Abstract
Background
Pluripotent states of embryonic stem cells (ESCs) with distinct transcriptional profiles affect ESC differentiative capacity and therapeutic potential. Although single-cell RNA sequencing has revealed additional subpopulations and specific features of naive and primed human pluripotent stem cells (hPSCs), the underlying mechanisms that regulate their specific transcription and that control their pluripotent states remain elusive.
Results
By single-cell analysis of high-resolution, three-dimensional (3D) genomic structure, we herein demonstrate that remodeling of genomic structure is highly associated with the pluripotent states of human ESCs (hESCs). The naive pluripotent state is featured with specialized 3D genomic structures and clear chromatin compartmentalization that is distinct from the primed state. The naive pluripotent state is achieved by remodeling the active euchromatin compartment and reducing chromatin interactions at the nuclear center. This unique genomic organization is linked to enhanced chromatin accessibility on enhancers and elevated expression levels of naive pluripotent genes localized to this region. In contradistinction, the primed state exhibits intermingled genomic organization. Moreover, active euchromatin and primed pluripotent genes are distributed at the nuclear periphery, while repressive heterochromatin is densely concentrated at the nuclear center, reducing chromatin accessibility and the transcription of naive genes.
Conclusions
Our data provide insights into the chromatin structure of ESCs in their naive and primed states, and we identify specific patterns of modifications in transcription and chromatin structure that might explain the genes that are differentially expressed between naive and primed hESCs. Thus, the inversion or relocation of heterochromatin to euchromatin via compartmentalization is related to the regulation of chromatin accessibility, thereby defining pluripotent states and cellular identity.
Similar content being viewed by others
Background
Although human pluripotent stem cells (hPSCs) and notably embryonic stem cells (ESCs) hold great promise in the regeneration of tissues, conventional hESCs are essentially classified as primed PSCs, like mouse epiblast stem cells (mEpiSCs) that exhibit lower differentiative capability [1,2,3,4]—limiting their potentially broad applicability. Significant progress has enabled the conversion of primed hESCs to the naive state of pluripotency, resembling well-characterized naive mouse ESCs (mESCs) [5,6,7,8,9,10,11]. Intriguingly, naive hPSCs can generate blastocyst-like structures in vitro under effective three-dimensional (3D) culture conditions [12]. Naive hPSCs express higher levels of specific pluripotent genes such as OCT4, NANOG, STELLA/DPPA3, and DPPA5—as well as the specific endogenous retrovirus HERVH—whereas primed hPSCs particularly express ZIC2, OTX2, and B3GAT1 [7, 8, 11, 13,18, 19], the mechanisms that regulate the specific transcription and control of these states have remained elusive. By taking advantage of a single-cell, diploid chromatin conformation-capture method termed Dip-C that reveals sufficiently high-resolution genomic structures [23], and our results revealed a difference from the previous assessment of CpG frequency. There was also a similar trend with respect to primed central aggregation of additional B components. We ascertained that the active euchromatin organization or compartmentalization at the nuclear center of the naive state and the repressive heterochromatic compartmentalization in the central region of the primed sate in hESCs appeared to be active or open chromatin (A) or repressed or closed chromatin (B) compartments, respectively, and that they were distinct from the A-B compartment switch reported previously [23,24,26, 27]. To assess whether the expression of pluripotent genes was linked to the enhancer chromatin structure, we analyzed the distribution of the enhancers at the gene loci in primed and naive hESCs. Our data revealed that more enhancers were enriched around the loci of the naive pluripotent genes in naive hESCs, and more enhancers were distributed around the loci of primed pluripotent genes in primed hESCs (Additional file 1: Fig. S4b). Moreover, primed pluripotent genes showed overlap of the loci with the enhancers in primed hESCs, and naive pluripotent genes overlapped at the loci with enhancers in naive hESCs (Fig. 6a, Additional file 1: Fig. S8). These results are consistent with the model in which chromatin is folded to form an enhancer–promoter loop, thus facilitating transcription.
