Abstract
Background
Head and neck squamous cell carcinoma (HNSCC) remains an unmet medical challenge. Metabolic reprogramming is a hallmark of diverse cancers, including HNSCC.
Methods
We investigated the metabolic profile in HNSCC by using The Cancer Genome Atlas (TCGA) (n = 481) and Gene Expression Omnibus (GEO) (n = 97) databases. The metabolic stratification of HNSCC samples was identified by using unsupervised k-means clustering. We analyzed the correlations of the metabolic subtypes in HNSCC with featured genomic alterations and known HNSCC subtypes. We further validated the metabolism-related subtypes based on features of ENO1, PFKFB3, NSDHL and SQLE expression in HNSCC by Immunohistochemistry. In addition, genomic characteristics of tumor metabolism that varied among different cancer types were confirmed.
Results
Based on the median expression of coexpressed cholesterogenic and glycolytic genes, HNSCC subtypes were identified, including glycolytic, cholesterogenic, quiescent and mixed subtypes. The quiescent subtype was associated with the longest survival and was distributed in stage I and G1 HNSCC. Mutation analysis of HNSCC genes indicated that TP53 has the highest mutation frequency. The CDKN2A mutation frequency has the most significant differences amongst these four subtypes. There is good overlap between our metabolic subtypes and the HNSCC subtype.
Conclusion
The four metabolic subtypes were successfully determined in HNSCC. Compared to the quiescent subtype, glycolytic, cholesterogenic and mixed subtypes had significantly worse outcome, which might offer guidelines for develo** a novel treatment strategy for HNSCC.
Similar content being viewed by others
Introduction
Head and neck squamous cell carcinoma (HNSCC) is a heterogeneous disease comprising tumors of the oral cavity, lip, oropharynx, nasopharynx, larynx, hypopharynx and salivary gland [1]. HNSCC is the sixth most common malignancy in humans worldwide, with 930,000 newly diagnosed cases and 467,000 deaths in 2020 [2]. An estimated 1.37 million new cases are projected to occur in 2040, representing a 32% increase [3]. Current research indicates that HNSCC is closely related to numerous factors, including smoking, drinking, and human papilloma virus [1]. Despite advances in treatment strategies and improved prognosis, HNSCC remains an incurable malignancy, with approximately half of patients relapsing and dying from the disease [4]. Additionally, histopathology and clinical stage are not sufficient to accurately predict the prognosis of a patient because of the heterogeneity of HNSCC [1].
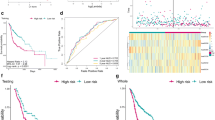
Recently, tumor-related metabolic reprogramming has been extensively studied, offering an approach to target cancers [5]. Metabolic reprogramming is recognized as a hallmark of cancer and presents opportunities for cancer diagnosis, prognosis, and therapy [6,7,8]. Cancer cells accumulate metabolic alterations to meet energetic demands and produce biosynthetic precursors, such as glucose, nucleotides, fatty acids and amino acids, for rapid tumor growth [8,9,10,11]. Such metabolic alterations can affect the fate of cancer. Recent studies have shown that the process of HNSCC emergence is related to tumor metabolism, which is mainly characterized by abnormal glycolysis and cholesterol synthesis. On the one hand, cancer cells utilize glycolysis for producing energy to promote the proliferation of cancer cells [12]. Molecular imaging studies using 18F-fluoro-2-deoxy-d-glucose positron emission tomography demonstrated increased glucose uptake and glycolysis in HNSCC [13, 14]. Increased glycolysis correlates with aggressive tumor progression, treatment resistance, and unfavorable prognosis in HNSCC [15, 16].On the other hand, cancer cells require high levels of cholesterol for membrane biogenesis and other functional needs, and subsequently promotes tumor development [17]. Meanwhile, certain investigations have indicated several key enzymes of cholesterol synthesis are closely related to poor prognosis of HNSCC [18]. Avasimibe, a specific inhibitor