Abstract
Background
Targeting the HGF/MET signaling pathway has been a viable therapeutic strategy for various cancer types due to hyperactivation of HGF/MET axis occurs frequently that leads to detrimental cancer progression and recurrence. Deciphering novel molecule mechanisms underlying complex HGF/MET signaling network is therefore critical to development of effective therapeutics for treating MET-dependent malignancies.
Results
Using isobaric mass tag-based quantitative proteomics approach, we identified IFITM3, an interferon-induced transmembrane protein that was highly expressed in micro-dissected gastric cancer (GC) tumor regions relative to adjacent non-tumor epithelia. Analyses of GC clinical specimens revealed that expression IFITM3 was closely correlated to advanced pathological stages. IFITM3 has been reported as a PIP3 scaffold protein that promotes PI3K signaling. In present study, we unprecedentedly unraveled that IFITM3 associated with MET and AKT to facilitate HGF/MET mediated AKT signaling crosstalk in suppressing FOXO3, consequently leading to c-MYC mediated GC progression. In addition, gene ontology analyses of the clinical GC cohort revealed significant correlation between IFITM3-associated genes and targets of c-MYC, which is a crucial downstream effector of HGF/MET pathway in cancer progression. Moreover, we demonstrated ectopic expression of IFITM3 suppressed FOXO3 expression, consequently led to c-MYC induction to promote tumor growth, cell metastasis, cancer stemness as well as chemoresistance. Conversely, depletion of IFITM3 resulted in suppression of HGF triggered cellular growth and migration via inhibition of AKT/c-MYC signaling in GC.
Conclusions
In summary, our present study unveiled a novel regulatory mechanism for c-MYC-driven oncogenesis underlined by IFITM3-mediated signaling crosstalk between MET associated AKT signaling cascade.
Similar content being viewed by others
Background
Gastric cancer (GC) ranks the fifth most common cancer and the third most common cause of cancer-related mortality worldwide [1]. Although early stage GC patients feasible for curative surgery retain 70 to 95% 5-year overall survival (OS) rate, more than two thirds of GC patients are diagnosed with advanced stages with unresectable diseases [2, 3]. For unresectable advanced or metastatic GC, the prognosis remains poor with median OS around 8 to 12 months after 1st-line chemotherapy and a 5-year OS at around 5% [4]. Since chemotherapy has been the major treatment option for advanced GC, development of chemoresistance has largely limited the effectiveness of chemotherapy and resulted in disease recurrence and grave prognosis [2]. The mechanisms of chemoresistance in GC are multifactorial that involve in drug efflux, drug interaction and dysregulation of cellular signaling as well as pathways that regulate cancer stemness [2, 5]. Hence, there has been an urgent need to explore new therapeutic targets to overcome chemoresistance in GC.
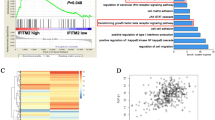
MET (Mesenchymal-epithelial transition factor) is a receptor tyrosine kinase that is proposed as a promising target in cancer therapy due to the predominant oncogenic signaling cascades that has been shown crucial to malignant progression and tumorigenesis of several cancer types [6,7,39,40,41], were identified to be significantly upregulated in GC tissues as compared to adjacent normal gastric tissues (Additional file 1: Fig. S1B). In addition to MCM2 to MCM7, six downstream oncogenic targets of c-MYC implicated in cancer survival and metabolism including PSMA1, PSMA6, PSMB2, HDAC2, LDHA and SPRING [45] were positively correlated with IFITM3 in 71 GC specimens using the Cho dataset [27] (Fig. 7B, and Additional file 1: Fig. S1C). Consistently, the gene expressions of MCM2, MEM3, MCM4, PSMA1, PSMA6, PSMB2, LDHA and SPRING were significantly elevated in IFITM3-overexpressing TMK-1 cells, while reduced expressions were observed in IFITM3-silenced TSGH cells (Fig. 7C and Additional file 1: Fig. S1D), further supporting IFITM3 as a positive regulator in MET/AKT/c-MYC signal axis in GC. To elaborate c-MYC induction is required during IFITM3-mediated tumor progression and chemoresistance acquisition, a c-MYC-silenced model was established using IFITM3-overexpressing GC cells (Fig. 7D). Our data demonstrated that both cellular proliferation and migration abilities of IFITM3 high-expressing cells were significantly compromised by the knockdown of c-MYC (Figs. 6F, 7E). Further, the diminished sphere formation by knockdown of c-MYC in IFITM3-overexpressing cells suggested c-MYC as a crucial target of IFITM3 (Fig. 7G). In line with these observations, c-MYC knockdown significantly lowered the 5’FU and cisplatin chemoresistance attributed by overexpression of IFITM3, confirming the crucial function of c-MYC in mediating the oncogenic effects of IFITM3 (Fig. 7H). Collectively, our present study revealed a novel mechanism modulated by IFITM3-associated HGF/MET/AKT signaling complexes that suppressed FOXO3, consequently leading to c-MYC upregulation to promote cell proliferation, metastasis, stemness and chemoresistance in GC (Fig. 8).