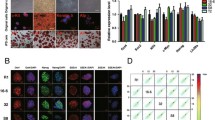
Transcriptional regulation and chromatin accessibility in naive and primed states. a Radial positioning along the genome of primed or naive genes and enhancers in naive or primed hESCs. b Heatmaps of ATAC-seq signal distribution around the transcriptional start site (TSS) ± 3000 bp of expressed genes and average profiles of the enrichment at the TSS in naive or primed hESCs (two replicates are shown for each state). c 3D structural differences between the two alleles of the representative marker-gene loci in primed and naive cells. d Joint analysis by ATAC-seq, CUT&Tag, and RNA-seq of the relationships between gene expression and chromatin accessibility in naive (NANOG) and primed states (ZIC2). Genome browser tracks of RNA-seq, ATAC-seq, and CUT&Tag-seq data of CTCF, H3K9me3, and H3K27me3 at the NANOG and ZIC2 loci in naive and primed hESCs
To form an enhancer–promoter loop, a relaxed open chromatin structure is required. Map** open chromatin using an assay for transposase-accessible chromatin (ATAC-seq) previously revealed distinct chromatin accessibility of naive and primed hPSCs [22]. To explore the consequences of distinct genomic organization of primed and naive states and to discern whether genomic spatial organization facilitated chromatin accessibility, we performed low (cell number) input ATAC-seq on our naive hESCs and compared them with primed hESCs. ATAC-seq detected more open euchromatin structure in naive cells (Fig. 6b), confirming the Dip-C analysis, showing that the naive state possessed a smaller number of chromatin contacts (Fig. 4i). When we simulated the contact morphology of primed and naive pluripotent genes localized in the chromatin based on a previously reported method [39]. In addition, TADs may represent a “population average” of individual loops that differ on a cell-to-cell basis. It is acknowledged that TADs are indeed present in individual cells and that their observed hierarchy may reflect multimeric associations between individual regions within the TAD [39]. 3D chromosomal structures based on Hi-C chromosomal conformation capture data in population cells show that TADs are largely preserved during the transition between the naive and primed states of hESCs [40], and TAD structure is also revealed by single-cell Dip-C. While enhancers are occupied by transcription factors, mediators, and cohesin—and their associated nucleosomes marked by H3K27ac [40]—histone modifications themselves may not be required for chromatin organization in these differential states. Loss of H3K27ac, however, perturbs transcriptional but not 3D chromatin architectural resetting [31].
mESCs and mEpiSCs can be distinguished by active and inactive compartmental organization and switching in sub-nuclear positioning that is associated with replication timing [41], where it appears that the heterochromatin (B) compartment is located in the nuclear periphery, whereas the active euchromatin (A) compartment is in the interior in both naive mESCs and mEpiSCs. We herein showed that primed hESCs form inactive compartments enriched with heterochromatin at the nuclear center and active euchromatin at the nuclear periphery that are distinct from the active euchromatin and naive pluripotent genes localized at the nuclear center in naive hESCs. Naive pluripotent gene networks between human and mouse PSCs are not well conserved and more closely resemble their respective blastocysts [42]. In fact, the naive pluripotent state observed for mouse ESCs has been difficult to capture in hESCs, appearing to be transitory in the human embryo itself. Thus, the direct application of mouse embryology to humans has not always been successful due to fundamental developmental differences between the two species [43]. There were also some differences between the naive hESC lines derived directly from the early embryo [5, 44] and those that we employed in the present study. The naive cells we used in this work were derived from the conversion of the primed hESC line H9 by an LTR7-GFP reporter, and we maintained the HERVH hyper-activation required for human pluripotency. The expression profile of naive cells enriched using the HERVH reporter in our analysis most closely resembled the inner cell mass when compared with the naive cells obtained by Gafni et al. [5] and showed a similar XIST expression and H3K27me3 distribution pattern around the X chromosome [26]. Cells were permeabilized in 500 μL of ice-cold Hi-C lysis buffer (10 mM Tris, pH 8.0; 10 mM NaCl; 0.2% IGEPAL CA 630) and 100 μL of protease inhibitor (Sigma, P8340) for ≥ 15 min on ice, washed in cold Hi-C lysis buffer on ice (with centrifugation at 2500 g for 5 min), and further permeabilized in 50 μL of 0.5% SDS at 62 °C for 10 min. The SDS was quenched by adding 145 μL of water and 25 μL of 10% Triton X-100 and incubated at 37 °C for 15 min with rotation. The cells were then digested by adding 25 μL of 10 × NEB Buffer 2 and 20 μL of 25 U/μL MboI (NEB, R0147M) and incubated overnight at 37 °C with rotation. On the second day, the cells were washed with 1 mL of ligation buffer (1 × T4 DNA ligase buffer, NEB B0202S) and 0.1 mg/mL BSA (NEB B9000S) and ligated in 1 mL of ligation buffer and 10 μL of 1 U/μL T4 DNA ligase (Life Tech, 15,224–025) at 16 °C for 20 h.