of ACAT, significantly inhibited tumor growth and prolonged survival by inhibiting the accumulation of cholesterol ester [ The GEO (https://www.icgc.org, GSE41613) [26] and TCGA (Illumina HiSeq Systems;) data portals were used to obtain HNSCC datasets together with related clinical data. In addition, the standard RNA sequencing data of the 481 TCGA-derived patients and 97 GSE41613-derived patients were collected. The human genome reference sequence GRCh37 formulated by the Genome Reference Consortium was used. In addition, somatic mutational data (SNVs, CNVs, and INDELs) were collected for each sample. RNA expression of every sample was normalized by the transcripts per million algorithms, which was later log-transformed into log10 ((normalized count*1e6145) + 1). A log2-fold change (FC) ≥ 1 was used as the threshold to select RNAs with significant differential expression. Samples whose tumor content was < 30% were eliminated from this work [27]. Genes obtained from the gene sets of the molecular signatures database (mSigDB) [28], namely, “REACTOME CHOLESTEROL BIOSYNTHESIS” (n = 24) and “REACTOME GLYCOLYSIS” (n = 29), were identified as cholesterogenic and glycolytic genes, respectively. Then, these genes were subjected to consensus clustering by adopting ConsensusClusterPlus (parameters: reps = 100, pFeature = 1, pItem = 0.8) [29]. Meanwhile, the Euclidean distance (ED) and Ward. D2 were adopted as the distance matrix and the clustering algorithm, respectively, with k = 4 (Additional file 1: Figure S1). In addition, the median expression of coexpressed cholesterogenic and glycolytic genes was utilized to assign the quiescent (glycolytic ≤ 0, cholesterogenic ≤ 0), cholesterogenic (glycolytic ≤ 0, cholesterogenic > 0), glycolytic (glycolytic > 0, cholesterogenic ≤ 0), or mixed (glycolytic > 0, cholesterogenic > 0) metabolic subtypes for every sample. Samples were classified by consistent clustering according to common tumor subtypes investigated by Weidong Zhang et al.[30] and Hongbo Zhou et al.[31]. Typically, subty** was classified according to 6 mRNA expression levels in the original paper by Weidong Zhang, whereas subty** was categorized according to 3 mRNAs from the original study by Hongbo Zhou. In the classification process for every subtype, each sample was consistently clustered according to mRNAs in every classifier, followed by semiautomatic subtype assignment. Gene sequences were identified from the GRCh37/hg19 human genome assembly. To identify oncogenic events among diverse HNSCC metabolic subtypes at the molecular level, the frequencies of SNVs, CNVs and INDELs were detected from frequently mutated HNSCC genes [32], and their associations with diverse HNSCC metabolic subtypes were also explored. For tumor ploidy, we defined DNA fragments with copy statuses ≥ 3 as amplified, whereas those ≤ 1 were defined as deleted [33]. Additionally, HNSCC copy number events were selected according to prior work using 10 or more supporting probes at the threshold of mean fragment > 0.2 (amplified) or < -0.2 (deleted). Afterward, the copy number event coordinates were mapped into the gene coding region using maftools, whereas contingency analysis was applied to test CNVs and SNVs for every gene. Finally, we tested those genes screened from every subgroup. The RNA-seq data [RNA-seq by expectation maximization (RSEM); GRCh37] of each TCGA-derived non-HNSCC sample were obtained using the GDC data portal. Then, samples of different cancer types that had 100 or more samples were screened, and 262 cancer types were obtained. The expression levels were subjected to log transformation (log10(RSEMþ1)), and then genewise location scaling was used for batch correction in every cancer type. Typically, consensus clustering (ConsensusClusterPlus, parameters: pFeature = 1, pItem = 0.8, reps = 100; ED and Ward. D2, k = 4) was repeated for every individual cancer type based on gene expression in the “REACTOME CHOLESTEROL BIOSYNTHESIS” and “REACTOME GLYCOLYSIS” gene sets. Moreover, we determined the percentages of cholesterogenic and glycolytic genes