Abstract
Heat stress affects muscle development and meat quality in food animals; however, little is known regarding its regulatory mechanisms at the epigenetic level, such as via DNA methylation. In this study, we aimed to compare the DNA methylation profiles between control and heat-stressed pigs to identify candidate genes for skeletal muscle development and meat quality. Whole-genome bisulfite sequencing was used to investigate the genome-wide DNA methylation patterns in the longissimus dorsi muscles of the pigs. Both groups showed similar proportions of methylation at CpG sites but exhibited different proportions at non-CpG sites. A total of 57,147 differentially methylated regions were identified between the two groups, which corresponded to 1,422 differentially methylated genes. Gene ontogeny and KEGG pathway analyses indicated that these were mainly involved in energy and lipid metabolism, cellular defense and stress responses and calcium signaling pathways. This study revealed the global DNA methylation pattern of pig muscle between normal and heat stress conditions. The result of this study might contribute to a better understanding of epigenetic regulation in pig muscle development and meat quality.
Similar content being viewed by others
Introduction
As a result of global warming, heat stress has become a factor negatively affecting animal performance and health and may jeopardize the development and economy of animal husbandry1,2. Exposure to high ambient temperatures has been reported to be detrimental to pig production and meat quality3. Moreover, heat stress affects protein anabolism and catabolism as well as satellite cell proliferation, all of which play important roles in determining muscle growth and development4. In addition, heat stress activates signaling pathways associated with the regulation of skeletal muscle growth and development5,6.
Because the influence of stress on the success of the pig industry has increased, scientists have studied the specific effects of heat stress on pigs, particularly meat quality7,8,9,10. Heat stress is thought to affect gene transcription; appropriate regulation of gene transcription is crucial for maintenance of cell identity and physiological function. In preliminary studies, we identified some differentially expressed genes that were closely related to skeletal development and meat quality parameters between constant heat stress and normal temperature in pigs (Hao et al. 2015, unpublished data). However, until now, no systematic studies have evaluated the underlying molecular mechanisms mediating the development of skeletal muscle under heat stress.
DNA methylation is a well-studied epigenetic regulatory mechanism that plays a key role in the regulation of gene expression11. Recently, multiple studies have been conducted to identify genome-wide methylation profiles of economically important animals12,13,Table 1).
DNA methylation patterns
In each group, approximately 4% of all genomic C sites were methylated. Methylation in pig was found to exist in three sequence contexts: CG, CHG (where H is A, C, or T) and CHH. We observed overall genome-wide levels of 67.62% CG, 0.59% CHG and 0.58% CHH methylation in the CN group and 67.54% CG, 0.21% CHG and 0.20% CHH methylation in the HE group (Table 2). Compared with the CN group, the rate of methylated CG18 in the HE group was increased from 86.04% to 94.45%. However, the rate of mCHH was decreased 2.5-fold, from 10.70% to 4.24% (Fig. 1).
In the genome of each group, over 60% of the CpG sites were methylated, which is the primary DNA sequence context of cytosine methylation (Fig. 2). In the CN group, we detected abundant DNA methylation in CHH contexts. Although heat stress did not change the methylation level of the CpG sites in the genome, heat stress decreased the methylation level of the CHH contexts, which accounted for most of the difference between the groups.
Sequence preferences analysis for methylation
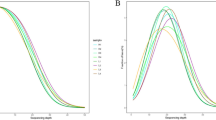
For CG context methylation, certain methylated C sites are defined as hypermethylation sites, at which the methylation level is over 75%; and some others are defined as sites of hypomethylation, at which the methylation level is less than 75%. For non-CG context methylation, hyper- and hypomethylation sites are defined as those at which the methylation levels are over or under 25%, respectively. In this study, we analyzed the relationship between sequence context and methylation preference. We calculated the percentage methylation of all possible 9-mer sequences in which the methylated cytosine was either in the fourth position (allowing an analysis of three nucleotides upstream of CG, CHG and CHH methylation) or in the first position (allowing analysis of five nucleotides following the methylated cytosine; Fig. 3). In heavily methylated regions, there was no distinct difference between the two groups in the sequence enrichment based on the genomic regions. In contrast, however, in poorly methylated regions, a high level of sequence context preferences for methylation was observed, especially in non-CG contexts.
Sequence preferences for methylation in CG, CHG and CHH contexts.
