Abstract
Motile cells inside living tissues often encounter junctions, where their path branches into several alternative directions of migration. We present a theoretical model of cellular polarization for a cell migrating along a one-dimensional line, arriving at a symmetric Y junction and extending protrusions along the different paths that originate at the junction. The model predicts the spontaneous emergence of deterministic oscillations of growth and cellular polarization between competing protrusions during the directional decision-making process. The oscillations are modified by cellular noise but remain a dominant feature that affects the time it takes the cell to migrate across the junction. These predictions are confirmed experimentally for two different cell types (non-cancerous endothelial and cancerous glioma cells) migrating on a patterned network of thin adhesive lanes with junctions.
Similar content being viewed by others
Main
Directed cell movement is central to processes such as organ development, tissue regeneration, cancer metastasis and immune response1,2,3. A moving cell must negotiate obstacles that frequently generate spatial junctions or bifurcations. On encountering such a bifurcation, the cell has to decide on a new direction of motion4,5,6,7,8. Although previous works on cells encountering bifurcations have focused on directed migration, in which the arms in the junction are biased due to hydraulic pressure4,6,7, chemical cues6 or size5, less is known about how a single cell behaves when encountering a spatial bifurcation in which the paths of the cell are symmetric, such that the cell is not directed by an external cue or geometrical constraints.
When migrating inside tissues, cells navigate through complex geometries along complex trajectories and have highly branched shapes9,10. Migrating cells face obstacles imposed by the surrounding extracellular matrix and cells of the surrounding tissue. Cells moving in such an environment extend protrusions that probe the alternative narrow paths11 and, therefore, need to efficiently choose the optimal direction of migration. When encountering a junction along the path, the cell may get stuck, go back or continue forward along one of the available paths12. As an example of this process, Fig. 1a shows cancerous cells (human glioma propagating cells, hGPCs) moving along a network of thin blood vessels in a mouse brain explant. Glioma cells often encounter bifurcating junctions as they invade the brain along the abluminal side of the network of blood vessels, which have frequent branches with Y-shaped junctions (Fig. 1a)13.
a, hGPCs migrating in a mouse brain slice (brain slice overlay assay47, Supplementary Video 1). The cells migrate along the surface of the blood vessels with a quasi-linear motion and encounter junctions where the blood vessels bifurcate. Inset, zoom-in of a bifurcating blood vessel. b, Phase-contrast time-lapse images of an hGPC migrating along a laminin-coated Y junction (left panels, Supplementary Video 2) and an endothelial cell (HUVEC) migrating along a fibronectin-coated Y junction (right panels, Supplementary Video 3). c, Model illustration. Three arms of length li with local polymerization speed vi (i = 1, 2 or 3) move along the confinement of a three-way symmetric Y-junction (black lines). All arms are connected by a spring. Green and yellow gradients denote areas of low or high concentration, respectively, of a polarity cue that inhibits the actin polymerization at the leading edges of the arms (red denotes the polymerization strength). At each edge, the friction force acts as a drag friction when the arm extends and as a slip-bond friction when the arm retracts20.
To decipher how motile cells pass through complex paths, several in vitro studies have constrained cells to migrate along networks of confining one-dimensional channels or adhesive stripes5,6,7,14,15,16,17,18. When moving through such a network, the cells encounter junctions, and they are often observed to form several protrusions along the alternative paths that leave the junction. Eventually, a cell has to choose one of the paths to leave the junction and continue its migration through the network. This migration may be random or oriented using an external chemotactic gradient that biases the motion. Examples of such in vitro experiments are shown in Fig. 1b, in which an hGPC and a human umbilical vein endothelial cell (HUVEC) are observed to migrate across a Y junction composed of adhesive lanes patterned over a non-adhesive background.