in every cluster, and clusters consisting of at least 50% of each gene set were identified as the “core” clusters. With regard to cancer types that had over one core cluster in one gene set, we chose the most homogenous cluster as the core cluster. Cancer types without 75% or higher homogeneity in the core cholesterogenic and glycolytic clusters were eliminated from subsequent analysis, giving rise to 12 cancer types. Meanwhile, for every cancer type, we further determined its metabolic subtypes according to the median expression of representative core cholesterogenic and glycolytic genes. For this study the ‘‘survminer’’ v.0.4.2 and ‘‘survival’’ v.2.4.2 R packages were employed to generate Kaplan–Meier plots. Cases whose overall survival (OS) was shorter than 1 month were eliminated from survival analysis. A total of 22 tissue samples were obtained from patients with HNSCC, who underwent surgical resection at ** for novel personalized therapies against pancreatic cancer. Clin Cancer Res. 2020;26(1):6–8." href="/article/10.1186/s12935-023-02880-3#ref-CR37" id="ref-link-section-d194387745e1078">37]. According to the prognosis models of Zhang [30] and Zhou [31], we determined the risk score of HNSCC prognosis and investigated their overlap with the metabolic subtype in TCGA HNSCC datasets. The quiescent subtype contained the highest frequency of low-risk cases, and the cholesterogenic subtype was also mainly composed of low-risk samples. In contrast, glycolysis and the mixed subtype predominantly consisted of the high-risk prognosis group (Fig. 7A, B). Relationships between metabolic subtypes and known HNSCC subtypes. A, B Bar plots illustrating the proportion of published HNSCC expression subtypes across each metabolic subgroup based on patient prognosis. C Factor analysis of four HNSCC subtypes based on mRNA-based subty**. D Factor analysis of four HNSCC subtypes based on miRNA-based subty**. E Sankey diagram showing overlay of the metabolic profiles with HNSCC expression subtypes based on mRNA-based subty** by Chung and miRNA-based subty** by Walter, as well as patient survival Moreover, Chung and Walter et al. identified molecular subtypes of HNSCC based on gene expression, termed basal, mesenchymal, classical and atypical [38,39,41]. However, to date, no research has defined the metabolic classification of HNSCC. Here, we successfully established four distinct subtypes of HNSCC, the quiescent, glycolytic, cholesterogenic and mixed subtypes, which affect tumor progression and patient survival. Furthermore, we found that the metabolic subtype of HNSCC was linked to clinicopathological features and, in particular, to clinical stage and grade. The quiescent subtype was mainly enriched in stage I and G1, suggesting a better outcome. Previous studies have shown that lactate levels are not correlated with presenting T stage or N stage [42]. In accordance with previous studies, our results showed that T stage and N stage were not significantly different in the distribution of the metabolic subgroups. Our study demonstrated that metabolomic profiling could be potentially useful for prognosis. Glycolytic metabolism is a common event in tumorigenesis, as indicated by the dramatic increase in glucose uptake [22]. The finding that tumors with increased glycolytic properties were related to the shortest overall survival confirms the role of glycolysis in tumor aggressiveness in HNSCC [43]. Chen et al. found that six glycolysis-based genes were identified and can be used as prognostic markers for patients with HNSCC. In addition, lipid metabolic reprogramming is one hallmark of cancer. Cholesterol plays a key role in pathways governing carcinogenesis and malignant progression. High expression levels of cholesterogenic genes were associated with human HNSCC development and supported poor prognosis [44,45,46,47]. Emerging evidence supports these observations. ENO-1 acts as a glycolytic enzyme and promotes invasion and metastasis formation in various cancers [48,49,50]. PFKFB3 is an essential glycolysis-activating enzyme, and its powerful kinase activity can increase glycolysis flux and was involved in the aggressive features of multiple malignances and correlates with poor survival [51,52, The original contributions presented in the study are included in the article/Supplementary Material.The datasets presented in this study can be found in online repositories. Johnson DE, Burtness B, Leemans CR, Lui VWY, Bauman JE, Grandis JR. Head and neck squamous cell carcinoma. Nat Rev Dis Primers. 