Single-cell isolation by flow cytometry
The ligated cells (in ligation buffer) were filtered through a 40-μm cell strainer (Falcon) and sorted into 0.2-mL UV-irradiated, DNA low-bind tubes (MAXYMum Recovery, Axygen) containing lysis buffer using a FACSAria III flow cytometer (BD, 85-μm nozzle). The area-scaling factor was set, and forward scatter (FSC)-A and side scatter (SSC)-A were used to exclude large-sized cellular structures or debris, and scatter SSC-W was set to avoid contamination by doublets or triplets. Single cells were sorted into a PCR tube by applying the “1.0 drop single” sorting mode. The collected single cells were stored for several months at − 80 °C.
Whole-genome amplification in Dip-C
Appreciable DNA contact information can be lost in traditional bulk Hi-C, but we greatly reduced any loss by inserting n different tags. As a result, only 1/n of input DNA contacts was lost in our study. We implemented META with n = 20 tags, and the sequences were treated according to previously described methods [Haplotype imputation (2D) In each round of imputation, contacts in an “evidence” set voted to impute unknown haplotypes of contacts in a “target” set. For each target contact, a list of compatible haplotype tuples was first enumerated. Each evidence contact would then vote for haplotype tuples from this list, if such contact fell within 10 Mb in L0.5 distance from the target contact and was compatible with one and only one haplotype tuple from the list. Imputation would occur if the winning haplotype tuple gathered ≥ 3 votes and ≥ 90% of all votes. Special care was taken for intrachromosomal contacts because intrahomologous contacts were far more frequent than interhomologous contacts, especially at short ranges (small genomic separation). A target contact would be assumed intrahomologous without voting, if its two legs were separated by ≤ 10 Mb; otherwise, voting still occurred but a winning interhomologous vote would only be accepted if two legs were separated by ≥ 100 Mb. In addition, intrachromosomal contacts that had unknown haplotypes on both legs were not imputed. One leg as both the target and the evidence sets was estimated for the contact location. Such imputation was repeated two more times, each time with previous results as the new evidence set. Results were subsequently cleaned by removal of isolated contacts (< 2 other contacts that had the same haplotypes within 10 Mb in L0.5 distance). Finally, cleaned results were used as the evidence set to impute a target set of all interchromosomal contacts that had unknown haplotypes on both legs. Simulated annealing was performed by nuc_dynamics [50] as a peak caller, with a q-value (FDR) threshold of 0.01. Based on a dynamic Poisson distribution, MACS2 can effectively capture local biases in the genome sequence, allowing for more sensitive and robust prediction of binding sites. Unique read for a position for peak calling is used to reduce false positive peaks, statically significant peaks are finally selected by calculated false discovery rate of reported peaks. Deeptools [51] is used for the heatmap plots. ATAC-seq peaks from all study samples were merged to create a union set of sites. Read densities were calculated for each peak for each sample, differential peaks between Naive and Primed were identified by DEseq2 [52] with adjusted P ≤ 0.05, |log2fold change|≥ 1. Cut-tag: use FastQC v0.11.9 for quality control of raw sequencing readings. Using TrimGalore v0.6.6 to remove raw readings from low-quality base and linker sequences(https://github.com/FelixKrueger/TrimGalore). Compare the filtered reading with the reference mouse genome assembly mm10 of the mouse sample and the human genome assembly GRCh38 of the human sample using Bowtie2 v2.4.4. The options are end-to-end, very sensitive, no mixing, no inconsistency, phred33-I 10-X 700. Use the sorting function of samtools v1.13 to sort aligned bam files based on chromosome coordinates. Use the genomecov function of bedtools v2.30 to summarize the sorted bam files into a bedgraph file (Quinlan et al., 2010) [53]. In the case of samples with multiple biological replicates, use the unionBeg function combination of bedtools v2.30 to replicate specific bed chart files. In strict mode, perform peak calls on all bedgraph files using SEACR v1.3 by selecting the top 1% of the call peak. SEACR is specifically developed for CUT&RUN and is also a recommended pipeline for chromatin analysis data with very low background, such as CUT&Tag. Perform visual QC on peak values of bam files and calls using Seqmonk [54].3D reconstruction
Availability of data and materials
Raw sequencing data reported in this paper were uploaded into NCBI with accession number PRJNA739271 [55]. And all the RNA-seq, CUT&Tag-seq and ATAC-seq processed data are available from Gene Expression Omnibus under accession number GEO: GSE260995 [56]. The bulk HiC data used in Fig. 4f are available from Genome Sequence Archive with the accession number CRA000852 [57]. The method to distinguish A/B parts in Fig. 4e is refereed to Lieberman Aiden [23]. No other scripts and software were used other than those mentioned in the “ Methods” section. All cell lines used in this study have been authenticated and are available upon request.