c-MYC and its target genes are regulated by IFITM3 in GC oncogenesis and chemoresistance. A Gene Set Enrichment Analysis (GSEA) was carried out to analyze the co-expression network and determine Pearson’s correlation coefficient between IFITM3 and other hallmarks of cancer in cBioPortal. The right panel represents an enrichment plot of IFITM3-correlated genes in c-MYC-regulated targets. B Spearman rank correlation coefficient of individual genes (including PSMA1, PSMA6, PSMB2, HDAC2, LDHA and SPRING) against IFITM3 in 71 GC tissues from Cho Gastric dataset (GSE138861). C Quantitative RT-PCR analyses were conducted on the six c-MYC target genes for their gene expression in TMK-1 IFITM3-overexpression or TSGH IFITM3-depletion models (*p < 0.05, **p < 0.01). D Western blot analyses of IFITM3 and c-MYC expression in control or IFITM3-overexpressing TMK-1 cells with or without c-MYC knockdown. Cellular growth curves (E) and migration rates (F) of the same experimental groups were determined by measuring viable and migrated cell numbers, respectively. G Sphere formation assays were performed to assess the influences c-MYC knockdown on IFITM3-overexpressing TMK-1 cells. (Left panel, scale bar, 500 μm; right panel is the statistical results, **p < 0.01) H Chemosensitivities of the same indicated groups of TMK-1 cells were determined based on the relative number of cells survived after treatment with 5’FU (20 μg/ml) or cisplatin (3 μM) for 72 h. *p < 0.05, **p < 0.01.
Cartoon illustration that depicts a model of IFITM3-mediated MET/ AKT/FOXO3/c-MYC cascade signaling pathway. Data presented in this study reveals a new signaling model in which IFITM3 associates with MET and AKT complex, leading to enhanced HGF/MET signaling as well as suppression of FOXO3, consequently resulting in upregulation of c-MYC to promote proliferation, metastasis, cancer stemness and chemoresistance of GC cells
Statistical analysis
All the in vitro experiments were reproducible and repeated at least three times, and the results are presented as means ± SEM. The Statistical analysis was performed with the GraphPad Prism software (GraphPad Software, CA) using Mann–Whitney U or Fisher’s exact test for between-group comparisons. The two-tailed paired or unpaired t-tests were performed to determine the significance between the groups compared. P values < 0.05 were considered statistically significant.
Availability of data and materials
The experimental procedure and results of iTRAQ quantitative proteomic analysis have been organized and deposited to Figshare (10. 6084/m9.figshare.20069969). Data could be available upon request to interested researchers. Please send data requests to Hsiang-Cheng Chi, PhD. Graduate Institute of Integrated Medicine, China Medical University, Taichung, Taiwan.
Abbreviations
- GC:
-
Gastric cancer
- OS:
-
Overall survival
- MET:
-
Mesenchymal-epithelial transition factor
- HGF:
-
Hepatocyte growth factor
- CSCs:
-
Cancer stem cells
- IFITM3:
-
Interferon induced transmembrane protein 3
- shRNA:
-
Hairpin RNA
- HCC:
-
Hepatocellular carcinoma
- FOXO3:
-
Forkhead Box O3
- c-MYC:
-
MYC proto-oncogene
- ALK:
-
Anaplastic lymphoma kinase
References
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68(6):394–424.
