships in 90 Chinese probands with Waardenburg syndrome. Hum Genet 1–14. https://doi.org/10.1007/S00439-021-02301-3/TABLES/4
Boissinot M, Ho A-H, Bergeron MG et al (2001) Detection of nucleic acids using novel polymers able to transduce hybridization into optical or electrical signal. Micro Total Anal Syst 2001:319–320. https://doi.org/10.1007/978-94-010-1015-3_135
Baoutina A, Bhat S (2021) Novel design of nucleic acid standards for hydrolysis probe-based PCR with melting analysis. Gene Ther 2021:1–6. https://doi.org/10.1038/s41434-021-00288-0
Lichtenstein AV, Melkonyan HS, Tomei LD, Umansky SR (2006) Novel applications of polymerase chain reaction to urinary nucleic acid analysis. Methods Mol Biol 336:145–154. https://doi.org/10.1385/1-59745-074-X:145
(2007) Nucleic acids hybridization: modern applications. Nucleic Acids Hybrid Mod Appl 1–314. https://doi.org/10.1007/978-1-4020-6040-3
Joshi VG, Chindera K, Bais MV et al (2021) Novel peptide (RATH) mediated delivery of peptide nucleic acids for antiviral interventions. Appl Microbiol Biotechnol 105:6669–6677. https://doi.org/10.1007/S00253-021-11502-9/FIGURES/4
Kulkarni JA, Witzigmann D, Thomson SB et al (2021) The current landscape of nucleic acid therapeutics. Nat Nanotechnol 166(16):630–643. https://doi.org/10.1038/s41565-021-00898-0
Yamada Y (2021) Nucleic acid drugs—current status, issues, and expectations for exosomes. Cancers (Basel) 13. https://doi.org/10.3390/CANCERS13195002
Balke D, Hieronymus R, M€ S et al (2017) Challenges and perspectives in nucleic acid enzyme engineering. Adv Biochem Eng Biotechnol 170:21–35. https://doi.org/10.1007/10_2017_21
Pêgo AP, Oliveira H, Moreno PM (2013) Biomaterial-based vectors for targeted delivery of nucleic acids to the nervous system. 185–224. https://doi.org/10.1007/978-94-007-6010-3_7
**ao W, Zhang C, Yadava P, Hughes J (2004) Nucleic acid cellular delivery. Cell Drug Deliv 81–94. https://doi.org/10.1007/978-1-59259-745-1_6
Wadhwa A, Aljabbari A, Lokras A et al (2020) Opportunities and challenges in the delivery of mRNA-based vaccines. Pharmaceutics 12. https://doi.org/10.3390/PHARMACEUTICS12020102
Elsabahy M, Nazarali A, Foldvari M (2011) Non-viral nucleic acid delivery: key challenges and future directions. Curr Drug Deliv 8:235–244. https://doi.org/10.2174/156720111795256174
Silva A, Lopes C, Sousa Lobo J, Amaral M (2015) Nucleic acids delivery systems: a challenge for pharmaceutical technologists. Curr Drug Metab 16:3–16. https://doi.org/10.2174/1389200216666150401110211
Thatcher SA (2018) Nucleic acid isolation. Princ Appl Mol Diagn 35–46. https://doi.org/10.1016/B978-0-12-816061-9.00003-5
Schwarzenbach H, Hoon DSB, Pantel K (2011) Cell-free nucleic acids as biomarkers in cancer patients. Nat Rev Cancer 116(11):426–437. https://doi.org/10.1038/nrc3066
Mullegama SV, Alberti MO, Au C et al (2019) Nucleic acid extraction from human biological samples. Methods Mol Biol 1897:359–383. https://doi.org/10.1007/978-1-4939-8935-5_30
Robinson NP (2013) Analysis of branched DNA replication and recombination intermediates from prokaryotic cells by two-dimensional (2D) native-native agarose gel electrophoresis. Methods Mol Biol 1054:45–61. https://doi.org/10.1007/978-1-62703-565-1_3
Sakatani Y, Ichihashi N, Kazuta Y, Yomo T (2015) A transcription and translation-coupled DNA replication system using rolling-circle replication. Sci Rep 51(5):1–9. https://doi.org/10.1038/srep10404
Saitou N (2018) Replication, transcription, and translation. 3–21. https://doi.org/10.1007/978-3-319-92642-1_1
Dales S, Pogo BGT (1981) Transcription and translation. 45–53. https://doi.org/10.1007/978-3-7091-8625-1_5
Translation: DNA to mRNA to Protein | Learn Science at Scitable. https://www.nature.com/scitable/topicpage/translation-dna-to-mrna-to-protein-393/. Accessed 24 Feb 2022