We present a theoretical framework that extends a model for the self-polarization of one-dimensional cells19,20 to include several competing cellular protrusions. Our simplified model of cell migration, which treats a cell as being composed of linear segments, allows us to obtain a deep understanding of the resulting complex dynamics without having to describe general cellular shapes21. Our model predicts that during the passage of a cell over a symmetric Y junction, the polarization of the cell undergoes deterministic oscillations between the competing cellular protrusions, resulting in cycles of elongation and retraction. These oscillations are an inevitable result of the polarization mechanism of the cell, which in our model is sensitive to the length changes of the competing protrusions. We find that the deterministic oscillations are modified by the noise inherent to the intracellular actin dynamics. These theoretical predictions motivated experimental studies in two different cell systems, non-cancerous endothelial cells (HUVECs) and cancerous cells (hGPCs). Both of these cell types face junctions while migrating through blood vessels during angiogenesis and repair22 or brain invasion23. In addition, endothelial cells and glioma cells present different migratory behaviours, which implies that there are different actin-polymerization regimes related to the speed of migration. We find that despite these differences, both cell types display oscillatory dynamics and exhibit motility modes along the junction that fit the different dynamical phases predicted by the model.
The model
We extend the model of a cell migrating along a linear track20 to a cell moving over a symmetric Y junction. When spanning the junction, the cell has three arms, which are free to extend or retract (Fig. 1c). Within the model, the dynamics of each arm (denoted by i) is described by three dynamical variables: (1) its length li, (2) the local actin-polymerization speed vi at the leading edge of the arm and (3) the concentration of adhesions ni at the arm’s leading edge:
where α is the actin sliding coefficient, which couples the actin-polymerization speed to the protrusive force exerted on the membrane, k is the mean elastic constant of the cell (of rest length 1), fs is the force susceptibility of the adhesion molecules (which behave as slip bonds), δ is the response rate of the actin-polymerization speed, r is the equilibrium dissociation constant of the adhesion molecules, which describes the effective adhesiveness of the surface, and L = ∑jlj is the total length of the cell. Note that the parameter k also takes into account the effect of active cell contractility. The term σξt refers to uncorrelated Gaussian noise with an amplitude σ that is added to the actin-polymerization speed. For a detailed derivation of the model, noise implementation and explanation of the numerical scheme used to solve equations (1)–(3), see Supplementary Information sections 1 and 2. Also note that equations (1)–(3) are non-dimensional and follow the scaling in ref. 20.
We describe the stick–slip dynamics through the friction term Γi in equation (1), in which the friction is a constant drag when an arm extends forward or a slip-bond adhesion friction when it retracts20:
where γ is the drag friction coefficient and κ is the elastic constant of the adhered slip bonds.
In the model, the net global actin treadmilling flow within an arm ui advects a polarity cue, which in turn affects the overall actin treadmilling flow in the cell. Examples of molecules that can serve as this polarity cue include those that contribute to the actin retrograde flow such as myosin II (refs. 19,24), those that debranch the actin network such as Arpin19, GMF25 and Coronin26, and other possible negative regulators of actin polymerization27,28. Here we assume that the polarity cue inhibits the local actin polymerization (or local flow) vi emanating from the leading edges of the arms (Fig. 1c). The steady-state global and local flows, as functions of the arm lengths and the local concentration of the polarity cue at the tips of the arms c(li, ui), are given by
where D is the diffusion coefficient of the polarity cue, and c is a dimensionless quantity that represents the ratio between the total concentration of the polarity cue and its dissociation constant (Supplementary Information section 2). A key parameter in our model is β, which is the maximal actin-polymerization speed produced at the leading edge of the cellular arms.
The steady-state local actin flow given by equation (5) describes the inhibitory effect of the polarity cue on the local actin-polymerization activity at each arm. The actin flow from each arm is assumed to split evenly at the junction and, therefore, contributes a half of its flux when calculating the net flow within each arm (equation (6)). It is this net flow that advects the polarity cue, which explains why the exponential function that describes the concentration of the polarity cue at the edge of an arm contains these net flow speeds (equation (7)). The total amount of polarity cue, namely its integral over the entire cell, is conserved. For the full derivation of equations (5)–(7), see Supplementary Information section 1.