References
Wu J, Izpisua Belmonte JC. Dynamic pluripotent stem cell states and their applications. Cell Stem Cell. 2015;17:509–25.
Yilmaz A, Benvenisty N. Defining human pluripotency. Cell Stem Cell. 2019;25:9–22.
Nichols J, Smith A. Naive and primed pluripotent states. Cell Stem Cell. 2009;4:487–92.
Li M, Belmonte JC. Ground rules of the pluripotency gene regulatory network. Nat Rev Genet. 2017;18:180–91.
Gafni O, Weinberger L, Mansour AA, Manor YS, Chomsky E, Ben-Yosef D, Kalma Y, Viukov S, Maza I, Zviran A, et al. Derivation of novel human ground state naive pluripotent stem cells. Nature. 2013;504:282–6.
Hackett JA, Surani MA. Regulatory principles of pluripotency: from the ground state up. Cell Stem Cell. 2014;15:416–30.
Theunissen TW, Powell BE, Wang H, Mitalipova M, Faddah DA, Reddy J, Fan ZP, Maetzel D, Ganz K, Shi L, et al. Systematic identification of culture conditions for induction and maintenance of naive human pluripotency. Cell Stem Cell. 2014;15:524–6.
Theunissen TW, Friedli M, He Y, Planet E, O’Neil RC, Markoulaki S, Pontis J, Wang H, Iouranova A, Imbeault M, et al. Molecular criteria for defining the naive human pluripotent state. Cell Stem Cell. 2016;19:502–15.
Takashima Y, Guo G, Loos R, Nichols J, Ficz G, Krueger F, Oxley D, Santos F, Clarke J, Mansfield W, et al. Resetting transcription factor control circuitry toward ground-state pluripotency in human. Cell. 2015;162:452–3.
Goke J, Lu X, Chan YS, Ng HH, Ly LH, Sachs F, Szczerbinska I. Dynamic transcription of distinct classes of endogenous retroviral elements marks specific populations of early human embryonic cells. Cell Stem Cell. 2015;16:135–41.
Guo G, von Meyenn F, Rostovskaya M, Clarke J, Dietmann S, Baker D, Sahakyan A, Myers S, Bertone P, Reik W, et al. Epigenetic resetting of human pluripotency. Development. 2017;144:2748–63.
Yu L, Wei Y, Duan J, Schmitz DA, Sakurai M, Wang L, Wang K, Zhao S, Hon GC, Wu J. Blastocyst-like structures generated from human pluripotent stem cells. Nature. 2021;591:620–6.
Lu X, Sachs F, Ramsay L, Jacques PE, Goke J, Bourque G, Ng HH. The retrovirus HERVH is a long noncoding RNA required for human embryonic stem cell identity. Nat Struct Mol Biol. 2014;21:423–5.
Wang J, **e G, Singh M, Ghanbarian AT, Rasko T, Szvetnik A, Cai H, Besser D, Prigione A, Fuchs NV, et al. Primate-specific endogenous retrovirus-driven transcription defines naive-like stem cells. Nature. 2014;516:405–9.
Weinberger L, Ayyash M, Novershtern N, Hanna JH. Dynamic stem cell states: naive to primed pluripotency in rodents and humans. Nat Rev Mol Cell Biol. 2016;17:155–69.