**a X (2021) Domains and functions of spike protein in SARS-Cov-2 in the context of vaccine design. Viruses 13. https://doi.org/10.3390/V13010109
Jackson LA, Anderson EJ, Rouphael NG et al (2020) An mRNA vaccine against SARS-CoV-2 - preliminary report. N Engl J Med 383:1920–1931. https://doi.org/10.1056/NEJMOA2022483
Baden LR, El Sahly HM, Essink B et al (2021) Efficacy and safety of the mRNA-1273 SARS-CoV-2 vaccine. N Engl J Med 384:403–416. https://doi.org/10.1056/NEJMOA2035389/SUPPL_FILE/NEJMOA2035389_DATA-SHARING.PDF
Cai X, Li JJ, Liu T et al (2021) Infectious disease mRNA vaccines and a review on epitope prediction for vaccine design. Brief Funct Genomics 20:289–303. https://doi.org/10.1093/BFGP/ELAB027
Wang Y, Nguyen K, Spitale RC, Chaput JC (2021) A biologically stable DNAzyme that efficiently silences gene expression in cells. Nat Chem 134(13):319–326. https://doi.org/10.1038/s41557-021-00645-x
Huang J, Chen M, Whitley MJ et al (2017) Generation and comparison of CRISPR-Cas9 and Cre-mediated genetically engineered mouse models of sarcoma. Nat Commun 81(8):1–11. https://doi.org/10.1038/ncomms15999
Noiman T, Kahana C (2018) A simple combined use of CRISPR-Cas9 and Cre-LoxP Technologies for generating conditional gene knockouts in mammalian cells. Cris J 1:278–285. https://doi.org/10.1089/CRISPR.2018.0010
Pereira GC, Malik S, Kis Z, Rocamonde B (2019) Computationally designed recombinant-DNA-based compounds production driven in plants during secondary metabolism and their implication in antimalarial therapies. In: Natural bio-active compounds. Springer Singapore, Singapore, pp 127–146
Pereira GC (2019) Application of biotechnology in producing plant bio-active compounds. In: Natural bio-active compounds. Springer, Singapore, pp 59–78
Goto H, Osaki T, Kijima T et al (2001) Gene therapy utilizing the Cre/loxP system selectively suppresses tumor growth of disseminated carcinoembryonic antigen-producing cancer cells. Int J Cancer 94:414–419
Methods and mechanisms for genetic manipulation of plants, animals, and microorganisms - safety of genetically engineered foods - NCBI Bookshelf. https://www.ncbi.nlm.nih.gov/books/NBK215771/. Accessed 24 Feb 2022
Wang Y, Wang Y, Song D et al (2021) An RNA-cleaving threose nucleic acid enzyme capable of single point mutation discrimination. Nat Chem 2021:1–10. https://doi.org/10.1038/s41557-021-00847-3
Ageely EA, Chilamkurthy R, Jana S et al (2021) Gene editing with CRISPR-Cas12a guides possessing ribose-modified pseudoknot handles. Nat Commun 121(12):1–15. https://doi.org/10.1038/s41467-021-26989-z
Dykstra PB, Kaplan M, Smolke CD (2022) Engineering synthetic RNA devices for cell control. Nat Rev Genet 2022:1–14. https://doi.org/10.1038/s41576-021-00436-7
Manzari MT, Shamay Y, Kiguchi H et al (2021) Targeted drug delivery strategies for precision medicines. Nat Rev Mater 64(6):351–370. https://doi.org/10.1038/s41578-020-00269-6
Vincent-Salomon A (2015) The key role of pathology in the context of precision medicine trials. Pan-cancer Integr Mol Portrait Towar a New Paradig Precis Med 9–14. https://doi.org/10.1007/978-3-319-22189-2_2
Rahat B, Ali T, Sapehia D et al (2020) Circulating cell-free nucleic acids as epigenetic biomarkers in precision medicine. Front Genet 11. https://doi.org/10.3389/FGENE.2020.00844
Bowman JC, Williams LD (2011) Nucleic acids. Encycl Astrobiol 1141–1147. https://doi.org/10.1007/978-3-642-11274-4_1079
Yamakawa K, Nakano-Narusawa Y, Hashimoto N et al (2019) Development and clinical trials of nucleic acid medicines for pancreatic cancer treatment. Int J Mol Sci 20. https://doi.org/10.3390/IJMS20174224