2020;6(1):92. Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, Bray F. Global cancer statistics 2020: globocan estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2021;71(3):209. World Health Organization. Global Cancer Observatory: Cancer Tomorrow. 2020. Available online at:
https://gco.iarc.fr/tomorrow/en/dataviz/isotype. Accessed 25 Mar 2021. Erkal H, Mendenhall W, Amdur R, Villaret D, Stringer S. Synchronous and metachronous squamous cell carcinomas of the head and neck mucosal sites. J Clin Oncol Off Journal Am Soc Clin Oncol. 2001;19(5):1358–62. Domingo-Vidal M, Whitaker-Menezes D, Martos-Rus C, Tassone P, Snyder CM, Tuluc M, Philp N, Curry J, Martinez-Outschoorn U. Cigarette smoke induces metabolic reprogramming of the tumor stroma in head and neck squamous cell carcinoma. Mol Cancer Res. 2019;17(9):1893–909. Martinez-Outschoorn UE, Peiris-Pages M, Pestell RG, Sotgia F, Lisanti MP. Cancer metabolism: a therapeutic perspective. Nat Rev Clin Oncol. 2017;14(1):11–31. Pavlova NN, Thompson CB. The emerging hallmarks of cancer metabolism. Cell Metab. 2016;23(1):27–47. Faubert B, Solmonson A, DeBerardinis RJ. Metabolic reprogramming and cancer progression. Science. 2020;368(6487):5473. Matsushita Y, Nakagawa H, Koike K. Lipid metabolism in oncology: why it matters, how to research, and how to treat. Cancers. 2021;13(3):474. Faubert B, Vincent EE, Griss T, Samborska B, Izreig S, Svensson RU, Mamer OA, Avizonis D, Shackelford DB, Shaw RJ, et al. Loss of the tumor suppressor LKB1 promotes metabolic reprogramming of cancer cells via HIF-1alpha. Proc Natl Acad Sci USA. 2014;111(7):2554–9. Svensson RU, Parker SJ, Eichner LJ, Kolar MJ, Wallace M, Brun SN, Lombardo PS, Van Nostrand JL, Hutchins A, Vera L, et al. Inhibition of acetyl-CoA carboxylase suppresses fatty acid synthesis and tumor growth of non-small-cell lung cancer in preclinical models. Nat Med. 2016;22(10):1108–19. Vander Heiden M, Cantley L, Thompson C. Understanding the Warburg effect: the metabolic requirements of cell proliferation. Science. 2009;324(5930):1029–33. Park G, Kim J, Roh J, Choi S, Nam S, Kim S. Prognostic value of metabolic tumor volume measured by 18F-FDG PET/CT in advanced-stage squamous cell carcinoma of the larynx and hypopharynx. Ann Oncol Off J European Soc Med Oncol. 2013;24(1):208–14. Paidpally V, Chirindel A, Lam S, Agrawal N, Quon H, Subramaniam R. FDG-PET/CT imaging biomarkers in head and neck squamous cell carcinoma. Imaging in medicine. 2012;4(6):633–47. Bonomo P, Merlotti A, Olmetto E, Bianchi A, Desideri I, Bacigalupo A, Franco P, Franzese C, Orlandi E, Livi L, et al. What is the prognostic impact of FDG PET in locally advanced head and neck squamous cell carcinoma treated with concomitant chemo-radiotherapy? A systematic review and meta-analysis. Eur J Nucl Med Mol Imaging. 2018;45(12):2122–38. Li Z, Zhang H. Reprogramming of glucose, fatty acid and amino acid metabolism for cancer progression. Cell Mol Life Sci: CMLS. 2016;73(2):377–92. Voisin M, de Medina P, Mallinger A, Dalenc F, Huc-Claustre E, Leignadier J, Serhan N, Soules R, Ségala G, Mougel A, et al. Identification of a tumor-promoter cholesterol metabolite in human breast cancers acting through the glucocorticoid receptor. Proc Natl Acad Sci USA. 2017;114(44):E9346–55. Xu X, Chen J, Li Y, Yang X, Wang Q, Wen Y, Yan M, Zhang J, Xu Q, Wei Y, et al. Targeting epigenetic modulation of cholesterol synthesis as a therapeutic strategy for head and neck squamous cell carcinoma. Cell Death Dis. 2021;12(5):482. Chen X, Song Q, **a L, Xu X. Synergy of dendritic cell vaccines and avasimibe in treatment of head and neck cancer in mice. Med Sci Monitor