Shi W-J, Gao J-B. Molecular mechanisms of chemoresistance in gastric cancer. World J Gastrointest Oncol. 2016;8(9):673–81.
Yang MD, Lin KC, Lu MC, Jeng LB, Hsiao CL, Yueh TC, Fu CK, Li HT, Yen ST, Lin CW, et al. Contribution of matrix metalloproteinases-1 genotypes to gastric cancer susceptibility in Taiwan. Biomedicine (Taipei). 2017;7(2):10.
Le DT, Ott PA, Korytowsky B, Le H, Le TK, Zhang Y, Maglinte GA, Abraham P, Patel D, Shangguan T, et al. Real-world treatment patterns and clinical outcomes across lines of therapy in patients with advanced/metastatic gastric or gastroesophageal junction cancer. Clin Colorectal Cancer. 2020;19(1):32-38.e33.
Bekaii-Saab T, El-Rayes B. Identifying and targeting cancer stem cells in the treatment of gastric cancer. Cancer. 2017;123(8):1303–12.
Sylvester PW. Targeting met mediated epithelial-mesenchymal transition in the treatment of breast cancer. Clin Transl Med. 2014;3(1):30–30.
Khater AR, Abou-Antoun T. Mesenchymal epithelial transition factor signaling in pediatric nervous system tumors: implications for malignancy and cancer stem cell enrichment. Front Cell Dev Biol. 2021. https://doi.org/10.3389/fcell.2021.654103.
Zhang Y, **a M, ** K, Wang S, Wei H, Fan C, Wu Y, Li X, Li X, Li G, et al. Function of the c-Met receptor tyrosine kinase in carcinogenesis and associated therapeutic opportunities. Mol Cancer. 2018;17(1):45.
Carneiro F, Sobrinho-Simoes M. The prognostic significance of amplification and overexpression of c-met and c-erb B-2 in human gastric carcinomas. Cancer. 2000;88(1):238–40.
Lee J, Seo JW, Jun HJ, Ki CS, Park SH, Park YS, Lim HY, Choi MG, Bae JM, Sohn TS, et al. Impact of MET amplification on gastric cancer: possible roles as a novel prognostic marker and a potential therapeutic target. Oncol Rep. 2011;25(6):1517–24.
Li C, Wu JJ, Hynes M, Dosch J, Sarkar B, Welling TH. Pasca di Magliano M, Simeone DM: c-Met is a marker of pancreatic cancer stem cells and therapeutic target. Gastroenterology. 2011;141(6):2218-2227 e2215.
Yashiro M, Nishii T, Hasegawa T, Matsuzaki T, Morisaki T, Fukuoka T, Hirakawa K. A c-Met inhibitor increases the chemosensitivity of cancer stem cells to the irinotecan in gastric carcinoma. Br J Cancer. 2013;109(10):2619–28.
O’Brien CA, Kreso A, Jamieson CHM. Cancer stem cells and self-renewal. Clin Cancer Res. 2010;16(12):3113–20.
Smith A. A glossary for stem-cell biology. Nature. 2006;441(7097):1060–1060.
Kaiser J. The cancer stem cell gamble. Science. 2015;347(6219):226.
Hou Y, Wang S, Gao M, Chang J, Sun J, Qin L, Li A. Interferon-induced transmembrane protein 3 expression upregulation is involved in progression of hepatocellular carcinoma. BioMed Res Int. 2021;2021:5612138.
Min J, Feng Q, Liao W, Liang Y, Gong C, Li E, He W, Yuan R, Wu L. IFITM3 promotes hepatocellular carcinoma invasion and metastasis by regulating MMP9 through p38/MAPK signaling. FEBS Open Bio. 2018;8(8):1299–311.
Liu X, Chen L, Fan Y, Hong Y, Yang X, Li Y, Lu J, Lv J, Pan X, Qu F, et al. IFITM3 promotes bone metastasis of prostate cancer cells by mediating activation of the TGF-β signaling pathway. Cell Death Dis. 2019;10(7):517.