Logos of sequence contexts that are preferentially methylated at the highest or lowest levels for 9-mer sequences in which the methylated cytosine is in the fourth position. (A) Regions of high methylation. (B) Regions of low methylation.
DNA methylation levels of different functional regions
To study the global DNA methylation profile differences in the pig genome between control and constant heat stress conditions, we analyzed the DNA methylation levels of different genomic regions (Fig. 4). The comparison of average methylation levels showed that there were differential methylation levels in different components of the genome. A major proportion of methylated sites were present in the regions of promoters and introns. Among all the classes, the average methylation level of exons was the lowest. Furthermore, all five regions in the HE group exhibited lower methylation levels than did those in the CN group.
DMR analysis
To characterize the differences of genome methylation levels between samples under different environmental conditions, DMRs and differentially methylated genes (DMGs) were detected. A total of 57,147 DMRs were identified between the two groups, which corresponded to 1,422 DMGs (Table 3 and Supplementary Table). The methylated regions were mainly located in introns, which proportion exceeded 70% (Fig. 5). The results of a boxplot analysis of DMRs showed that the methylation level of the HE group was lower than that of the CN group (Fig. 6). Furthermore, we also found that some of these DMGs were involved in biological processes important for skeletal muscle development: energy metabolism (such as 6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 1 [PFKFB], phosphoglycerate kinase 1 [PGK1] and pyruvate dehydrogenase kinase, isozyme 3 [PDK3]), transcription factors (such as cAMP responsive element binding [CREB], NF-κB- activating protein [NKAP] and signal transducer and activator of transcription 2 [STAT2]), Ca2+ homeostasis (chloride intracellular channel 2 [CLIC2], ryanodine receptor 3 [RYR3] and calcium channel, voltage-dependent, alpha 2/delta subunit 2 [CACNA2D]), protein kinases (such as calpain 2 [CAPN2] and protein kinase C [PRKCA]) and repair and stabilization of stressed proteins (such as αB-crystallin [CRYAB] and DnaJ [Hsp40] homolog, subfamily C, member 5 [DNAJC5]). However, gene ontogeny (GO) assignments and KEGG pathway analysis showed that these methylated genes were not significantly enriched in predicted process and signaling pathway (Supplementary Fig. 1 and Fig. 2).
Association analysis between the DMGs and the differentially expressed genes (DEGs)
To explore the relationship between these DMGs and the DEGs found at the transcriptome level, an association analysis was performed. This identified four overlapped genes: RYR3, PGK1, CRYAB and four and a half LIM domains 1 protein, isoform C (FHL1C; Fig. 7). In addition, DEGs were divided into highly regulated types and lowly regulated types, according to their gene expression level. For each type of gene, we analyzed the methylation level in five functional element regions (Fig. 8). For highly regulated genes, we observed a visible difference in methylation level between the CN and HE groups in the promoter regions for both CG and non-CG contexts. However, whereas a visible difference in methylation level was found in the exon regions of CHG contexts, differences in methylation levels were found in the 5′UTR, intron and 3′UTR regions of CHH contexts. On the other hand, for lowly regulated genes, there was a visible difference in methylation level between CN and HE groups in the 5′UTR regions of CG context. A visible difference in methylation level was also found in the 3′UTR regions of CHG contexts and a difference in methylation level was found in the promoter, 5′UTR and 3′UTR regions of CHH contexts.
Association analysis of DMGs and DEGs.
DEGs were divided into highly and lowly regulated types, according to their gene expression level. For each type of gene, methylation levels were analyzed in five functional element regions. (A) Highly regulated genes. (B) Lowly regulated genes. DEG, differentially expressed gene; DMG, differentially methylated gene. HE, heat-exposed group. CN, control group.