Int Med J Exp Clin Res. 2017;23:4471–6. Nakajima E, Laymon C, Oborski M, Hou W, Wang L, Grandis J, Ferris R, Mountz J, Van Houten B. Quantifying metabolic heterogeneity in head and neck tumors in real time: 2-DG uptake is highest in hypoxic tumor regions. PLoS ONE. 2014;9(8): e102452. Karasinska J, Topham J, Kalloger S, Jang G, Denroche R, Culibrk L, Williamson L, Wong H, Lee M, O’Kane G, et al. Altered gene expression along the glycolysis-cholesterol synthesis axis is associated with outcome in pancreatic cancer. Clin Cancer Res Off Journal Am Assoc Cancer Res. 2020;26(1):135–46. Sandulache V, Ow T, Pickering C, Frederick M, Zhou G, Fokt I, Davis-Malesevich M, Priebe W, Myers J. Glucose, not glutamine, is the dominant energy source required for proliferation and survival of head and neck squamous carcinoma cells. Cancer. 2011;117(13):2926–38. Rocken C. Molecular classification of gastric cancer. Expert Rev Mol Diagn. 2017;17(3):293–301. Rodriguez-Canales J, Parra-Cuentas E, Wistuba II. Diagnosis and molecular classification of lung cancer. Cancer Treat Res. 2016;170:25–46. Provenzano E, Ulaner G, Chin S. Molecular classification of breast cancer. PET clinics. 2018;13(3):325–38. Lohavanichbutr P, Méndez E, Holsinger F, Rue T, Zhang Y, Houck J, Upton M, Futran N, Schwartz S, Wang P, et al. A 13-gene signature prognostic of HPV-negative OSCC: discovery and external validation. Clin Cancer Res Off Journal Am Assoc Cancer Res. 2013;19(5):1197–203. Karasinska JM, Topham JT, Kalloger SE, Jang GH, Denroche RE, Culibrk L, Williamson LM, Wong HL, Lee MKC, O’Kane GM, et al. Altered gene expression along the glycolysis-cholesterol synthesis axis is associated with outcome in pancreatic cancer. Clin Cancer Res. 2020;26(1):135–46. Liberzon A, Subramanian A, Pinchback R, Thorvaldsdóttir H, Tamayo P, Mesirov J. Molecular signatures database (MSigDB) 3 0. Bioinformatics. 2011;27(12):1739–40. Wilkerson M, Hayes D. Consensus cluster plus: a class discovery tool with confidence assessments and item tracking. Bioinformatics. 2010;26(12):1572–3. Tian S, Meng G, Zhang W. A six-mRNA prognostic model to predict survival in head and neck squamous cell carcinoma. Cancer Manag Res. 2019;11:131–42. Cao R, Wu Q, Li Q, Yao M, Zhou H. A 3-mRNA-based prognostic signature of survival in oral squamous cell carcinoma. PeerJ. 2019;7: e7360. Duitama J, Quintero J, Cruz D, Quintero C, Hubmann G, Foulquié-Moreno M, Verstrepen K, Thevelein J, Tohme J. An integrated framework for discovery and genoty** of genomic variants from high-throughput sequencing experiments. Nucleic Acids Res. 2014;42(6): e44. Laddha S, Ganesan S, Chan C, White E. Mutational landscape of the essential autophagy gene BECN1 in human cancers. Mol Cancer Res MCR. 2014;12(4):485–90. Remmele W, Stegner H. Recommendation for uniform definition of an immunoreactive score (IRS) for immunohistochemical estrogen receptor detection (ER-ICA) in breast cancer tissue. Pathologe. 1987;8(3):138–40. Wilkie MD, Anaam EA, Lau AS, Rubbi CP, Jones TM, Boyd MT, Vlatkovic N. TP53 mutations in head and neck cancer cells determine the Warburg phenotypic switch creating metabolic vulnerabilities and therapeutic opportunities for stratified therapies. Cancer Lett. 2020;478:107–21. Shi S, Ji S, Qin Y, Xu J, Zhang B, Xu W, Liu J, Long J, Liu C, Liu L, et al. Metabolic tumor burden is associated with major oncogenomic alterations and serum tumor markers in patients with resected pancreatic cancer. Cancer Lett. 2015;360(2):227–33. Mehla K, Singh PK. Metabolic subty** for novel personalized therapies against pancreatic cancer. Clin Cancer Res. 2020;26(1):6–8. Walter V, Yin X, Wilkerson MD, Cabanski CR, Zhao N, Du Y, Ang MK, Hayward MC, Salazar AH, Hoadley KA, et al. Molecular subtypes in head and neck cancer exhibit distinct patterns of chromosomal gain and loss of canonical cancer genes. PLoS ONE. 2013;8(2): e56823. Cancer Genome Atlas N. Comprehensive genomic characterization of head and neck squamous cell carcinomas. Nature. 