Gao X, Nowak-Imialek M, Chen X, Chen D, Herrmann D, Ruan D, Chen ACH, Eckersley-Maslin MA, Ahmad S, Lee YL, et al. Establishment of porcine and human expanded potential stem cells. Nat Cell Biol. 2019;21:687–99.
Kilens S, Meistermann D, Moreno D, Chariau C, Gaignerie A, Reignier A, Lelievre Y, Casanova M, Vallot C, Nedellec S, et al. Parallel derivation of isogenic human primed and naive induced pluripotent stem cells. Nat Commun. 2018;9:360.
Han X, Chen H, Huang D, Chen H, Fei L, Cheng C, Huang H, Yuan GC, Guo G. Map** human pluripotent stem cell differentiation pathways using high throughput single-cell RNA-sequencing. Genome Biol. 2018;19:47.
Messmer T, von Meyenn F, Savino A, Santos F, Mohammed H, Lun ATL, Marioni JC, Reik W. Transcriptional heterogeneity in naive and primed human pluripotent stem cells at single-cell resolution. Cell Rep. 2019;26(815–824):e814.
Tan L, **ng D, Chang CH, Li H, **e XS. Three-dimensional genome structures of single diploid human cells. Science. 2018;361:924–8.
Szczerbinska I, Gonzales KAU, Cukuroglu E, Ramli MNB, Lee BPG, Tan CP, Wong CK, Rancati GI, Liang H, Goke J, et al. A chemically defined feeder-free system for the establishment and maintenance of the human naive pluripotent state. Stem Cell Reports. 2019;13:612–26.
Pastor WA, Liu W, Chen D, Ho J, Kim R, Hunt TJ, Lukianchikov A, Liu X, Polo JM, Jacobsen SE, Clark AT. TFAP2C regulates transcription in human naive pluripotency by opening enhancers. Nat Cell Biol. 2018;20:553–64.
Lieberman-Aiden E, van Berkum NL, Williams L, Imakaev M, Ragoczy T, Telling A, Amit I, Lajoie BR, Sabo PJ, Dorschner MO, et al. Comprehensive map** of long-range interactions reveals folding principles of the human genome. Science. 2009;326:289–93.
Vilarrasa-Blasi R, Soler-Vila P, Verdaguer-Dot N, Russinol N, Di Stefano M, Chapaprieta V, Clot G, Farabella I, Cusco P, Kulis M, et al. Dynamics of genome architecture and chromatin function during human B cell differentiation and neoplastic transformation. Nat Commun. 2021;12:651.
Dixon JR, Jung I, Selvaraj S, Shen Y, Antosiewicz-Bourget JE, Lee AY, Ye Z, Kim A, Rajagopal N, **e W, et al. Chromatin architecture reorganization during stem cell differentiation. Nature. 2015;518:331–6.
Rao SS, Huntley MH, Durand NC, Stamenova EK, Bochkov ID, Robinson JT, Sanborn AL, Machol I, Omer AD, Lander ES, Aiden EL. A 3D map of the human genome at kilobase resolution reveals principles of chromatin loo**. Cell. 2014;159:1665–80.
Haberle V, Stark A. Eukaryotic core promoters and the functional basis of transcription initiation. Nat Rev Mol Cell Biol. 2018;19:621–37.
Lu JY, Chang L, Li T, Wang T, Yin Y, Zhan G, Han X, Zhang K, Tao Y, Percharde M, et al. Homotypic clustering of L1 and B1/Alu repeats compartmentalizes the 3D genome. Cell Res. 2021;31:613–30.
Zhang Y, Li T, Preissl S, Amaral ML, Grinstein JD, Farah EN, Destici E, Qiu Y, Hu R, Lee AY, et al. Transcriptionally active HERV-H retrotransposons demarcate topologically associating domains in human pluripotent stem cells. Nat Genet. 2019;51:1380–8.
Stadhouders R, Vidal E, Serra F, Di Stefano B, Le Dily F, Quilez J, Gomez A, Collombet S, Berenguer C, Cuartero Y, et al. Transcription factors orchestrate dynamic interplay between genome topology and gene regulation during cell reprogramming. Nat Genet. 2018;50:238–49.