**g X, Zhang F, Pan M et al (2019) Solidifying framework nucleic acids with silica. Nat Protoc 148(14):2416–2436. https://doi.org/10.1038/s41596-019-0184-0
Mirkin CA, Stegh AH (2014) Spherical nucleic acids for precision medicine. Oncotarget 5:9–10. https://doi.org/10.18632/ONCOTARGET.1757
Sgro A, Blancafort P (2020) Epigenome engineering: new technologies for precision medicine. Nucleic Acids Res 48:12453–12482. https://doi.org/10.1093/NAR/GKAA1000
Saminathan A, Zajac M, Anees P, Krishnan Y (2021) Organelle-level precision with next-generation targeting technologies. Nat Rev Mater 2021:1–17. https://doi.org/10.1038/s41578-021-00396-8
Kibirige CN, Manak M, King D et al (2022) Development of a sensitive, quantitative assay with broad subtype specificity for detection of total HIV-1 nucleic acids in plasma and PBMC. Sci Rep 121(12):1–16. https://doi.org/10.1038/s41598-021-03016-1
Regnery RL, Johnson KM, Kiley MP (1980) Virion nucleic acid of Ebola virus. J Virol 36:465–469. https://doi.org/10.1128/JVI.36.2.465-469.1980
Wang L, Wang X, Wu Y et al (2022) Rapid and ultrasensitive electromechanical detection of ions, biomolecules and SARS-CoV-2 RNA in unamplified samples. Nat Biomed Eng 2022:1–10. https://doi.org/10.1038/s41551-021-00833-7
Zheng Y, Deng J, Han L et al (2022) SARS-CoV-2 NSP5 and N protein counteract the RIG-I signaling pathway by suppressing the formation of stress granules. Signal Transduct Target Ther 71(7):1–12. https://doi.org/10.1038/s41392-022-00878-3
Katti A, Diaz BJ, Caragine CM et al (2022) CRISPR in cancer biology and therapy. Nat Rev Cancer 2022:1–21. https://doi.org/10.1038/s41568-022-00441-w
Baharlooi H, Mansourabadi AH, Minbashi Moeini M et al (2021) Nucleic acids as novel therapeutic modalities to address multiple sclerosis onset and progression. Cell Mol Neurobiol 2021:1–17. https://doi.org/10.1007/S10571-021-01158-4
Lee H, Park C, Na W et al (2020) Precision cell-free DNA extraction for liquid biopsy by integrated microfluidics. npj Precis Oncol 41(4):1–10. https://doi.org/10.1038/s41698-019-0107-0
Jani MS, Veetil AT, Krishnan Y (2019) Precision immunomodulation with synthetic nucleic acid technologies. Nat Rev Mater 46(4):451–458. https://doi.org/10.1038/s41578-019-0105-4
Puray-Chavez M, Tedbury PR, Huber AD et al (2017) Multiplex single-cell visualization of nucleic acids and protein during HIV infection. Nat Commun 81(8):1–11. https://doi.org/10.1038/s41467-017-01693-z
Jacob ST, Crozier I, Fischer WA et al (2020) Ebola virus disease. Nat Rev Dis Prim 61(6):1–31. https://doi.org/10.1038/s41572-020-0147-3
Long QX, Liu BZ, Deng HJ et al (2020) Antibody responses to SARS-CoV-2 in patients with COVID-19. Nat Med 266(26):845–848. https://doi.org/10.1038/s41591-020-0897-1
Vogel AB, Kanevsky I, Che Y et al (2021) BNT162b vaccines protect rhesus macaques from SARS-CoV-2. Nature 592:283–289. https://doi.org/10.1038/S41586-021-03275-Y
Wu Y, Huang X, Yuan L et al (2021) A recombinant spike protein subunit vaccine confers protective immunity against SARS-CoV-2 infection and transmission in hamsters. Sci Transl Med eabg1143. https://doi.org/10.1126/SCITRANSLMED.ABG1143
WHO Coronavirus (COVID-19) Dashboard | WHO Coronavirus (COVID-19) Dashboard With Vaccination Data. https://covid19.who.int/. Accessed 24 Feb 2022
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Pereira, G.C. (2023). Latest Trends in Nucleic Acids’ Engineering Techniques Applied to Precision Medicine. In: Pereira, G.C. (eds) Gene, Drug, and Tissue Engineering. Methods in Molecular Biology, vol 2575. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2716-7_2
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2716-7_2
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2715-0
Online ISBN: 978-1-0716-2716-7
eBook Packages: Springer Protocols