2015;517(7536):576–82. Chung C, Parker J, Karaca G, Wu J, Funkhouser W, Moore D, Butterfoss D, **ang D, Zanation A, Yin X, et al. Molecular classification of head and neck squamous cell carcinomas using patterns of gene expression. Cancer Cell. 2004;5(5):489–500. Gong Y, Ji P, Yang YS, ** of triple-negative breast cancer reveals potential therapeutic targets. Cell Metab. 2021;33(1):51–64. Brizel D, Schroeder T, Scher R, Walenta S, Clough R, Dewhirst M, Mueller-Klieser W. Elevated tumor lactate concentrations predict for an increased risk of metastases in head-and-neck cancer. Int J Radiat Oncol Biol Phys. 2001;51(2):349–53. Kumar D, New J, Vishwakarma V, Joshi R, Enders J, Lin F, Dasari S, Gutierrez WR, Leef G, Ponnurangam S, et al. Cancer-associated fibroblasts drive glycolysis in a targetable signaling loop implicated in head and neck squamous cell carcinoma progression. Cancer Res. 2018;78(14):3769–82. Lang L, Loveless R, Dou J, Lam T, Chen A, Wang F, Sun L, Juarez J, Qin Z, Saba N, et al. ATAD3A mediates activation of RAS-independent mitochondrial ERK1/2 signaling, favoring head and neck cancer development. J Exp Clin Cancer Res CR. 2022;41(1):43. Dickinson A, Saraswat M, Joenväärä S, Agarwal R, Jyllikoski D, Wilkman T, Mäkitie A, Silén S. Mass spectrometry-based lipidomics of oral squamous cell carcinoma tissue reveals aberrant cholesterol and glycerophospholipid metabolism —a pilot study. Transl Oncol. 2020;13(10): 100807. Liu Y, Fang L, Liu W. High SQLE expression and gene amplification correlates with poor prognosis in head and neck squamous cell carcinoma. Cancer Manage Res. 2021;13:4709–23. Zhao R, Tian L, Zhao B, Sun Y, Cao J, Chen K, Li F, Li M, Shang D, Liu M. FADS1 promotes the progression of laryngeal squamous cell carcinoma through activating AKT/mTOR signaling. Cell Death Dis. 2020;11(4):272. Principe M, Borgoni S, Cascione M, Chattaragada M, Ferri-Borgogno S, Capello M, Bulfamante S, Chapelle J, Di Modugno F, Defilippi P, et al. Alpha-enolase (ENO1) controls alpha v/beta 3 integrin expression and regulates pancreatic cancer adhesion, invasion, and metastasis. J Hematol Oncol. 2017;10(1):16. Fu Q, Liu Y, Fan Y, Hua S, Qu H, Dong S, Li R, Zhao M, Zhen Y, Yu X, et al. Alpha-enolase promotes cell glycolysis, growth, migration, and invasion in non-small cell lung cancer through FAK-mediated PI3K/AKT pathway. J Hematol Oncol. 2015;8:22. Song Y, Luo Q, Long H, Hu Z, Que T, Zhang X, Li Z, Wang G, Yi L, Liu Z, et al. Alpha-enolase as a potential cancer prognostic marker promotes cell growth, migration, and invasion in glioma. Mol Cancer. 2014;13:65. Gao W, Zhang Y, Luo H, Niu M, Zheng X, Hu W, Cui J, Xue X, Bo Y, Dai F, et al. Targeting SKA3 suppresses the proliferation and chemoresistance of laryngeal squamous cell carcinoma via impairing PLK1-AKT axis-mediated glycolysis. Cell Death Dis. 2020;11(10):919. O’Neal J, Clem A, Reynolds L, Dougherty S, Imbert-Fernandez Y, Telang S, Chesney J, Clem B. Inhibition of 6-phosphofructo-2-kinase (PFKFB3) suppresses glucose metabolism and the growth of HER2+ breast cancer. Breast Cancer Res Treat. 2016;160(1):29–40. Shi L, Pan H, Liu Z, **e J, Han W. Roles of PFKFB3 in cancer. Signal Transduct Target Ther. 2017;2:17044. Kalogirou C, Linxweiler J, Schmucker P, Snaebjornsson M, Schmitz W, Wach S, Krebs M, Hartmann E, Puhr M, Müller A, et al. MiR-205-driven downregulation of cholesterol biosynthesis through SQLE-inhibition identifies therapeutic vulnerability in aggressive prostate cancer. Nat Commun. 2021;12(1):5066. Li C, Wang Y, Liu D, Wong C, Coker O, Zhang X, Liu C, Zhou Y, Liu Y, Kang W, et al. Squalene epoxidase drives cancer cell proliferation and promotes gut dysbiosis to accelerate colorectal carcinogenesis. Gut. 2022. https://doi.org/10.1136/gutjnl-2021-325851. Gabitova L, Restifo D, Gorin A, Manocha K, Handorf E, Yang D, Cai K, Klein-Szanto A, Cunningham D, Kratz L, et al. Endogenous sterol metabolites regulate growth of EGFR/KRAS-dependent tumors via LXR. Cell Rep. 2015;12(11):1927–38. DeBerardinis R, Chandel N. Fundamentals of cancer metabolism. Sci Adv. 2016;2(5): e1600200. Baschnagel AM, Wobb JL, Dilworth JT, Williams L, Eskandari M, Wu D, Pruetz BL, Wilson GD. The association of (18)F-FDG PET and glucose metabolism biomarkers GLUT1 and HK2 in p16 positive and negative head and neck squamous cell carcinomas. Radiother Oncol. 