Abstract
Background
Lung cancer (LC) is one of the most devastating diseases worldwide, there is growing studies confirm the role of impaired lung function in LC susceptibility. Moreover, gut microbiota dysbiosis is associated with LC severity. Whether alterations in gut microbiota and metabolites are associated with long-term lung dysfunction in LC patients remain unclear. Our study aimed to analyze the risk factors in LC patients with impaired pulmonary function based on the characteristics of the gut microbiome and metabolites.
Methods
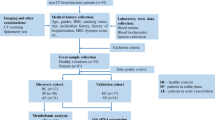
Fecal samples from 55 LC patients and 28 benign pulmonary nodules patients were collected. Pulmonary ventilation function was graded according to the American Thoracic Society/ European Respiratory Society (ATS/ERS) method. LC patients were divided into 3 groups, including 20 patients with normal lung ventilation, 23 patients with mild pulmonary ventilation dysfunction and 12 patients with moderate or above pulmonary ventilation dysfunction. The fecal samples were analyzed using 16 S rRNA gene amplicon sequencing and metabolomics.
Results
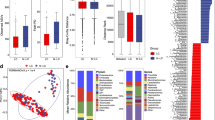
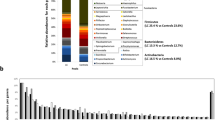
The gut microbiome composition between LC patients and benign pulmonary nodules patients presented clearly differences based on Partial Least Squares Discriminant Analysis (PLS-DA). Pulmonary ventilation function was positively correlated with LC tumor stage, the richness and diversity of the gut microbiota in LC patients with moderate or above pulmonary ventilation dysfunction increased significantly, characterized by increased abundance of Subdoligranulum and Romboutsia. The metabolomics analysis revealed 69 differential metabolites, which were mainly enriched in beta-Alanine metabolism, styrene degradation and pyrimidine metabolism pathway. The area under the curve (AUC) combining the gut microbiome and metabolites was 90% (95% CI: 79-100%), indicating that the two species and four metabolites might regarded as biomarkers to assess the prediction of LC patients with impaired pulmonary function.
Conclusions
Our results showed that microbiome and metabolomics analyses provide important candidate to be used as clinically diagnostic biomarkers and therapeutic targets related to lung cancer with impaired pulmonary function.
Similar content being viewed by others
Introduction
Lung cancer ( LC ) is one of the most common and the leading cause of cancer deaths worldwide [1]. Genetic susceptibility, gut microbiome and smoking are hypothesized to increase the risks of LC by sha** the tumor microenvironment and promoting the tumorigenesis [2]. Gut microbiota can influence the immune status of the host, which in turn increases susceptibility to malignancy. The gut microbiota and metabolites can enter the blood through the intestinal barrier, leading to a chronic inflammatory state in the organism [3]. Dysfunction of gut microbiota is believed to be associated with the occurrence and development of cancers, and studies have identified potential fecal biomarkers, satisfactory performances of these markers have been shown for diagnosing pancreatic cancer (AUC = 0.78–0.94) [4, 5], hepatocellular carcinoma (AUC = 0.8064) [6], lung adenocarcinoma (AUC = 0.76–0.976) [7, 8] and so on. Emerging studies have indicated that significant changes in the composition and function of the gut microbiota in patients with pulmonary diseases compared to healthy individuals [7, 9, 10]. The detection of differences in gut microbial communities between healthy individuals and LC patients could be used as a predictive tool for LC progression [11, 12]. Research on the role and mechanisms of intestinal flora and its metabolites in LC is beginning to receive widespread attention.