Throughout this work, the model equations are normalized by the timescale of the inverse of the focal adhesion disassembly rate (5–30 min; refs. 29,30,31,32,33,34) and by the length scale of the rest length of the cell, which is when the cell is stationary and does not protrude extensions along the one-dimensional track (10–100 μm; ref. 20). The friction coefficients α and γ are normalized to α = γ = 1, which set the force scale of the force-related parameters in the model. For details of the parameter estimation and scaling, see Supplementary Information section 3.
Symmetry breaking
Within our model, cell migration occurs when the cell spontaneously self-polarizes due to the feedback between the actin flow and the intracellular distribution of a polarity cue. The polarity cue is advected by the actin flow, which in turn determines the flow by inhibiting actin polymerization at the leading edges of the arms (equation (5)). We investigate the conditions in our model under which cells polarize and turn migratory by exploring the k–β parameter space (Fig. 2a) and analyse the conditions that allow cells to migrate across the Y junction, for two limiting cases: (1) a symmetric case, in which the cell lands at the centre of the junction and spreads symmetrically in all directions (Fig. 2b–d), and (2) a moving case, in which the cell migrates at a steady state along a linear track20, enters the junction and grows two symmetric protrusions along the new arms (Fig. 2e–g). The first case, where a cell lands at the centre of the junction, is simpler to solve analytically but is not often observed experimentally, whereas the second case is the more common experimental observation.
a, Phase diagram of the cellular dynamics according to the model, as functions of the parameters β and k. The white region is where cells remain trapped in the junction, that is, β < βc (equation (10)). The light red region, above the solid red line, is where cells escape the junction, that is, β > βc (equation (10)). The darker red region above the dashed red line is where cells can remain in a metastable configuration (the slow process) before esca** the junction (Supplementary Information section 4). b, The symmetric spreading case. All arms initially extend symmetrically with the same local flow v. c, For β = 5 < βc, the cell reaches a stationary shape, in which all the arms are of equal length, with the same local actin-polymerization activity. d, For β = 6.5 > βc, the cell polarizes and leaves the junction while the local flows in the arms undergo seesaw oscillations. (i)–(iii) Time series of the arm lengths and the local and global actin flows. e, The moving case. A cell of length l enters the junction. At the front, the cell extends two symmetric protrusions, initially of equal lsmall length ϵ with the same local actin-polymerization speed v. At the rear (in red), the actin polymerization is v1. f, For β = 5 < βc, the migrating cell stops at the junction. g, For β = 6.5 > βc, the cell migrates past the junction, leaving it through one of the new protruding arms. (i)–(iii) Time series of the arm lengths and the local and global flows. Red, green and blue curves represent arms 1, 2 and 3, respectively. The solid black line in (i) indicates the total cell length. Dashed black and dashed grey lines in (i) indicate Lc and Lp, respectively. Parameters: δ = 250, c = 3.85, D = 3.85, k = 0.8, r = 5, fs = 5, κ = 20 and σ = 10−7 (noise amplitude).
The polarization of the cell is critically dependent in our model on its length20. For the symmetrically spreading cell (Fig. 2b), we can calculate the length (Lp) for which there is a balance between the protrusive forces and the elasticity of the cell20:
and we can also calculate the critical length (Lc) at which the feedback between the treadmilling flow and the polarity cue destabilizes the uniform solution (in the limit ui → 0) and the cell becomes polarized:
By equating equations (8) and (9), we obtain the critical polarization speed:
For β < βc (white region below the solid red curve in Fig. 2a), the cell spreads and reaches a symmetric, stationary shape that is stable at the junction (Fig. 2c). For β > βc (red regions above the solid red curve in Fig. 2a), the cell spreads and spontaneously polarizes along one of the arms when its total length exceeds Lc (Fig. 2d(i)). The cell eventually migrates away from the junction, as indicated by two of the arms shrinking to zero (Fig. 2d(i)).