Discussion
DNA methylation profiles in skeletal muscle
Although global DNA methylation surveys have been performed on pigs15,17, this study is the first to systematically compare the genome-wide skeletal muscle methylation profiles between normal and heat-stressed pigs, which show different overall meat quality. Skeletal muscle is the major portion the animal used for human food and its features have a direct impact on meat quality. Moreover, the DNA methylation status of promoter and gene body regions can affect gene expression via changes in chromatin structure or transcription efficiency18,19,20; therefore, we aimed to compare the genome-wide methylation patterns between heat-exposed and normal pigs to identify differentially methylated genes that may affect the quality of meat exposed to heat stress. From these analyses, we found that the intronic regions of the pig genome comprised a large proportion of DMRs (>70%) and that only a small proportion of DMRs were located in the 3′UTR, 5′UTR and promoters. Furthermore, our previous study showed that the average daily gain (ADG) and average daily feed intake (ADFI) of heat-exposed pigs were both significantly decreased compared with those of control pigs21. These results could affect the observed changes in DNA methylation, in addition to direct heat stress. For decreased feed intake in heat-exposed pigs, food composition may be altered to compensate for decreased nutrients, which could offset a proportion of the observed methylation changes.
In mammals, most DNA methylation occurs at CpG sites and asymmetric non-CpG methylation has been detected at appreciable levels in a few cell types22,23,24. Methylation in symmetrical sequences is preserved through cycles of DNA replication by maintenance DNA methyltransferases, which show a preference for hemimethylated substrates and methylate cytosines in the newly synthesized strand. Maintenance mechanisms for asymmetric methylation patterns are unknown, but must include de novo methylation after each cell division. Results of a recent study by Guo et al.25 showed that CpG methylation is always symmetrically methylated, whereas non-CpG sites are strand-specifically methylated in introns, short interspersed nuclear elements and long interspersed nuclear elements25. They also showed that the skew of non-CpG methylation in introns is more pronounced at the boundaries and more significant for highly expressed genes. The results of this study provide further details regarding the biological significance of the various methylation patterns (CG, CHG and CHH), particularly in the context of pigs.
Key differentially methylated genes related to muscle development and meat quality
Porcine skeletal muscle development and growth have been shown to be associated with the differential expression of genes. In this study, several genes involved in muscle growth and development, including small muscle protein X-linked (SMPX), myosin, heavy chain 11 (MYH11), collagen, type XVI, alpha 1 (COL16A1), and collagen, type IV, alpha 3 (COL4A3), exhibited hypomethylation following constant heat stress. This result indicated that the expression of these genes might be upregulated. SMPX is upregulated in skeletal muscle by stretching and has been suggested to serve as both a transcription factor and a mechanosensor, possibly giving rise to changes in muscle fiber size and type. Thus, SMPX is thought to be a candidate gene for regulating muscle size. Schindeler et al.26 also found that SMPX might participate in the regulation of cytoskeletal dynamics through the Rac1/p38 pathway26. Additionally, differentiating C2C12 myoblasts overexpressing SMPX increase their susceptibility to fuse, forming large “myosacs” in an IGF-1 dependent manner27 further indicating a role for SMPX in the regulation of fiber size.
COL16A1 is a gene suggested to have importance for cell adhesion28. COL4A3 was also identified as a structural component of collagen, which is known to affect meat tenderness. In turn, myosin is a major structural protein of the thick filament of the sarcomere28. A previous study demonstrated that MyHC isoform composition affects the quality of the meat originating from bovine muscles55 was used to perform alignments of bisulfite-treated reads to a reference genome using the default parameters. The reference genome was first transformed into a bisulfite-converted version (C-to-T and G-to-A conversions) and then indexed using bowtie256. Sequence reads were also transformed into fully bisulfite-converted versions (C-to-T and G-to-A conversions) before they were aligned to similarly converted versions of the genome in a directional manner. Sequence reads that produced a unique best alignment from the two alignment processes (original top and bottom strand) were then compared to the normal genomic sequence and the methylation state of all cytosine positions in the read was inferred. The same reads that aligned to the same regions of the genome were regarded as duplicated reads. The sequencing depth and coverage were summarized using deduplicated reads. The sodium bisulfite non-conversion rate was calculated as the percentage of cytosines sequenced at cytosine reference positions in the lambda genome.
