Abstract
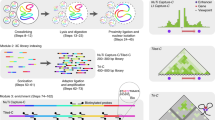
Targeted chromosome conformation capture (HiCap) is an experimental method for detecting spatial interactions of genomic features such as promoters and/or enhancers. The protocol first describes the design of sequence capture probes. After that, it provides details on the chromosome conformation capture adapted for next-generation sequencing (Hi-C). Finally, the methodology for coupling Hi-C with sequence capture technology is described.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Dekker J, Rippe K, Dekker M et al (2002) Capturing chromosome conformation. Science 295(5558):1306–1311
Lieberman-Aiden E, van Berkum NL, Williams L et al (2009) Comprehensive map** of long-range interactions reveals folding principles of the human genome. Science 326(5950):289–293
Schmitt AD, Hu M, Ren B (2016) Genome-wide map** and analysis of chromosome architecture. Nat Rev Mol Cell Biol 17(12):743–755
Hughes JR, Roberts N, McGowan S et al (2014) Analysis of hundreds of cis-regulatory landscapes at high resolution in a single, high-throughput experiment. Nat Genet 46(2):205–212
Sahlen P, Abdullayev I, Ramskold D et al (2015) Genome-wide map** of promoter-anchored interactions with close to single-enhancer resolution. Genome Biol 16:156
Dryden NH, Broome LR, Dudbridge F et al (2014) Unbiased analysis of potential targets of breast cancer susceptibility loci by capture Hi-C. Genome Res 24(11):1854–1868
Zhao Z, Tavoosidana G, Sjolinder M et al (2006) Circular chromosome conformation capture (4C) uncovers extensive networks of epigenetically regulated intra- and interchromosomal interactions. Nat Genet 38(11):1341–1347
Simonis M, Klous P, Splinter E et al (2006) Nuclear organization of active and inactive chromatin domains uncovered by chromosome conformation capture-on-chip (4C). Nat Genet 38(11):1348–1354
Fullwood MJ, Liu MH, Pan YF et al (2009) An oestrogen-receptor-alpha-bound human chromatin interactome. Nature 462(7269):58–64
Javierre BM, Burren OS, Wilder SP et al (2016) Lineage-specific genome architecture links enhancers and non-coding disease variants to target gene promoters. Cell 167(5):1369–84 e19
Sahlen P, Spalinskas R, Asad S et al (2021) Chromatin interactions in differentiating keratinocytes reveal novel atopic dermatitis and psoriasis-associated genes. J Allergy Clin Immunol 147(5):1742–1752
Pradhananga S, Spalinskas R, Poujade FA et al (2020) Promoter anchored interaction landscape of THP-1 macrophages captures early immune response processes. Cell Immunol 355:104148
Akerborg O, Spalinskas R, Pradhananga S et al (2019) High-resolution regulatory maps connect vascular risk variants to disease-related pathways. Circ Genom Precis Med 12(3):e002353
Beesley J, Sivakumaran H, Moradi Marjaneh M et al (2020) Chromatin interactome map** at 139 independent breast cancer risk signals. Genome Biol 21(1):8
Anil A, Spalinskas R, Akerborg O et al (2018) HiCapTools: a software suite for probe design and proximity detection for targeted chromosome conformation capture applications. Bioinformatics 34(4):675–677
Cairns J, Freire-Pritchett P, Wingett SW et al (2016) CHiCAGO: robust detection of DNA loo** interactions in capture Hi-C data. Genome Biol 17(1):127
Yang Y (2011) Thermodynamics of DNA binding and break repair by the Pol I DNA polymerases from Escherichia coli and Thermus aquaticus
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Zhigulev, A., Sahlén, P. (2022). Targeted Chromosome Conformation Capture (HiCap). In: Sexton, T. (eds) Spatial Genome Organization. Methods in Molecular Biology, vol 2532. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2497-5_5
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2497-5_5
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2496-8
Online ISBN: 978-1-0716-2497-5
eBook Packages: Springer Protocols