In recent years, many studies focus on the relationship between lung function and lung health. Kachuri et al. found that immune-mediated genetic pathways led to impaired lung function, with reduced FEV1 increasing the risk of squamous cell carcinoma and reduced FEV1/FVC increasing the risk of adenocarcinoma [13]. A large observational literature has found an increased risk of LC in patients with pulmonary insufficiency [14], but the possible relationship between LC and respiratory dysfunction has not been established. Li et al. revealed that the diversity of pulmonary microbiota in chronic obstructive pulmonary disease (COPD) patients with impaired lung function was similar [35, 36]. Mendelian randomization analysis revealed histological-specific effects of reduced FEV1 and FEV1/FVC on LC susceptibility, suggesting that these indicators of impaired lung function may be pathogenic risk factors [13]. In our study, pulmonary ventilation function is positively correlated with tumor stage. Reduced FEV1 has been shown to increase the risk of squamous cell carcinoma and reduced the ratios of FEV1 to FVC increase the risk of adenocarcinoma and lung cancer in never smokers [13]. Gut microbiota is believed to play an important role in altering lung function [37]. However, no studies have investigated whether alterations in gut microbiota and metabolites are associated with long-term lung dysfunction in LC patients. Our study showed that ZC group and QD group had a similar α-diversity, but the α-diversity of ZZD group was significantly reduced (P < 0.05). In addition, these groups could be significantly separated from each other based on PLS-DA. Of which, Hungatella was significantly decreased in the ZZD group, suggesting a negative correlation with the aggravation of tumor stage in LC patients. However, Subdoligranulum and Romboutsia were significantly elevated in ZZD group. Chriswell et al. suggested that subdoligranulum stimulated joint swelling and inflammation [38]. Ni et al. demonstrated that subdoligranulum is the dominant biomarker that distinguish vitiligo patients from healthy controls [39]. In contrast, Lloyd-Price at al. showed that subdoligranulum was markedly increased in inflammatory bowel diseases (IBD), considering as a butyrate producer [40]. Due to the variability of findings, therefore further research is needed to demonstrate the role of Subdoliganulum. It has been reported that invasive mechanical ventilation leads to lung microbiota changes in rat models, which mainly characterization by the Romboutsia and Tubriciactor genera [41]. We speculate that the alterations of genera may be related to the patient dietary habits and a history of nasal oxygen therapy. So the role of these genera in LC patients with pulmonary dysfunction needs to be further investigated in a larger sample cohort.
Then, in the metabolomics analysis of fecal samples, bile acids were significantly decreased in ZZD group. Bile acid biosynthesis changed is a collaborative effect between the host and gut microbiome [42]. The reduction of secondary bile acids may alter the composition of gut microbiota and promote an intestinal inflammation profile [43]. Nie et al. revealed that bile acid metabolism is related to poor prognosis, and may potentiate migration of lung adenocarcinoma (LUAD) [44]. TGR5, the bile acid receptor, as a negative regulator of the NF-κB and AKT pathway, may effectively inhibit the progression of non-small cell lung cancer (NSCLC) [45].
The ROC curve analysis identified four potential biomarkers of diagnostic significance, Stearoylethanolamide, Serylthreonine, Xestoaminol C and Farnesyl acetone with AUC of 0.8750, 0.8625, 0.8583 and 0.8583, respectively. Stearoylethanolamide, an endogenous cannabinoid-like compound with pro-apoptotic activity, is found in the human brain to support the blood-brain barrier in acute systemic inflammation [46, 47]. Terpenoids related compounds exhibit high antibacterial activity against gram-negative bacteria and the very high cytotoxic activity profile of farnesol gives it potential as an anticancer agent [48]. ROC curve analysis between the gut microbes and metabolites showed that the diagnostic power was significantly increased (AUC = 0.9). Previous studies revealed that combining of the bacteria and the clinical tumor markers showed a higher ROC value for predicting LC [49]. Besides, the diagnostic power for predicting colorectal cancer was significantly increased based on the combination of metabolites and bacterial markers [50]. These studies suggesting that joint multi-dimensional data could be used to better predict disease risks. In many cases, gut flora structure has been correlated with the severity of respiratory diseases, such as NSCLC, COVID-19 and COPD [51,52,53].
In conclusion, the intestinal microecology of patients with LC and benign pulmonary diseases patients were characterized in this study. It is revealed that impaired lung ventilation may influence disease severity in LC patients and a predictive model could be developed based on the combination of gut microbes and metabolites for assessing the severity of lung cancer. In the future, a larger sample of the validation queue is needed to validate that alterations in gut microbiota and metabolites are associated with long-term lung dysfunction in LC patients.
Conclusions
Our results showed that microbiome and metabolomics analyses provide important candidate to be used as clinically diagnostic biomarkers and therapeutic targets related to lung cancer with impaired pulmonary function.
Data availability
The datasets presented in this study can be found in online repositories. The names of the repository/repositories and accession number(s) can be found below: https://www.ncbi.nlm.nih.gov/, PRJNA956658.
References
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, et al. Global Cancer statistics 2020: GLOBOCAN estimates of incidence and Mortality Worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2020;71(3):209–49.
Zhai R, Yu X, Wei Y, Su L, Christiani DC. Smoking and Smoking cessation in relation to the development of co-existing non-small cell Lung cancer with Chronic Obstructive Pulmonary Disease. Int J Cancer. 2014;134(4):961–70.