Zhao X, Li J, Winkler CA, An P, Guo J-T. IFITM genes, variants, and their roles in the control and pathogenesis of viral infections. Front Microbiol. 2019; 9(3228).
Yang M, Gao H, Chen P, Jia J, Wu S. Knockdown of interferon-induced transmembrane protein 3 expression suppresses breast cancer cell growth and colony formation and affects the cell cycle. Oncol Rep. 2013;30(1):171–8.
Min J, Hu J, Luo C, Zhu J, Zhao J, Zhu Z, Wu L, Yuan R. IFITM3 upregulates c-myc expression to promote hepatocellular carcinoma proliferation via the ERK1/2 signalling pathway. Biosci Trends. 2020;13(6):523–9.
Zhao B, Wang H, Zong G, Li P. The role of IFITM3 in the growth and migration of human glioma cells. BMC Neurol. 2013;13:210.
Lee J, Robinson ME, Ma N, Artadji D, Ahmed MA, **ao G, Sadras T, Deb G, Winchester J, Cosgun KN, et al. IFITM3 functions as a PIP3 scaffold to amplify PI3K signalling in B cells. Nature. 2020;588(7838):491–7.
Li D, Peng Z, Tang H, Wei P, Kong X, Yan D, Huang F, Li Q, Le X, Li Q, et al. KLF4-mediated negative regulation of IFITM3 expression plays a critical role in colon cancer pathogenesis. Clin Cancer Res. 2011;17(11):3558–68.
Hu J, Wang S, Zhao Y, Guo Q, Zhang D, Chen J, Li J, Fei Q, Sun Y. Mechanism and biological significance of the overexpression of IFITM3 in gastric cancer. Oncol Rep. 2014;32(6):2648–56.
Cheng JCCBM, Gonzales FA, Ye W, Greer S, Marquez VE, Jones PA, Selker EU. Inhibition of DNA methylation and reactivation of silenced genes by Zebularine. J Natl Cancer Inst. 2003.
Cho JY, Lim JY, Cheong JH, Park YY, Yoon SL, Kim SM, Kim SB, Kim H, Hong SW, Park YN, et al. Gene expression signature-based prognostic risk score in gastric cancer. Clin Cancer Res. 2011;17(7):1850–7.
Takashima A, Yamada Y, Nakajima TE, Kato K, Hamaguchi T, Shimada Y. standard first-line chemotherapy for metastatic gastric cancer in Japan has met the global standard: evidence from recent phase Iii trials. Gastrointest Cancer Res GCR. 2009;3(6):239–44.
Tung SL, Huang WC, Hsu FC, Yang ZP, Jang TH, Chang JW, Chuang CM, Lai CR, Wang LH. miRNA-34c-5p inhibits amphiregulin-induced ovarian cancer stemness and drug resistance via downregulation of the AREG-EGFR-ERK pathway. Oncogenesis. 2017;6: e326.
Dallas NA, **a L, Fan F, Gray MJ, Gaur P, van Buren G, Samuel S, Kim MP, Lim SJ, Ellis LM. Chemoresistant colorectal cancer cells, the cancer stem cell phenotype, and increased sensitivity to insulin-like growth factor-I receptor inhibition. Cancer Res. 2009;69(5):1951–7.
El Darsa H, El Sayed R, Abdel-Rahman O. MET inhibitors for the treatment of gastric cancer: what’s their potential? J Exp Pharmacol. 2020;12:349–61.
Huang WC, Jang TH, Tung SL, Yen TC, Chan SH, Wang LH. A novel miR-365-3p/EHF/keratin 16 axis promotes oral squamous cell carcinoma metastasis, cancer stemness and drug resistance via enhancing beta5-integrin/c-met signaling pathway. J Exp Clin Cancer Res. 2019;38(1):89.