We highlight here that βc (equation (10)) is larger than the critical value of β required for spontaneous motion along a straight line20. This is due to an additional leading edge that the cell has on the Y junction, which increases the dilution of the polarity cue, which we consider to be a limited resource that is needed for spontaneous polarization. For a comparison between the critical values of β during symmetry breaking at a Y junction and along a one-dimensional track, see Supplementary Information section 5.
For the moving cell, we find that below βc, the moving cell stops at the junction and remains stationary (Fig. 2f), whereas above βc, the cell migrates across the junction. The two new protrusions compete until the symmetry breaks and the cell decides on a new path (Fig. 2f). The critical value of β is slightly lower in the moving case than in the symmetric case (equation (10); red curve in Fig. 2a) due to the different condition on the force balance that gives rise to Lp (equation (8)). In Supplementary Information section 5, we provide our approximations for Lp and βc for the moving case.
Note that we find another critical value of \(\beta ={\beta }_\mathrm{c}^\mathrm{slow}\) (dashed red curve in Fig. 2a), above which the system can dwell for long periods of time in a metastable configuration (the slow process) during which two competing arms are in a force balance (while the third arm has shrunk to zero length; see Supplementary Information section 4). This transition will be discussed further in a later section.
Deterministic escape patterns of seesaw oscillations
In both cases, of symmetric spreading and migration to the junction, we find that above βc, the local actin-polymerization activity flips at the tips of the competing arms like a seesaw as the cell moves past the junction (Fig. 2c,f). This seesaw-oscillation competition of the actin-polymerization activity affects the growth and retraction of the arms, as seen in their length dynamics. This behaviour is a distinct deterministic mechanism, not driven by noise, which is negligible in these simulations.
The origin of the seesaw-oscillation mechanism is due to the redistribution of the conserved amount of polarity cue over the entire length of the cell, which is highly sensitive to the length changes. We analyse the origin of these oscillations in detail for a symmetrically spreading cell that undergoes symmetry breaking (Extended Data Fig. 1 at t ≈ 2). When the symmetry breaks, one arm retracts (green curve in Extended Data Fig. 1a(i)) as the local actin-polymerization speed at its tip rapidly decreases (green curve in Extended Data Fig. 1a(ii)), while the two remaining arms compete for the cell polarization direction (blue and red curves in Extended Data Fig. 1a(i,ii)). This symmetry-breaking event is then followed by two cycles of seesaw oscillations between one of the competing arms and the retracting arm (red and green curves in Extended Data Fig. 1a(i,ii)). These cycles are denoted by the dashed vertical lines in Extended Data Fig. 1a. During these seesaw oscillations, the local flows (vi) undergo rapid and large changes (Extended Data Fig. 1a(ii)), driving corresponding changes to the lengths of the arms (Extended Data Fig. 1a(i)).
We analyse the seesaw-oscillation mechanism by plotting the steady-state solutions of the local flows in the proximity of the seesaw events (Extended Data Fig. 1b,c). For each seesaw event, we chose a time just before and after the event (vertical dashed lines in Extended Data Fig. 1a) and use the simulations to calculate the ratio between the lengths of the arms at these times (Extended Data Fig. 1a(iii)). We then plot the self-consistent solutions for the actin-polymerization speeds (equation (5)) as a function of the length of the winning arm (l3), while maintaining the ratios fixed (Extended Data Fig. 1b,c). The solutions associated with the value of l3 from the simulations are denoted by the black circles in Extended Data Fig. 1b,c. The steady-state solutions are obtained using a continuation method35.