Atlasi Y, Megchelenbrink W, Peng T, Habibi E, Joshi O, Wang SY, Wang C, Logie C, Poser I, Marks H, Stunnenberg HG. Epigenetic modulation of a hardwired 3D chromatin landscape in two naive states of pluripotency. Nat Cell Biol. 2019;21:568–78.
Barakat TS, Halbritter F, Zhang M, Rendeiro AF, Perenthaler E, Bock C, Chambers I. Functional dissection of the enhancer repertoire in human embryonic stem cells. Cell Stem Cell. 2018;23(276–288):e278.
Feng Y, Wang Y, Wang X, He X, Yang C, Naseri A, Pederson T, Zheng J, Zhang S, **ao X, et al. Simultaneous epigenetic perturbation and genome imaging reveal distinct roles of H3K9me3 in chromatin architecture and transcription. Genome Biol. 2020;21:296.
Falk M, Feodorova Y, Naumova N, Imakaev M, Lajoie BR, Leonhardt H, Joffe B, Dekker J, Fudenberg G, Solovei I, Mirny LA. Heterochromatin drives compartmentalization of inverted and conventional nuclei. Nature. 2019;570:395–9.
Lynch CJ, Bernad R, Martinez-Val A, Shahbazi MN, Nobrega-Pereira S, Calvo I, Blanco-Aparicio C, Tarantino C, Garreta E, Richart-Gines L, et al. Global hyperactivation of enhancers stabilizes human and mouse naive pluripotency through inhibition of CDK8/19 Mediator kinases. Nat Cell Biol. 2020;22:1223–38.
Atlasi Y, Stunnenberg HG. The interplay of epigenetic marks during stem cell differentiation and development. Nat Rev Genet. 2017;18:643–58.
Schlesinger S, Meshorer E. Open chromatin, epigenetic plasticity, and nuclear organization in pluripotency. Dev Cell. 2019;48:135–50.
Zheng H, **e W. The role of 3D genome organization in development and cell differentiation. Nat Rev Mol Cell Biol. 2019;20:535–50.
Dixon JR, Gorkin DU, Ren B. Chromatin domains: the unit of chromosome organization. Mol Cell. 2016;62:668–80.
Ji X, Dadon DB, Powell BE, Fan ZP, Borges-Rivera D, Shachar S, Weintraub AS, Hnisz D, Pegoraro G, Lee TI, et al. 3D chromosome regulatory landscape of human pluripotent cells. Cell Stem Cell. 2016;18:262–75.
Miura H, Takahashi S, Poonperm R, Tanigawa A, Takebayashi SI, Hiratani I. Single-cell DNA replication profiling identifies spatiotemporal developmental dynamics of chromosome organization. Nat Genet. 2019;51:1356–68.
Huang K, Maruyama T, Fan G. The naive state of human pluripotent stem cells: a synthesis of stem cell and preimplantation embryo transcriptome analyses. Cell Stem Cell. 2014;15:410–5.
Rossant J, Tam PPL. New insights into early human development: lessons for stem cell derivation and differentiation. Cell Stem Cell. 2017;20:18–28.
Ware CB, Nelson AM, Mecham B, Hesson J, Zhou W, Jonlin EC, Jimenez-Caliani AJ, Deng X, Cavanaugh C, Cook S, et al. Derivation of naive human embryonic stem cells. Proc Natl Acad Sci U S A. 2014;111:4484–9.
Amano T, Sagai T, Tanabe H, Mizushina Y, Nakazawa H, Shiroishi T. Chromosomal dynamics at the Shh locus: limb bud-specific differential regulation of competence and active transcription. Dev Cell. 2009;16:47–57.
Kaya-Okur HS, Wu SJ, Codomo CA, Pledger ES, Bryson TD, Henikoff JG, Ahmad K, Henikoff S. CUT&Tag for efficient epigenomic profiling of small samples and single cells. Nat Commun. 1930;2019:10.
Wu J, Xu J, Liu B, Yao G, Wang P, Lin Z, Huang B, Wang X, Li T, Shi S, et al. Chromatin analysis in human early development reveals epigenetic transition during ZGA. Nature. 2018;557:256–60.
Mei S, Qin Q, Wu Q, Sun H, Zheng R, Zang C, Zhu M, Wu J, Shi X, Taing L, et al. Cistrome Data Browser: a data portal for ChIP-Seq and chromatin accessibility data in human and mouse. Nucleic Acids Res. 2017;45:D658-d662.