Usatyuk PV, Fu P, Mohan V, Epshtein Y, Jacobson JR, Gomez-Cambronero J, Wary KK, Bindokas V, Dudek SM, Salgia R, et al. Role of c-Met/phosphatidylinositol 3-kinase (PI3k)/Akt signaling in hepatocyte growth factor (HGF)-mediated lamellipodia formation, reactive oxygen species (ROS) generation, and motility of lung endothelial cells. J Biol Chem. 2014;289(19):13476–91.
Liu Y, Ao X, Ding W, Ponnusamy M, Wu W, Hao X, Yu W, Wang Y, Li P, Wang J. Critical role of FOXO3a in carcinogenesis. Mol Cancer. 2018;17(1):104.
T** EP, Groen RW, Vogelzang I, Derksen PW, Klok MD, Meijer HP, van Eeden S, Pals ST, Spaargaren M. Functional analysis of HGF/MET signaling and aberrant HGF-activator expression in diffuse large B-cell lymphoma. Blood. 2006;107(2):760–8.
Matsuzaki H, Daitoku H, Hatta M, Tanaka K, Fukamizu A. Insulin-induced phosphorylation of FKHR (Foxo1) targets to proteasomal degradation. Proc Natl Acad Sci U S A. 2003;100(20):11285–90.
Peck B, Ferber EC, Schulze A. Antagonism between FOXO and MYC regulates cellular powerhouse. Front Oncol. 2013;3:96.
Shen A, Wang L, Huang M, Sun J, Chen Y, Shen YY, Yang X, Wang X, Ding J, Geng M. c-Myc alterations confer therapeutic response and acquired resistance to c-Met inhibitors in MET-addicted cancers. Cancer Res. 2015;75(21):4548–59.
Zeller KI, Zhao X, Lee CW, Chiu KP, Yao F, Yustein JT, Ooi HS, Orlov YL, Shahab A, Yong HC, et al. Global map** of c-Myc binding sites and target gene networks in human B cells. Proc Natl Acad Sci U S A. 2006;103(47):17834–9.
Blanco-Bose WE, Murphy MJ, Ehninger A, Offner S, Dubey C, Huang W, Moore DD, Trumpp A. C-Myc and its target FoxM1 are critical downstream effectors of constitutive androstane receptor (CAR) mediated direct liver hyperplasia. Hepatology. 2008;48(4):1302–11.
Lei M. The MCM complex: its role in DNA replication and implications for cancer therapy. Curr Cancer Drug Targets. 2005;5(5):365–80.
Li Y, Huang J, Sun J, **ang S, Yang D, Ying X, Lu M, Li H, Ren G. The transcription levels and prognostic values of seven proteasome alpha subunits in human cancers. Oncotarget. 2017;8(3):4501–19.
Kim JK, Noh JH, Eun JW, Jung KH, Bae HJ, Shen Q, Kim MG, Chang YG, Kim SJ, Park WS, et al. Targeted inactivation of HDAC2 restores p16INK4a activity and exerts antitumor effects on human gastric cancer. Mol Cancer Res. 2013;11(1):62–73.
Feng Y, **ong Y, Qiao T, Li X, Jia L, Han Y. Lactate dehydrogenase A: a key player in carcinogenesis and potential target in cancer therapy. Cancer Med. 2018;7(12):6124–36.
Bayraktar EC, La K, Karpman K, Unlu G, Ozerdem C, Ritter DJ, Alwaseem H, Molina H, Hoffmann HH, Millner A, et al. Metabolic coessentiality map** identifies C12orf49 as a regulator of SREBP processing and cholesterol metabolism. Nat Metab. 2020;2(6):487–98.
Jia Y, **ao Z, Jiang W, Chen G, Wang Z. Overexpression of IFITM3 predicts poor prognosis in stage IIA esophageal squamous cell carcinoma after Ivor Lewis esophagectomy. Thorac Cancer. 2017;8(6):592–9.
Liu Y, Lu R, Cui W, Pang Y, Liu C, Cui L, Qian T, Quan L, Dai Y, Jiao Y, et al. High IFITM3 expression predicts adverse prognosis in acute myeloid leukemia. Cancer Gene Ther. 2020;27(1–2):38–44.