This analysis shows that the seesaw-oscillation mechanism lies in transitions between the different solution branches as the arms elongate or retract and the ratios of their lengths change. Just before the seesaw events, the solutions of the losing arms (for v1 and v2 in red and green, respectively) are at the tips of a solution-branch saddle (Extended Data Fig. 1b(i,ii),c(i,ii)). As the winning arm l3 grows larger, the ratios between l3 and the losing arms l1 and l2 decrease (Extended Data Fig. 1a(iii)). As a result, the saddle shifts slightly yet enough to force the solutions to discontinuously switch between the different solution branches. These branch transitions are most evident in the local flows in the losing arms v1 and v2 but also occur for the local flow in the winning arm v3 in the first seesaw event (Extended Data Fig. 1b(iii)).
We emphasize that this nonlinear deterministic pattern does not arise due to growing arm(s) simply pulling on the remaining arms and causing them to retract. Rather, the mechanism arises due to the sensitive interdependence of the distribution of the finite amount of polarity cue over the whole cell and its effect on the local actin-polymerization activity at the ends of the competing arms. Due to this competition, the actin-polymerization activity shifts between the leading edges of the competing arms, such that they switch to be out of phase with each other. Using the same analysis, we also find another oscillating seesaw pattern, with a smaller amplitude, which is associated with the stabilization of the local flows in the losing arms before the cell leaves the junction (Supplementary Information section 6).
Figure 3 shows examples of a HUVEC and a hGPC performing seesaw oscillations of the local actin-polymerization activity at the tips of the arms, including model predictions (for the experimental methods, see Supplementary Information section 7). We quantify the actin-polymerization activity at the tips of the arms in the experiments as follows. We define a region extending over a distance of 10 μm back from the edge of each arm. Within this region, we measure the overall membrane area (tip area), which captures the amount of protrusive lamellipodia activity. We also measure the total actin fluorescence intensity (tip actin intensity) within this region as a measure of the actin-polymerization activity at the leading edge of the arm. Both measures are compared to the local flows in the model (vi, equation (2)), and both exhibited the predicted seesaw-oscillation cycles in the losing arms (Figs. 3a(ii,iii),f(ii,iii)).
a–e, HUVEC experiment and simulation a, Experiment. Total area of the arms (i). Lamellipodia (arm tip) area (ii). Lamellipodia actin intensity (iii). b, Simulation with (β, σ) = (6.7, 0.5). Arm length (i). Local actin flows (ii). Red, green and blue curves indicate arms 1, 2 and 3. The black curves indicate the total cell length. Grey sections are regions where the local polymerization speed of the losing arms display seesaw oscillations. Grey dashed lines in a correspond to the time-lapse images in d and the arrows in c. Grey dashed lines in b correspond to the pictograms in e. c, Arm kymographs. d, Time-lapse images (Supplementary Video 4). Upper panels, whole cell. Lower panels, enlarged arm protrusion tips. The yellow lines indicate a 10 μm window at which the intensity and area at the tips were measured. e, Pictograms of the simulation. f–i, As a–e but for the hGPC experiment and simulation. f, Experiment. Total area of the arms (i). Lamellipodia (arm tip) area (ii). Lamellipodia actin intensity (iii). g, Simulation with (β, σ) = (8.7, 0.3). Arm length (i). Local actin flows (ii). h, Arm kymographs. i, Time-lapse images (Supplementary Video 5). j, Pictograms of the simulation. Other simulation parameters: c = 3.85, D = 3.85, k = 0.8, fs = 5, r = 5, κ = 20 and δ = 250. Conversion of simulation units (Supplementary Table 1): 1 time unit = 5–30 min, 1 length unit = 10–100 μm.