Fellows R, Denizot J, Stellato C, Cuomo A, Jain P, Stoyanova E, et al. Microbiota derived short chain fatty acids promote histone crotonylation in the colon through histone deacetylases. Nat Commun. 2018;9(1):105.
Nagata N, Nishijima S, Kojima Y, Hisada Y, Imbe K, Miyoshi-Akiyama T, et al. Metagenomic identification of Microbial signatures Predicting Pancreatic Cancer from a multinational study. Gastroenterology. 2022;163(1):222–38.
Kartal E, Schmidt TSB, Molina-Montes E, Rodríguez-Perales S, Wirbel J, Maistrenko OM, et al. A faecal microbiota signature with high specificity for Pancreatic cancer. Gut. 2022;71(7):1359–72.
Ren Z, Li A, Jiang J, Zhou L, Yu Z, Lu H, et al. Gut microbiome analysis as a tool towards targeted non-invasive biomarkers for early hepatocellular carcinoma. Gut. 2019;68(6):1014–23.
Lim MY, Hong S, Hwang KH, Lim EJ, Han JY, Nam YD. Diagnostic and prognostic potential of the oral and gut microbiome for lung adenocarcinoma. Clin Transl Med. 2021;11(9):e508.
Zheng Y, Fang Z, Xue Y, Zhang J, Zhu J, Gao R, et al. Specific gut microbiome signature predicts the early-stage Lung cancer. Gut Microbes. 2020;11(4):1030–42.
Stokholm J, Blaser MJ, Thorsen J, Rasmussen MA, Waage J, Vinding RK, et al. Maturation of the gut microbiome and risk of Asthma in childhood. Nat Commun. 2018;9(1):141.
Hu Y, Feng Y, Wu J, Liu F, Zhang Z, Hao Y, et al. The gut Microbiome signatures discriminate healthy from pulmonary Tuberculosis patients. Front Cell Infect Microbiol. 2019;9:90.
Zhuang H, Cheng L, Wang Y, Zhang YK, Zhao MF, Liang GD, et al. Dysbiosis of the gut microbiome in Lung Cancer. Front Cell Infect Microbiol. 2019;9:112.
Liu F, Li J, Guan Y, Lou Y, Chen H, Xu M, et al. Dysbiosis of the gut microbiome is associated with Tumor biomarkers in Lung Cancer. Int J Biol Sci. 2019;15(11):2381–92.
Kachuri L, Johansson M, Rashkin SR, Graff RE, Bossé Y, Manem V, et al. Immune-mediated genetic pathways resulting in pulmonary function impairment increase Lung cancer susceptibility. Nat Commun. 2020;11(1):27.
Dong Y, Kumar H, Tawhai M, Veiga C, Szmul A, Landau D, et al. In Silico Ventilation within the dose-volume is predictive of lung function post-radiation therapy in patients with Lung Cancer. Ann Biomed Eng. 2021;49(5):1416–31.
Li W, Wang B, Tan M, Song X, **e S, Wang C. Analysis of sputum microbial metagenome in COPD based on exacerbation frequency and lung function: a case control study. Respir Res. 2022;23(1):321.
Yan Z, Chen B, Yang Y, Yi X, Wei M, Ecklu-Mensah G, et al. Multi-omics analyses of airway host-microbe interactions in Chronic Obstructive Pulmonary Disease identify potential therapeutic interventions. Nat Microbiol. 2022;7(9):1361–75.
Liu X, Cheng Y, Zang D, Zhang M, Li X, Liu D, et al. The role of Gut Microbiota in Lung Cancer: from carcinogenesis to Immunotherapy. Front Oncol. 2021;11:720842.
Miller MR, Hankinson J, Brusasco V, Burgos F, Casaburi R, Coates A, et al. Standardisation of spirometry. Eur Respir J. 2005;26(2):319–38.
Kanehisa M, Furumichi M, Sato Y, Kawashima M, Ishiguro-Watanabe M. KEGG for taxonomy-based analysis of pathways and genomes. Nucleic Acids Res. 2023;51(D1):D587–92.
Li M, Liu J, Zhu J, Wang H, Sun C, Gao NL, et al. Performance of gut microbiome as an Independent Diagnostic Tool for 20 Diseases: Cross-cohort Validation of Machine-Learning classifiers. Gut Microbes. 2023;15(1):2205386.