Tian T, Zhang Y, Wang S, Zhou J, Xu S. Sox2 enhances the tumorigenicity and chemoresistance of cancer stem-like cells derived from gastric cancer. J Biomed Res. 2012;26(5):336–45.
Jonasson E, Ghannoum S, Persson E, Karlsson J, Kroneis T, Larsson E, Landberg G, Ståhlberg A. Identification of breast cancer stem cell related genes using functional cellular assays combined with single-cell RNA sequencing in MDA-MB-231 cells. Front Genet. 2019;10:500.
Chi H-C, Tsai C-Y, Wang C-S, Yang H-Y, Lo C-H, Wang W-J, Lee K-F, Lai L-Y, Hong J-H, Chang Y-F, et al. DOCK6 promotes chemo- and radioresistance of gastric cancer by modulating WNT/β-catenin signaling and cancer stem cell traits. Oncogene. 2020;39(37):5933–49.
Zhang F, Li K, Yao X, Wang H, Li W, Wu J, Li M, Zhou R, Xu L, Zhao L. A miR-567-PIK3AP1-PI3K/AKT-c-Myc feedback loop regulates tumour growth and chemoresistance in gastric cancer. EBioMedicine. 2019;44:311–21.
Madden SK, de Araujo AD, Gerhardt M, Fairlie DP, Mason JM. Taking the Myc out of cancer: toward therapeutic strategies to directly inhibit c-Myc. Mol Cancer. 2021;20(1):3.
Matsumoto K, Umitsu M, De Silva DM, Roy A, Bottaro DP. Hepatocyte growth factor/MET in cancer progression and biomarker discovery. Cancer Sci. 2017;108(3):296–307.
van der Schans JJ, van de Donk NWCJ, Mutis T. Dual targeting to overcome current challenges in multiple myeloma CAR T-cell treatment. Front Oncol. 2020. https://doi.org/10.3389/fonc.2020.01362.
Kleppe M, Koche R, Zou L, van Galen P, Hill CE, Dong L, De Groote S, Papalexi E, HanasogeSomasundara AV, Cordner K, et al. Dual targeting of oncogenic activation and inflammatory signaling increases therapeutic efficacy in myeloproliferative neoplasms. Cancer Cell. 2018;33(1):29-43.e27.
Jiang L, Zawacka-Pankau J. The p53/MDM2/MDMX-targeted therapies—a clinical synopsis. Cell Death Dis. 2020;11(4):237.
Yang J, Nie J, Ma X, Wei Y, Peng Y, Wei X. Targeting PI3K in cancer: mechanisms and advances in clinical trials. Mol Cancer. 2019;18(1):26.
Cui Q, Cai C-Y, Gao H-L, Ren L, Ji N, Gupta P, Yang Y, Shukla S, Ambudkar SV, Yang D-H, et al. Glesatinib, a c-MET/SMO dual inhibitor, antagonizes P-glycoprotein mediated multidrug resistance in cancer cells. Front Oncol. 2019. https://doi.org/10.3389/fonc.2019.00313.
Rimassa L, Bozzarelli S, Pietrantonio F, Cordio S, Lonardi S, Toppo L, Zaniboni A, Bordonaro R, Di Bartolomeo M, Tomasello G, et al. Phase II study of tivantinib and cetuximab in patients with KRAS wild-type metastatic colorectal cancer with acquired resistance to EGFR inhibitors and emergence of MET overexpression: lesson learned for future trials with EGFR/MET dual inhibition. Clin Colorectal Cancer. 2019;18(2):125-132.e122.
Tsai CY, Wang CS, Tsai MM, Chi HC, Cheng WL, Tseng YH, Chen CY, Lin CD, Wu JI, Wang LH, et al. Interleukin-32 increases human gastric cancer cell invasion associated with tumor progression and metastasis. Clin Cancer Res. 2014;20(9):2276–88.
Chi HC, Chen SL, Lin SL, Tsai CY, Chuang WY, Lin YH, Huang YH, Tsai MM, Yeh CT, Lin KH. Thyroid hormone protects hepatocytes from HBx-induced carcinogenesis by enhancing mitochondrial turnover. Oncogene. 2017;36(37):5274–84.