When comparing the simulations to the experimental observations (Fig. 3), we fitted two parameters of the model to capture the basic qualitative features of the dynamics of the arms lengths: the maximal actin-polymerization speed β and its noise amplitude σ. For the HUVEC (Fig. 3a), we see that the tail retracts smoothly (red curve), and it makes two small seesaw oscillations with the losing new arm (green curve) at the end of the process. The simulation captures these dynamic features (Fig. 3b) using a relatively low value of β = 6.7, which is only slightly larger than the critical βc = 5.95 (Fig. 2a). For the hGPC (Fig. 3f), we observe larger length variations, with the losing new arm (green curve in Fig. 3f) growing larger than the original cell tail (red curve in Fig. 3f), and the tail vanishes before it recovers temporarily with a large-amplitude seesaw oscillation. We recover these features in the simulations, as well as the much shorter escape time by using a larger value of β = 8.7 (Fig. 3g).
Extended Data Fig. 2 presents examples of hGPCs migrating along blood vessels with bifurcations in an ex vivo mouse brain slice as well as a comparison with simulations. We find the same oscillatory dynamics observed for the patterned lanes in Fig. 3a–g, suggesting the validity of our model for the cellular dynamics of in vivo physiology.
Supplementary Information section 8 provides further examples of cells migrating past the junction, which exhibit seesaw oscillations. Supplementary Information section 9 provides further quantification of the seesaw-oscillation mechanism and demonstrates that the seesaw oscillations are a robust and prevalent feature of the dynamics of both cell types when migrating over a junction.
Escape times and the metastable slow process
Next, we investigate the dependence of the average time it takes a cell to migrate past the junction (escape time, Tescape). We define the escape time as the time from when the cell first arrives to when it leaves the junction completely. We study the escape time behaviour with respect to changes in the maximal actin-polymerization speed β for a finite value of the actin response rate δ (equation (2)) and a fixed value of the cell elasticity k. For δ → ∞ (in which the actin flow instantaneously realizes its steady-state value) and for the dependence of the escape time on the cell elasticity k, see Supplementary Information section 10.
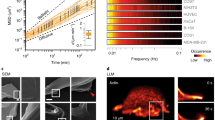
Our model predicts that above βc, the escape time decreases as β increases (black curve in Fig. 4a), an expected behaviour as faster polymerization speeds translate to faster moving cells. However, above a critical value of β (β ≈ 8.8 in Fig. 4a), a new dynamical mode appears, which is characterized by longer escape times (red curves in Fig. 4a,b). We define the cell escape over short or long times as the fast or slow process, respectively. Around the critical value (β ≈ 8.8), the probability that the cell will escape in a fast (slow) process decreases (increases) as β increases (inset in Fig. 4b). Figure 4c is the probability density function (pdf) of the escape times at β = 9, a region where the fast and the slow processes co-exist, highlighting that these processes are indeed distinct from one another.
a,b, The mean escape time as a function of β for a cell that migrates past the junction for β from 6 to 9 (a) and β from 9 to 12 (b). Red and black curves represent the fast and the slow processes, respectively. Black arrows in a denote the values of β that were used to simulate the hGPC and HUVEC in Fig. 3. The inset in b is the escape probability in the vicinity of the transition point. The grey dashed line in a indicates the \({\beta }_\mathrm{c}^\mathrm{slow}\) estimate. c, Escape time distribution for β = 9 (yellow point in a). d, Illustration of the theoretical model in which two competing arms of length l and polymerization speed v are balanced by the spring force, while the third arm is of negligible length ϵ with a local flow v1. Green and yellow gradients indicate a high or low concentration of the inhibitory polarity cue, respectively. The red at the tips of the arms indicates the actin-polymerization strength. e, Time series of the length (upper panels) and local actin flow (lower panels) for the slow process (β = 9). The panels on the right are zoom-ins of the final stage during which the balance between the long arms is resolved. Red, green and blue curves indicate arms 1, 2 and 3. f, The steady-state v2 versus v3 flow fields for the cross sections (i)–(iv) in e showing that the slow process corresponds to a local metastable solution that eventually becomes unstable. Blue points indicate the values of v1/2/3 taken from the simulation shown in e. g, Violin bar plot of the escape times of hGPCs (n = 32) and HUVECs (n = 44) in green and blue, respectively. The asterisks correspond to a two-tailed t-test where P < 0.0001. h, Time series of the arm lengths of an hGPC exhibiting the metastable slow process. Red, green and blue indicate arms 1, 2 and 3. Black indicates the total length of the cell. Grey dashed lines correspond to the time-lapse images in i (Supplementary Video 6) and to the arrows in the arm kymographs in j. Simulation parameters: c = 3.85, D = 3.85, k = 0.8, fs = 5, r = 5, κ = 20, δ = 250 and σ = 10−7. Conversion of simulation units (Supplementary Table 1): 1 time unit = 5–30 min, 1 length unit = 10–100 μm.