2015;117(1):118–24. Ju HQ, Ying H, Tian T, Ling J, Fu J, Lu Y, Wu M, Yang L, Achreja A, Chen G, et al. Mutant Kras- and p16-regulated NOX4 activation overcomes metabolic checkpoints in development of pancreatic ductal adenocarcinoma. Nat Commun. 2017;8:14437. Ahn C, Metallo C. Mitochondria as biosynthetic factories for cancer proliferation. Cancer Metab. 2015;3(1):1. Eriksson M, Ambroise G, Ouchida A, Lima Queiroz A, Smith D, Gimenez-Cassina A, Iwanicki M, Muller P, Norberg E, Vakifahmetoglu-Norberg H. Effect of mutant p53 proteins on glycolysis and mitochondrial metabolism. Mol Cell Biol. 2017. https://doi.org/10.1128/MCB.00328-17. Broadfield L, Pane A, Talebi A, Swinnen J, Fendt S. Lipid metabolism in cancer: new perspectives and emerging mechanisms. Dev Cell. 2021;56(10):1363–93. We thank TCGA project and NCBI GEO for providing the RNA-sequencing data and clinical data of patients with HNSCC. This work was supported by the grant from Key Research and Development Program of Hunan Province (No. 2020SK2056) and Natural Science Foundation of Hunan Province of China (No. S2021JJMSXM2780) and the Fundamental Research Funds for the Central Universities of Central South University (No. 2021zzts0947). HZ and XD; Investigation, ZZ; visualization, ZZ and JT; Supervision, OL; Validation, XD; Funding acquisition, HZ; Project administration, OL and HZ; Software and Methodology, KS and YL; Resources, JJ and TL; Writing–Original Draft Preparation, ZZ and JT; Writing–Review and Editing, ZZ, OL and XD All authors have read and agreed to the published version of the manuscript. All authors read and approved the final manuscript. The study was conducted in accordance with the Declaration of Helsinki, and approved by the Ethics Committee of **angya Stomatological Hospital of Central South University (20190067). Written informed consent for publication was obtained from all participants. The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest. Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations. The consensus clustering of HNSCC samples classification. A–D The color‐coded heatmap corresponding to the consensus matrix for k = 2,3,4,5 obtained by applying consensus clustering. The color gradients were from 0 to 1, representing the degree of consensus, with white corresponding to 0 and dark blue to 1. E–F Delta area curve of consensus clustering, indicating the relative change in area under the cumulative distribution function (CDF) curve for 2 each category number k compared with k–1. The horizontal axis represents the category number k and the vertical axis represents the relative change in area under CDF curve. Figure S2. This maftools plot showing most mutated genes, SNV class, and variant classification distributions in HNSCC. Table S1. Clinicopathological characteristics of patients with head and neck squamous cell carcinoma (N = 22). Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data. Zhou, Z., Tang, J., Lu, Y. et al. Prognosis-related molecular subty** in head and neck squamous cell carcinoma patients based on glycolytic/cholesterogenic gene data.
Cancer Cell Int 23, 37 (2023). https://doi.org/10.1186/s12935-023-02880-3 Received: Accepted: Published: DOI: https://doi.org/10.1186/s12935-023-02880-3Materials and methods
HNSCC dataset acquisition and processing
RNA sequencing data analysis
Metabolic gene subgroup classification
Pre-existing HNSCC subgroup classification
Mutation analysis for HNSCC genes
Pan-TCGA RNA-seq analysis
Survival of HNSCC cases
Patient specimens and Immunohistochemistry (IHC)
Availability of data and materials
References
Acknowledgements
Funding
Author information
Authors and Affiliations
Contributions
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
Consent for publication
Competing interests
Additional information
Publisher's Note
Supplementary Information
Additional file 1: Figure S1.
Rights and permissions
About this article
Cite this article
Keywords