Abstract
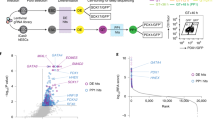
The CRISPR/Cas technology has revolutionized forward genetic screening, and thereby facilitated genetic dissection of cellular processes and pathways. TGF-β signaling is a highly conserved cascade involved in development, regeneration, and diseases such as cancer. Even though many core components of the signaling cascade have already been described, several context-dependent pathway modulators remain unknown. To address this knowledge gap, we have recently developed a CRISPR screening approach for identifying TGF-β pathway regulators in three-dimensional organoid culture systems. Here, we provide a detailed protocol describing this approach in human intestinal organoids. With adaptations, this screening method could also be applied to other organoid types, and to other signaling cascades such as EGF or WNT signaling, thereby uncovering important mechanism in regeneration and disease.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
David CJ, Massagué J (2018) Contextual determinants of TGFβ action in development, immunity and cancer. Nat Rev Mol Cell Biol 19:419–435
Massagué J (2008) TGFβ in cancer. Cell 134:215–230
Akhurst RJ, Derynck R (2001) TGF-β signaling in cancer - a double-edged sword. Trends Cell Biol 11:S44–S51
Pickup M, Novitskiy S, Moses HL (2013) The roles of TGFβ in the tumour microenvironment. Nat Rev Cancer 13:788–799
Drost J, Van Jaarsveld RH, Ponsioen B et al (2015) Sequential cancer mutations in cultured human intestinal stem cells. Nature 521:43–47. https://doi.org/10.1038/nature14415
Muzny DM, Bainbridge MN, Chang K et al (2012) Comprehensive molecular characterization of human colon and rectal cancer. Nature 487:330–337. https://doi.org/10.1038/nature11252
Guinney J, Dienstmann R, Wang X et al (2015) The consensus molecular subtypes of colorectal cancer. Nat Med 21:1350–1356. https://doi.org/10.1038/nm.3967
Sato T, Vries RG, Snippert HJ et al (2009) Single Lgr5 stem cells build crypt-villus structures in vitro without a mesenchymal niche. Nature 459:262–265
Ringel T, Frey N, Ringnalda F et al (2020) Genome-scale CRISPR screening in human intestinal organoids identifies drivers of TGF-β resistance. Cell Stem Cell 26:431–440.e8. https://doi.org/10.1016/J.STEM.2020.02.007
Hart T, Chandrashekhar M, Aregger M et al (2015) High-resolution CRISPR screens reveal fitness genes and genotype-specific cancer liabilities. Cell 163:1515–1526. https://doi.org/10.1016/j.cell.2015.11.015
Shifrut E, Carnevale J, Tobin V et al (2018) Genome-wide CRISPR screens in primary human T cells reveal key regulators of immune function. Cell 175:1958–1971.e15. https://doi.org/10.1016/j.cell.2018.10.024
Shalem O, Sanjana NE, Hartenian E et al (2014) Genome-scale CRISPR-Cas9 knockout screening in human cells. Science 343:84–87. https://doi.org/10.1126/science.1247005
Wang T, Wei JJ, Sabatini DM, Lander ES (2014) Genetic screens in human cells using the CRISPR-Cas9 system. Science 343:80–84. https://doi.org/10.1126/science.1246981
Sato T, Stange DE, Ferrante M et al (2011) Long-term expansion of epithelial organoids from human colon, adenoma, adenocarcinoma, and Barrett’s epithelium. Gastroenterology 141:1762–1772. https://doi.org/10.1053/j.gastro.2011.07.050
Doench JG, Fusi N, Sullender M et al (2016) Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR-Cas9. Nat Biotechnol 34:184–191. https://doi.org/10.1038/nbt.3437
Mizutani T, Clevers H (2020) Primary intestinal epithelial organoid culture. Methods Mol Biol 171:185–200
Protocol: PCR of sgRNAs for Illumina sequencing MATERIALS
Wang B, Wang M, Zhang W et al (2019) Integrative analysis of pooled CRISPR genetic screens using MAGeCKFlute. Nat Protoc 14:756–780. https://doi.org/10.1038/s41596-018-0113-7
Li W, Xu H, **ao T et al (2014) MAGeCK enables robust identification of essential genes from genome-scale CRISPR/Cas9 knockout screens. Genome Biol 15:554. https://doi.org/10.1186/s13059-014-0554-4
Ran FA, Hsu PD, Wright J et al (2013) Genome engineering using the CRISPR-Cas9 system. Nat Protoc 8:2281–2308
Sanjana NE, Shalem O, Zhang F (2014) Improved vectors and genome-wide libraries for CRISPR screening. Nat Methods 11:783–784
Joung J, Konermann S, Gootenberg JS et al (2017) Genome-scale CRISPR-Cas9 knockout and transcriptional activation screening. Nat Protoc 12:828–863. https://doi.org/10.1038/nprot.2017.016
Esk C, Lindenhofer D, Haendeler S et al (2020) A human tissue screen identifies a regulator of ER secretion as a brain-size determinant. Science 370:935–941. https://doi.org/10.1126/science.abb5390
Acknowledgements
We thank Till Ringel for development of the method and valuable discussion on the protocol. Also, we thank Patrik Simmler for proof-reading the manuscript. Nina Frey is supported by an ETH PhD grant.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Frey, N., Schwank, G. (2022). CRISPR-Based Screening in Three-Dimensional Organoid Cultures to Identify TGF-β Pathway Regulators. In: Zi, Z., Liu, X. (eds) TGF-Beta Signaling. Methods in Molecular Biology, vol 2488. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2277-3_8
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2277-3_8
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2276-6
Online ISBN: 978-1-0716-2277-3
eBook Packages: Springer Protocols