Li H, Durbin R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics. 2009;25:1754–60.
Zhang Y, Liu T, Meyer CA, Eeckhoute J, Johnson DS, Bernstein BE, Nusbaum C, Myers RM, Brown M, Li W, Liu XS. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 2008;9:R137.
Ramírez F, Ryan DP, Grüning B, Bhardwaj V, Kilpert F, Richter AS, Heyne S, Dündar F, Manke T. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 2016;44:W160-165.
Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15:550.
Quinlan A, Hall I. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics. 2010;26:841–2.
Galle E, Wong CW, Ghosh A, Desgeorges T, Melrose K, Hinte LC, Castellano-Castillo D, Engl M, de Sousa JA, Ruiz-Ojeda FJ, et al. H3K18 lactylation marks tissue-specific active enhancers. Genome Biol. 2022;23:207.
Li N, ** K, Liu B, Yang M, Shi P, Heng D, Wang J, Liu L: Single-cell 3D genome structure reveals distinct human pluripotent states. Raw sequencing data. https://www.ncbi.nlm.nih.gov/bioproject/PRJNA739271. (2024).
Li N, ** K, Liu B, Yang M, Shi P, Heng D, Wang J, Liu L: Single-cell 3D genome structure reveals distinct human pluripotent states. Expression profiling and genome binding/occupancy profiling sequencing data. https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE260995. (2024).
Chen X, Ke Y, Wu K, Zhao H, Sun Y, Gao L, Liu Z, Zhang J, Tao W, Hou Z, et al. Key role for CTCF in establishing chromatin structure in human embryos. Nature. 2019;576:306–10.
Acknowledgements
We thank Zhenzhen Lin, Zhe Liu, and Ge Zhan for the kind assistance with DNA FISH protocol.
Peer review information
Wen**g She was the primary editor of this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Review history
The review history is available as Additional file 4.
Funding
This study was supported by Projects of International Cooperation and Exchanges NSFC (32261160571), National Natural Science Foundation of China (Key Program, 82230052), National Natural Science Foundation for Youth (32100418), and Haihe Laboratory of Cell Ecosystem Innovation Fund (22HHXBSS00029).
Author information
Authors and Affiliations
Contributions
N.L. performed major experiments, ESC culture and experiment, RNA-seq, ATAC-seq, Dip-C, CUT&Tag and data analysis, and prepared the figures and the manuscript draft. B.L. helped RNA-seq, ATAC-seq, Dip-C, and data analysis. K.J. performed immunofluorescence microcopy, DNA-FISH, RNA-FISH, Dip-C, ATAC-seq, CUT&Tag and western blot and participated in the revision of manuscript draft. M.Y. provided materials and consulted hESC cultures. P.S. and D.H. assisted with the western blot and immunofluorescence microscopy. J.W. generated and supervised extended naive human ESC cultures and primed hESCs, discussed the project, and revised the manuscript. L.L. conceptualized and supervised the study and designed the experiments, and revised the manuscript. All authors approved and contributed to the final version of the manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
Ethical approval is not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
13059_2024_3268_MOESM1_ESM.xlsx
Additional file 1: Fig S1. Transcriptome of naive and primed hESCs. Fig S2. Simulation of 3D structure of single-cell chromatin in naive and primed hESCs. Fig S3. Distinct 3D structures of chromosomes in naive and primed hESCs. Fig S4. Features of localization of marker gene locus and regulation in primed and naive hESCs. Fig S5. Fluorescence intensity distribution of epigenetic histone modifications in the nucleus. Fig S6. Distribution of epigenetic histone modifications and LINE1/Alu in primed and naive hESCs. Fig S7. Genomic localization of marker genes, HERVH, and expression of LINE-1-orf-1p in primed and naive hESCs. Fig S8. Radial positioning along the genome of primed or naive genes and enhancer in naive or primed hESCs. Fig S9. Uncropped scans of Western blot.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Li, N., **, K., Liu, B. et al. Single-cell 3D genome structure reveals distinct human pluripotent states. Genome Biol 25, 122 (2024). https://doi.org/10.1186/s13059-024-03268-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s13059-024-03268-w