Estimating methylation levels
To identify the sites of methylation, we modeled the sum  of methylated counts as a binomial (Bin) random variable with a methylation rate as follows:
of methylated counts as a binomial (Bin) random variable with a methylation rate as follows:

We employed a sliding-window approach, which is conceptually similar to approaches that have been used for bulk bisulfite sequencing (http://www.bioconductor.org/packages/2.13/bioc/html/bsseq.html). Using a window size w = 3,000 bp and step size of 600 bp57, the sum of methylated and unmethylated read counts in each window were calculated. The methylation level (ML) for each C site showed the fraction of methylated Cs and was defined as:

The calculated ML value was further corrected using the bisulfite non-conversion rate according to previous studies58. Given the bisulfite non-conversion rate r, the corrected ML was estimated as:

Differentially methylated region analysis
Differentially methylated regions (DMRs) were identified using swDMR software (http://122.228.158.106/swDMR/), using a sliding-window approach. The window was set to 1000 bp and the step length was 100 bp. A Fisher test was then implemented to detect the DMRs. We defined regions as DMRs that had corrected P-value of less than 0.05, reads coverage of greater than 5, fold-change values of greater than 2 and false discovery rate of less than 0.05.
GO and KEGG enrichment analysis of DMR-related genes
GO enrichment analysis of genes related to DMRs was implemented by the GOseq R package59, in which gene length bias was corrected. KOBAS software60 was used to test the statistical enrichment of DMR-related genes in KEGG pathways. GO terms and KEGG with corrected P-values of less than 0.05 were considered significantly enriched by DMR-related genes.
Transcriptome analysis
Sample preparation for RNA-seq analysis
mRNA was purified from total RNA using poly-T oligo-attached magnetic beads. Fragmentation was performed using divalent cations under elevated temperatures in a proprietary Illumina fragmentation buffer. First-strand cDNA was synthesized using random oligonucleotides and SuperScript II (Life Technologies Inc, Rockville, MD, USA). Second-strand cDNA synthesis was then performed using DNA polymerase I and RNase H. Remaining overhangs were converted into blunt ends using exonuclease and polymerase. After the adenylation of the 3′ ends of the DNA fragments, Illumina PE adapter oligonucleotides were ligated in preparation for hybridization. To preferentially select cDNA fragments 200 bp in length, the library fragments were purified using the AMPure XP system (Beckman Coulter, Beverly, MA, USA). DNA fragments with adaptors ligated on both ends were selectively enriched using the Illumina PCR Primer Cocktail with a 10-cycle PCR protocol. The products were then purified (AMPure XP system) and quantified using Agilent high-sensitivity DNA assays on the Agilent Bioanalyzer 2100 system.
Clustering and sequencing
Clustering of the index-coded samples was performed on a cBot Cluster Generation System using the TruSeq PE Cluster Kit v3-cBot-HS (Illumina). After cluster generation, the library preparations were sequenced on an Illumina Hiseq 2000 platform and 100-bp paired-end reads were generated (Novogene, Bei**g, China).
Quantification of gene expression levels
After quality control and filtering of raw data, clean reads were aligned to the S. scrofa 10.2 reference genome using TopHat v1.4.061, which was chosen as it could generate a database of splice junctions based on the gene model annotation file to provide better map** results.
HTSeq v0.5.3 was used to count the read numbers mapped to each gene. The reads per kilobase of exon model per million mapped reads (RPKM) of each gene was calculated based on the length of the gene and the number of reads mapped to this gene.
Differential gene expression analysis
Differential gene expression between the control and heat stress groups was analyzed using the DESeq R package (1.8.3). The P-values were adjusted using the Benjamini & Hochberg method62. A corrected P-value of 0.05 was set as the threshold for significantly different expression.
Association analysis
We estimated the associations between the DMGs and the differentially expressed genes (DEGs) by Pearson correlation. In addition, DEGs were divided into highly regulated types and lowly regulated types, according to the results of transcriptome analysis. For each type of gene, we analyzed the methylation level in five functional element regions. Each gene function element was divided into 20 bins according to the location. The methylation level of each bin was calculated sequentially and the average methylation levels in bin1, bin2, bin3 … bin20 of all genes were then calculated.
Additional Information
How to cite this article: Hao, Y. et al. Genome-wide DNA methylation profiles changes associated with constant heat stress in pigs as measured by bisulfite sequencing. Sci. Rep. 6, 27507; doi: 10.1038/srep27507 (2016).
References
St-Pierre, C. B. & Schnitkey, G. Economic Losses from Heat Stress by US Livestock Industries. J. Dairy Sci. 86, E52–E77 (2003).
Quinteiro-Filho, W. M. et al. Heat stress impairs performance parameters, induces intestinal injury and decreases macrophage activity in broiler chickens. Poult. Sci. 89, 1905–1914 (2010).