Wirbel J, Pyl PT, Kartal E, Zych K, Kashani A, Milanese A, et al. Meta-analysis of fecal metagenomes reveals global microbial signatures that are specific for Colorectal cancer. Nat Med. 2019;25(4):679–89.
Jiang P, Wu S, Luo Q, Zhao XM, Chen WH. Metagenomic analysis of Common Intestinal Diseases reveals relationships among Microbial signatures and Powers Multidisease Diagnostic models. mSystems. 2021;6(3):e00112–21.
Lu H, Gao NL, Tong F, Wang J, Li H, Zhang R, et al. Alterations of the human lung and gut microbiomes in Non-small Cell Lung carcinomas and distant Metastasis. Microbiol Spectr. 2021 Dec;9(3):e0080221.
Qin N, Yang F, Li A, Prifti E, Chen Y, Shao L, et al. Alterations of the human gut microbiome in liver Cirrhosis. Nature. 2014;513(7516):59–64.
Zhang WQ, Zhao SK, Luo JW, Dong XP, Hao YT, Li H, et al. Alterations of fecal bacterial communities in patients with Lung cancer. Am J Transl Res. 2018;10(10):3171–85.
Liu X, Mao B, Gu J, Wu J, Cui S, Wang G, et al. Blautia-a new functional genus with potential probiotic properties? Gut Microbes. 2021;13(1):1–21.
Benítez-Páez A, Gómez D, Pugar EM, López-Almela I, Moya-Pérez Á, Codoñer-Franch P, Sanz Y. Depletion of Blautia Species in the microbiota of obese children relates to intestinal inflammation and metabolic phenotype worsening. mSystems. 2020;5(2):e00857–19.
Tyler AD, Knox N, Kabakchiev B, Milgrom R, Kirsch R, Cohen Z, et al. Characterization of the gut-associated microbiome in inflammatory pouch Complications following ileal pouch-anal anastomosis. PLoS ONE. 2013;8(9):e66934.
Kakiyama G, Pandak WM, Gillevet PM, Hylemon PB, Heuman DM, Daita K, et al. Modulation of the fecal bile acid profile by gut microbiota in Cirrhosis. J Hepatol. 2013;58(5):949–55.
Hosomi K, Saito M, Park J, Murakami H, Shibata N, Ando M, et al. Oral administration of Blautia wexlerae ameliorates obesity and type 2 Diabetes via metabolic remodeling of the gut microbiota. Nat Commun. 2022;13(1):4477.
Zhao F, An R, Wang L, Shan J, Wang X. Specific gut microbiome and serum metabolome changes in Lung Cancer patients. Front Cell Infect Microbiol. 2021;11:725284.
Lobo LA, Jenkins AL, Jeffrey Smith C, Rocha ER. Expression of Bacteroides fragilis hemolysins in vivo and role of HlyBA in an intra-abdominal Infection model. Microbiologyopen. 2013;2(2):326–37.
Yu T, Guo F, Yu Y, Sun T, Ma D, Han J, et al. Fusobacterium nucleatum promotes Chemoresistance to Colorectal Cancer by modulating Autophagy. Cell. 2017;170(3):548–563e16.
Li HL, Lu L, Wang XS, Qin LY, Wang P, Qiu SP, et al. Alteration of Gut Microbiota and Inflammatory Cytokine/Chemokine profiles in 5-Fluorouracil Induced Intestinal Mucositis. Front Cell Infect Microbiol. 2017;7:455.
Mannino DM, Aguayo SM, Petty TL, Redd SC. Low lung function and incident Lung cancer in the United States: data from the First National Health and Nutrition Examination Survey follow-up. Arch Intern Med. 2003;163(12):1475–80.
Brenner DR, McLaughlin JR, Hung RJ. Previous lung Diseases and Lung cancer risk: a systematic review and meta-analysis. PLoS ONE. 2011;6(3):e17479.
Ver Heul A, Planer J, Kau AL. The human microbiota and Asthma. Clin Rev Allergy Immunol. 2019;57(3):350–63.
Chriswell ME, Lefferts AR, Clay MR, Hsu AR, Seifert J, Feser ML, et al. Clonal IgA and IgG autoantibodies from individuals at risk for rheumatoid arthritis identify an arthritogenic strain of Subdoligranulum. Sci Transl Med. 2022;14(668):eabn5166.