Chi HC, Tsai CY, Wang CS, Yang HY, Lo CH, Wang WJ, Lee KF, Lai LY, Hong JH, Chang YF, et al. DOCK6 promotes chemo- and radioresistance of gastric cancer by modulating WNT/beta-catenin signaling and cancer stem cell traits. Oncogene. 2020;39(37):5933–49.
Acknowledgements
We would like to thank Graduate Institute of Integrated Medicine & Chinese Medicine Research Center of China Medical University for providing us with materials to conduct experiments in the present study. This work were supported by the Ministry of Science and Technology, Taiwan (MOST 109-2320-B-039 -067, MOST 110-2320-B-039 -061, MOST 109-2314-B442-001), and China Medical University, Taiwan (CMU110-N-10), and Tainan Municipal An-Nan Hospital-China Medical University, Taiwan (ANHRF111-01), and the “Chinese Medicine Research Center, China Medical University” from The Featured Areas Research Center Program within the framework of the Higher Education Sprout Project by the Ministry of Education (MOE) in Taiwan (CMRC-CENTER-0), and the Show Chwan Memorial Hospital, Taiwan (SRD-109023 and SRD-109035) and Linko Chang-Gung Memorial Hospital, Taoyuan (CMRPD1L0111 to KHL).
Author information
Authors and Affiliations
Contributions
Conceptualization: HCC, KHL and PYC; methodology: WCH and CYT; formal analysis: PYC, WCH, SLT and CYT; investigation: CWL and YHL; data curation: WHL, CYC and HYL; writing the original draft preparation: HCC and CYT; writing, reviewing and editing: KHL, CCC, HYL and CHH. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of Interest
The authors have no conflicts to disclose.
Ethics approval and consent to participate
All experimental and research procedures were in accordance with relevant regulations. All patients diagnosed pathologically with gastric cancer (GC) at the Chang Gung Memorial Hospital (CGMH) from 2000 to 2005 were enlisted after informed consent. None of the patients had received radiotherapy and chemotherapy before surgery and all were undergone gastric resection. All the pathologic analyses and biological examination were conducted with informed consent. The research protocol was approved and verified by the Medical Ethics and Human Clinical Trial Committee of the Chang Gung Memorial Hospital (IRB NO. 201702000B0C101). The patients were followed regularly at the outpatient department in the Chang Gung Memorial Hospital every 3 months in the first 2 years, every 6 months between 3 to 5 years, and once a year thereafter. All animal experiments were performed in accordance with the Guide for Care and Use of Laboratory Animals issued by the Institutional Animal Care and Use Committee of Chang Gung University and the National Institutes of Health of United States (CGU106-142).
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Additional file 1: Fig. S1.
MCM2-7 proteins are regulated by IFITM3 in GC proliferation. (A) MetaCore analysis of 415 overexpressed proteins in GC tumors from our proteomic database. The signature genes involved in cell cycle regulation, metabolism and transcriptional regulation were shown. (B) Relative MCM2-7 proteins in GC tissues as compared to adjacent normal gastric tissues from our iTRAQ quantitative proteomic analysis (C) Spearman rank correlation coefficient of individual genes (including MCM2, MCM3, MCM4, MCM5, MCM6 and MCM7) against IFITM3 in 71 GC tissues from Cho Gastric dataset (GSE138861). (D) Quantitative RT-PCR analyses were conducted on MCM2-7 genes for their transcript expression in TMK-1 IFITM3-overexpression or TSGH IFITM3-depletion models (*p<0.05, **p<0.01).
Additional file 2: Table S1.
The 415 overexpressed proteins in GC tumors from our iTRAQ quantitative proteomic database.
Additional file 3: Table S2.
The repeated presented protein candidates in iTRAQ quantitative proteomic analysis.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Chu, PY., Huang, WC., Tung, SL. et al. IFITM3 promotes malignant progression, cancer stemness and chemoresistance of gastric cancer by targeting MET/AKT/FOXO3/c-MYC axis. Cell Biosci 12, 124 (2022). https://doi.org/10.1186/s13578-022-00858-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s13578-022-00858-8