Patience, J., Umboh, J., Chaplin, R. & Nyachoti, C. Nutritional and physiological responses of growing pigs exposed to a diurnal pattern of heat stress. Livest. Prod. Sci. 96, 205–214 (2005).
Kamanga-Sollo, E., Pampusch, M. S., White, M. E., Hathaway, M. R. & Dayton, W. R. Effects of heat stress on proliferation, protein turnover and abundance of heat shock protein messenger ribonucleic acid in cultured porcine muscle satellite cells. J. Anim. Sci. 89, 3473–3480 (2011).
Mustafi, S. B., Chakraborty, P. K., Dey, R. S. & Raha, S. Heat stress upregulates chaperone heat shock protein 70 and antioxidant manganese superoxide dismutase through reactive oxygen species (ROS), p38MAPK and Akt. Cell Stress Chaperone 14, 579–589 (2009).
Yoshihara, T. et al. Heat stress activates the Akt/mTOR signalling pathway in rat skeletal muscle. Acta. Physiol (Oxf). 207, 416–426 (2013).
Foury, A. et al. Stress hormones, carcass composition and meat quality in Large White x Duroc pigs. Meat Sci. 69, 703–707 (2005).
Leheska, J. M., Wulf, D. M. & Maddock, R. J. Effects of fasting and transportation on pork quality development and extent of postmortem metabolism. J Anim Sci. 80, 3194–202 (2002).
Terenina, E. et al. Association study of molecular polymorphisms in candidate genes related to stress responses with production and meat quality traits in pigs. Domest. Anim. Endocrinol 44, 81–97 (2013).
Yang, P. G. et al. The expression of carnosine and its effect on the antioxidant capacity of Longissimus dorsi muscle in finishing pigs exposed to constant heat stress. Asian-Australas J. Anim. Sci. 27, 1763–1672 (2014).
Gao, F. et al. Comparison of gene expression and genome-wide DNA methylation profiling between phenotypically normal cloned pigs and conventionally bred controls. PLoS One 6, e25901 (2011).
Li, Q. et al. Genome-wide map** of DNA methylation in chicken. PLoS One 6, e19428 (2011).
Hu, Y. et al. Comparison of the genome-wide DNA methylation profiles between fast-growing and slow-growing broilers. PLoS One 8, e56411 (2013).
Su, J., Wang, Y., **ng, X., Liu, J. & Zhang, Y. Genome-wide analysis of DNA methylation in bovine placentas. BMC Genomics 15, 12 (2014).
Kwak, W. et al. Genome-wide DNA methylation profiles of small intestine and liver in fast-growing and slow-growing weaning piglets. Asian-Australas J. Anim. Sci. 27, 1532–1539 (2014).
Moresi, V., Marroncelli, N., Coletti, D. & Adamo, S. Regulation of skeletal muscle development and homeostasis by gene imprinting, histone acetylation and microRNA. Biochim. Biophys. Acta. 1849, 309–316 (2015).
Li, M. et al. An atlas of DNA methylomes in porcine adipose and muscle tissues. Nat. Commun. 3, 850 (2012).
Lorincz, M. C., Dickerson, D. R., Schmitt, M. & Groudine, M. Intragenic DNA methylation alters chromatin structure and elongation efficiency in mammalian cells. Nat. Struct. Mol. Biol. 11, 1068–1075 (2004).
Klose, R. J. & Bird, A. P. Genomic DNA methylation: the mark and its mediators. Trends Biochem. Sci. 31, 89–97 (2006).
Suzuki, M. M. & Bird, A. DNA methylation landscapes: provocative insights from epigenomics. Nat. Rev. Genet. 9, 465–476 (2008).
Hao, Y. et al. Nutritional and physiological responses of finishing pigs exposed to a permanent heat exposure during three weeks. Arch. Anim. Nutr. 68, 296–308 (2014).
Lister, R. et al. Hotspots of aberrant epigenomic reprogramming in human induced pluripotent stem cells. Nature 471, 68–73 (2011).
Ziller, M. J. et al. Genomic distribution and inter-sample variation of non-CpG methylation across human cell types. PLoS Genet. 7, e1002389 (2011).
Tomizawa, S. et al. Dynamic stage-specific changes in imprinted differentially methylated regions during early mammalian development and prevalence of non-CpG methylation in oocytes. Development 138, 811–820 (2011).