Ni Q, Ye Z, Wang Y, Chen J, Zhang W, Ma C, et al. Gut microbial dysbiosis and plasma Metabolic Profile in individuals with Vitiligo. Front Microbiol. 2020;11:592248.
Lloyd-Price J, Arze C, Ananthakrishnan AN, Schirmer M, Avila-Pacheco J, Poon TW, et al. Multi-omics of the gut microbial ecosystem in inflammatory bowel Diseases. Nature. 2019;569(7758):655–62.
Liu C, Wu K, Sun T, Chen B, Yi Y, Ren R, et al. Effect of invasive mechanical ventilation on the diversity of the pulmonary microbiota. Crit Care. 2022;26(1):252.
Winston JA, Theriot CM. Diversification of host bile acids by members of the gut microbiota. Gut Microbes. 2020;11(2):158–71.
Sinha SR, Haileselassie Y, Nguyen LP, Tropini C, Wang M, Becker LS, et al. Dysbiosis-Induced secondary bile Acid Deficiency promotes intestinal inflammation. Cell Host Microbe. 2020;27(4):659–670e5.
Nie M, Yao K, Zhu X, Chen N, **ao N, Wang Y, et al. Evolutionary metabolic landscape from preneoplasia to invasive lung adenocarcinoma. Nat Commun. 2021;12(1):6479.
Ma L, Yang F, Wu X, Mao C, Guo L, Miao T, et al. Structural basis and molecular mechanism of biased GPBAR signaling in regulating NSCLC cell growth via YAP activity. Proc Natl Acad Sci U S A. 2022;119(29):e2117054119.
Kasatkina LA, Heinemann A, Hudz YA, Thomas D, Sturm EM. Stearoylethanolamide interferes with retrograde endocannabinoid signalling and supports the blood-brain barrier integrity under acute systemic inflammation. Biochem Pharmacol. 2020;174:113783.
Maccarrone M, Pauselli R, Di Rienzo M, Finazzi-Agrò A. Binding, degradation and apoptotic activity of stearoylethanolamide in rat C6 glioma cells. Biochem J. 2002;366(Pt 1):137–44.
Bonikowski R, Świtakowska P, Sienkiewicz M, Zakłos-Szyda M. Selected compounds structurally related to Acyclic Sesquiterpenoids and their antibacterial and cytotoxic activity. Molecules. 2015;20(6):11272–96.
Cheng C, Wang Z, Wang J, Ding C, Sun C, Liu P, et al. Characterization of the lung microbiome and exploration of potential bacterial biomarkers for Lung cancer. Transl Lung Cancer Res. 2020;9(3):693–704.
Mujagic Z, Kasapi M, Jonkers DM, Garcia-Perez I, Vork L, Weerts ZZRM, et al. Integrated fecal microbiome-metabolome signatures reflect stress and serotonin metabolism in irritable bowel syndrome. Gut Microbes. 2022;14(1):2063016.
Qian X, Zhang HY, Li QL, Ma GJ, Chen Z, Ji XM, et al. Integrated microbiome, metabolome, and proteome analysis identifies a novel interplay among commensal bacteria, metabolites and candidate targets in non-small cell Lung cancer. Clin Transl Med. 2022;12(6):e947.
Bowerman KL, Rehman SF, Vaughan A, Lachner N, Budden KF, Kim RY, et al. Disease-associated gut microbiome and metabolome changes in patients with Chronic Obstructive Pulmonary Disease. Nat Commun. 2020;11(1):5886.
Guo M, Wu G, Tan Y, Li Y, ** X, Qi W, et al. Guild-Level Microbiome Signature Associated with COVID-19 severity and prognosis. mBio. 2023;6:e0351922.
Funding
This work was supported by the Taishan Scholars Program of Shandong Province (tsqn202103196).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection and analysis were performed by Jiahui Luan, Fuxin Zhang, Lijun Suo and Hongyun Cao. The first draft of the manuscript was written by Jiahui Luan, Fuxin Zhang, Lijun Suo, Bo Liu and Hongyun Cao and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
This study was approved by Ethics Committee of Zibo Municipal Hospital (Ethics No.20220311) and informed consents were obtained from all patients.
Competing interests
The authors declare no competing interests.
Consent for publication
Not applicable.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Luan, J., Zhang, F., Suo, L. et al. Analyzing lung cancer risks in patients with impaired pulmonary function through characterization of gut microbiome and metabolites. BMC Pulm Med 24, 1 (2024). https://doi.org/10.1186/s12890-023-02825-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12890-023-02825-6