Abstract
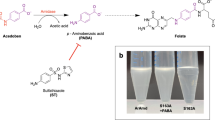
CRISPR/Cas13a nucleases are currently considered to be the basis for the development of a new generation of biosensors for the ultrasensitive, in-field detection of bacterial and viral pathogens. A recombinant Cas13a nuclease with functional affinity was obtained as a result of heterologous expression in E. coli with a single-step purification process via metal-chelating chromatography with the N-terminal polyhistidine tag. The simplified procedure of Cas13a nuclease purification broadens the possibilities for the development and practical application of diagnostic biosensing systems based on it. Moreover, our results indicate that the currently uncharacterized protein U2PWF1 of Leptotrichia wadei represents Cas13a nuclease.




Similar content being viewed by others
REFERENCES
van der Oost, J., Westra, E.R., Jackson, R.N., and Wiedenheft, B., Nat. Rev. Microbiol., 2014, vol. 12, no. 7, pp. 479–492.
Koonin, E.V., Makarova, K.S., and Zhang, F., Curr. Opin. Microbiol., 2017, vol. 37, pp. 67–78.
Hsu, P.D., Lander, E.S., and Zhang, F., Cell, 2014, vol. 157, no. 6, pp. 1262–1278.
Abudayyeh, O.O., Gootenberg, J.S., Konermann, S., Joung, J., Slaymaker, I.M., Cox, D.B., Shmakov, S., Makarova, K.S., Semenova, E., Minakhin, L., Severinov, K., Regev, A., Lander, E.S., Koonin, E.V., and Zhang, F., Science, 2016, vol. 353, no. 6299, p. aaf5573.
Pickar-Oliver, A. and Gersbach, C.A., Nat. Rev. Mol. Cell. Biol., 2019, vol. 20, no. 8, pp. 490–507.
Abudayyeh, O.O., Gootenberg, J.S., Essletzbichler, P., Han, S., Joung, J., Belanto, J.J., Verdine, V., Cox, D.B.T., Kellner, M.J., Regev, A., Lander, E.S., Voytas, D.F., Ting, A.Y., and Zhang, F., Nature, 2017, vol. 550, no. 7675, pp. 280–284.
Cox, D.B.T., Gootenberg, J.S., Abudayyeh, O.O., Franklin, B., Kellner, M.J., Joung, J., and Zhang, F., Science, 2017, vol. 358, no. 6366, pp. 1019–1027.
Shmakov, S., Abudayyeh, O.O., Makarova, K.S., Wolf, Y.I., Gootenberg, J.S., Semenova, E., Minakhin, L., Joung, J., Konermann, S., Severinov, K., Zhang, F., and Koonin, E.V., Mol. Cell, 2015, vol. 60, no. 3, pp. 385–397.
Tambe, A., East-Seletsky, A., Knott, G.J., Doudna, J.A., and O’Connell, M.R., Cell Rep., vol. 24, no. 4, pp. 1025–1036.
East-Seletsky, A., O’Connell, M.R., Knight, S.C., Burstein, D., Cate, J.H., Tjian, R., and Doudna, J.A., Nature, 2016, vol. 538, no. 7624, pp. 270–273.
Gootenberg, J.S., Abudayyeh, O.O., Lee, J.W., Essletzbichler, P., Dy, A.J., Joung, J., Verdine, V., Donghia, N., Daringer, N.M., Freije, C.A., Myhrvold, C., Bhattacharyya, R.P., Livny, J., Regev, A., Koonin, E.V., Hung, D.T., Sabeti, P.C., Collins, J.J., and Zhang, F., Science, 2017, vol. 356, no. 6336, pp. 438–442.
Gootenberg, J.S., Abudayyeh, O.O., Kellner, M.J., Joung, J., Collins, J.J., and Zhang, F., Science, 2018, vol. 360, no. 6387, pp. 439–444.
Kellner, M.J., Koob, J.G., Gootenberg, J.S., Abudayyeh, O.O., and Zhang, F., Nat. Protoc., 2019, vol. 14, no. 10, pp. 2986–3012.
Myhrvold, C., Freije, C.A., Gootenberg, J.S., Abudayyeh, O.O., Metsky, H.C., Durbin, A.F., et al., Science, 2018, vol. 360, no. 6387, pp. 444–448.
Li, Y., Li, S., Wang, J., and Liu, G., Trends Biotechnol., 2019, vol. 37, no. 7, pp. 730–743.
Shevchenko, A., Tomas, H., Havlis, J., Olsen, J.V., and Mann, M., Nat. Protoc., 2006, vol. 1, no. 6, pp. 2856–2860.
Barsnes, H. and Vaudel, M., J. Proteome Res., 2018, vol. 17, no. 7, pp. 2552–2555.
Vaudel, M., Burkhart, J.M., Zahedi, R.P., Oveland, E., Berven, F.S., Sickmann, A., Martens, L., and Barsnes, H., Nat. Biotechnol., 2015, vol. 33, no. 1, pp. 22–24.
Bjornson, R.D., Carriero, N.J., Colangelo, C., Shifman, M., Cheung, K.H., Miller, P.L., and Williams, K., Proteome Res., 2008, vol. 7, no. 1, pp. 293–299.
Kim, S. and Pevzner, P.A., Nat. Commun., 2014, vol. 5, p. 5277.
Geer, L.Y., Markey, S.P., Kowalak, J.A., Wagner, L., Xu, M., Maynard, D.M., Yang, X., Shi, W., and Bryant, S.H., J. Proteome Res., 2004, vol. 3, no. 5, pp. 958–964.
Zhu, H., Richmond, E., and Liang, C., Bioinformatics, 2018, vol. 34, no. 1, pp. 117–119.
Zuker, M., Nucleic Acids Res., 2003, vol. 31, no. 13, pp. 3406–3415.
Savinova, A.S., Koptev, E.Yu., Usachev, E.V., Tkachuk, A.P., and Gushchin, V.A., Vestn. RGMU, 2018, vol. 2, p. 21.
ACKNOWLEDGMENTS
The authors thank Yu.V Kotelevtsev (Skoltech) for the discussion of the possibilities of CRISPR detectors and I.Yu. Toropygin and V.G Zgoda (N.V. Orekhovich Institute of Biomedical Chemistry) for their assistance with mass-spectrometric analysis. In this work, we used the equipment of the Human Proteome Center for Collective Use (N.V. Orekhovich Institute of Biomedical Chemistry).
Funding
This study was supported by the Program of Fundamental Research for State Academies of Sciences for 2013–2020.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
The authors declare that they have no conflict of interest. This article does not contain any studies involving animals or human participants performed by any of the authors.
Additional information
Translated by M. Novikova
Rights and permissions
About this article
Cite this article
Kurbatov, L.K., Radko, S.P., Kravchenko, S.V. et al. Single Stage Purification of CRISPR/Cas13a Nuclease via Metal-Chelating Chromatography Following Heterologous Expression with the Preservation of Collateral Ribonuclease Activity. Appl Biochem Microbiol 56, 671–677 (2020). https://doi.org/10.1134/S0003683820060071
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0003683820060071