Guo, W., Chung, W. Y., Qian, M., Pellegrini, M. & Zhang, M. Q. Characterizing the strand-specific distribution of non-CpG methylation in human pluripotent cells. Nucleic Acids Res. 42, 3009–3016 (2014).
Schindeler, A., Lavulo, L. & Harvey, R. P. Muscle costameric protein, Chisel/Smpx, associates with focal adhesion complexes and modulates cell spreading in vitro via a Rac1/p38 pathway. Exp. Cell Res. 307, 367–380 (2005).
Palmer, S. et al. The small muscle-specific protein Csl modifies cell shape and promotes myocyte fusion in an insulin-like growth factor 1-dependent manner. J. Cell Biol. 153, 985–998 (2001).
Zhang, Y. Y. et al. Differentially expressed genes in skeletal muscle tissues from castrated Qinchuan cattle males compared with those from intact males. Livest. Sci. 135, 76–83 (2001).
**ong, Y. L. Myofibrillar protein from different muscle fiber types: implications of biochemical and functional properties in meat processing. Crit. Rev. Food Sci. Nutr. 34, 293–320 (1994).
Bai, Q. et al. Development of a porcine skeletal muscle cDNA microarray: analysis of differential transcript expression in phenotypically distinct muscles. BMC Genomics 4, 8 (2003).
Tang, S., Yang, L., Tang, X. & Liu, M. The role of oxidized ATM in the regulation of oxidative stress-induced energy metabolism reprogramming of CAFs. Cancer Lett. 353, 133–144 (2014).
Xu, Y. et al. Differential proteome and transcriptome analysis of porcine skeletal muscle during development. J. Proteomics 75, 2093–2108 (2012).
Sugden, M. C. & Holness, M. Interactive regulation of the pyruvate dehydrogenase complex and the carnitine palmitoyltransferase system. FASEB J. 8, 54–61 (1994).
Kim, T. et al. Carnitine Palmitoyltransferase 1b Deficient Mice Develop Severe Insulin Resistance After Prolonged High Fat Diet Feeding. J. Diabetes Metab. 5, 10.4172/2155-6156.1000401 (2014).
Hirose, K. et al. Evaluation of effects of multiple candidate genes (LEP, LEPR, MC4R, PIK3C3 and VRTN) on production traits in Duroc pigs. Anim. Sci. J. 85, 198–206 (2014).
Noble, E. G., Milne, K. J. & Melling, C. W. Heat shock proteins and exercise: a primer. Appl. Physiol. Nutr. Metab. 33, 1050–1065 (2008).
Paulsen, G. et al. Maximal eccentric exercise induces a rapid accumulation of small heat shock proteins on myofibrils and a delayed HSP70 response in humans. Am. J. Physiol. Regul. Integr. Comp. Physiol. 293, R844–853 (2007).
Koh, T. J. Do small heat shock proteins protect skeletal muscle from injury? Exerc. Sport Sci. Rev. 30, 117–121 (2002).
Nussbaum, E. L. & Locke, M. Heat shock protein expression in rat skeletal muscle after repeated applications of pulsed and continuous ultrasound. Arch. Phys. Med. Rehabil. 88, 785–790 (2007).
Pan, P. J., Hsu, C. F., Tsai, J. J. & Chiu, J. H. The role of oxidative stress response revealed in preconditioning heat stimulation in skeletal muscle of rats. J. Surg. Res. 176, 108–113 (2012).
Bukau, B. & Horwich, A. L. The Hsp70 and Hsp60 chaperone machines. Cell 92, 351–366 (1998).
Kelley, W. L. Molecular chaperones: How J domains turn on Hsp70s. Curr. Biol. 9, R305–308 (1999).
MacLennan, D. H. & Phillips, M. S. Malignant hyperthermia. Science 256, 789–794 (1992).
Davis, M. C. et al. Dexamethasone-induced inositol 1,4,5-trisphosphate receptor elevation in murine lymphoma cells is not required for dexamethasone-mediated calcium elevation and apoptosis. J. Biol. Chem. 283, 10357–10365 (2008).
Fujii, J. et al. Identification of a mutation in porcine ryanodine receptor associated with malignant hyperthermia. Science 253, 448–451 (1991).
Board, P. G., Coggan, M., Watson, S., Gage, P. W. & Dulhunty, A. F. CLIC-2 modulates cardiac ryanodine receptor Ca2+ release channels. Int. J. Biochem. Cell Biol. 36, 1599–1612 (2004).
Cao, G. et al. Molecular cloning and characterization of a novel human cAMP response element-binding (CREB) gene (CREB4). J. Hum. Genet. 47, 373–376 (2002).
Yossifoff, M., Kisliouk, T. & Meiri, N. Dynamic changes in DNA methylation during thermal control establishment affect CREB binding to the brain-derived neurotrophic factor promoter. Eur. J. Neurosci. 28, 2267–2277 (2008).
Chen, D. et al. Identification of a nuclear protein that promotes NF-kappaB activation. Biochem. Biophys. Res. Commun. 310, 720–724 (2003).
Pahl, H. L. Activators and target genes of Rel/NF-kappaB transcription factors. Oncogene 18, 6853–6866 (1999).
Eizirik, D. L. et al. Cytokines suppress human islet function irrespective of their effects on nitric oxide generation. J. Clin. Invest. 93, 1968–1974 (1994).
Bendtzen, K. et al. Cytotoxicity of human pI 7 interleukin-1 for pancreatic islets of Langerhans. Science 232, 1545–1547 (1986).
Mahdi, T. et al. Secreted frizzled-related protein 4 reduces insulin secretion and is overexpressed in type 2 diabetes. Cell Metab. 16, 625–633 (2012).
Hall, E. et al. Sex differences in the genome-wide DNA methylation pattern and impact on gene expression, microRNA levels and insulin secretion in human pancreatic islets. Genome Biol. 15, 522 (2014).
Krueger, F. & Andrews, S. R. Bismark: a flexible aligner and methylation caller for Bisulfite-Seq applications. Bioinformatics 27, 1571–1572 (2011).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Smallwood, S. A. et al. Single-cell genome-wide bisulfite sequencing for assessing epigenetic heterogeneity. Nat. Methods 11, 817–820 (2014).
Lister, R. et al. Global epigenomic reconfiguration during mammalian brain development. Science 341, 1237905 (2013).
Young, M. D., Wakefield, M. J., Smyth, G. K. & Oshlack, A. Gene ontology analysis for RNA-seq: accounting for selection bias. Genome Biol. 11, R14 (2010).
Mao, X., Cai, T., Olyarchuk, J. G. & Wei, L. Automated genome annotation and pathway identification using the KEGG Orthology (KO) as a controlled vocabulary. Bioinformatics 21, 3787–3793 (2005).
Trapnell, C., Pachter, L. & Salzberg, S. L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111 (2009).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010).
Acknowledgements
The authors thank Novogene Company for excellent technical help. This research was supported by the National Key Basic Research Program of China (National 973 Project; 2012CB124706), National Natural Science of China (31501961), the Agricultural Science and Technology Innovation Program (ASTIP-IAS07) and Key Project in the National Science & Technology Pillar Program (2012BAD39B0202).
Author information
Authors and Affiliations
Contributions
Y.H. conducted experimental design and prepared the manuscript. Y.C. provided help for sample collection and carried out DNA extraction and preparation. X.G. conceived the project, participated in experimental design and was in charge of the overall project. All authors reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Hao, Y., Cui, Y. & Gu, X. Genome-wide DNA methylation profiles changes associated with constant heat stress in pigs as measured by bisulfite sequencing. Sci Rep 6, 27507 (2016). https://doi.org/10.1038/srep27507
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep27507
- Springer Nature Limited
This article is cited by
-
Joint Analysis of Genome-wide DNA Methylation and Transcription Sequencing Identifies the Role of BAX Gene in Heat Stress–Induced-Sertoli Cells Apoptosis
Reproductive Sciences (2024)
-
Transcriptome and DNA Methylation Responses in the Liver of Yellowfin Seabream Under Starvation Stress
Marine Biotechnology (2023)
-
Influence of e-waste exposure on DNA damage and DNA methylation in people living near recycling sites
Environmental Science and Pollution Research (2023)
-
Gestational heat stress alters skeletal muscle gene expression profiles and vascularity in fetal pigs in a sexually dimorphic manner
Journal of Animal Science and Biotechnology (2022)
-
A draft genome of Drung cattle reveals clues to its chromosomal fusion and environmental adaptation
Communications Biology (2022)