In the simulations we find that the slow process is characterized by two arms that extend symmetrically with very similar lengths, while the third arm shrinks to essentially zero length (as illustrated in Fig. 4d,e). The critical value of \(\beta ={\beta }_\mathrm{c}^\mathrm{slow}\) for which the slow process begins (displayed in the k–β phase diagram in Fig. 2a) is found by taking the limit at which two arms of equal length l with equal local flows v compete and balance the spring force (for the full derivation and solution of \({\beta }_\mathrm{c}^\mathrm{slow}\), see Supplementary Information section 5). Note that the slow process ends with the completion of a seesaw-oscillation event, of the same type that appears during the fast escape (Fig. 4e), and its metastable nature is shown in Fig. 4f (additional analysis of the slow mode is given in Supplementary Information section 11).
When comparing the dynamics of the two cell types across the junction (Fig. 3 and Supplementary Information sections 8 and 9), we found that the cancerous hGPCs correspond on average to a higher value of the parameter β compared to the non-cancerous HUVECs (Fig. 4g) (the noise in both cell types was fitted to be of similar amplitude). From Fig. 4a we, therefore, predict that the hGPCs should have a shorter mean escape time across the junction, compared to HUVECs (Fig. 4g). The experimental observations of the escape times of the two cell types shown in Fig. 4g are in excellent agreement with our prediction. In addition, the value of β that is typical for hGPCs places them close to the onset of the slow process (Fig. 4a). In agreement with this prediction, we find that hGPCs exhibit more often events that correspond to the slow process predicted by the model for cells with higher values of β (Fig. 4h–j). Note that in the experiments, some of these events are resolved when the cell detaches from the junction and remains stretched over the non-adhesive region between the two competing arms. This detachment is beyond the scope of the model.
The effect of cytoskeleton-modifying drugs
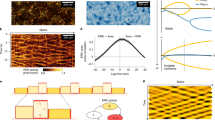
To further test our model, we treated the cells with different drugs that affect the acto-myosin cytoskeleton and quantified their migration patterns along linear one-dimensional tracks and across the Y junction (Fig. 5). Both hGPCs and HUVECs were treated with three drugs: (1) Latrunculin A (Lat. A), which inhibits actin polymerization with the highest turnover. That is, it mainly inhibits lamellipodia whereas the stress fibres are less affected36,37. In the model, the addition of Lat. A is, therefore, equivalent to a decrease in the maximal polymerization speed β (blue in Fig. 5). (2) Cytochalasin D (Cyto. D), which acts mainly as a cap** protein targeting the barbed ends (the plus end or the fast-polymerizing end) of actin filaments. This drug, therefore, affects the cortical actin layer below the cell membrane and the stress fibres, thereby softening the cell in addition to inhibiting protrusion38. In the model, the addition of Cyto. D is, therefore, equivalent to reducing both the overall cell elasticity k and the maximal polymerization speed β (green in Fig. 5). (3) Blebbistatin (Blebb.), which is an inhibitor of myosin II (refs. 39,40), reduces the overall contractility of the cell, making the cell softer. In the model, the Blebb. treatment is equivalent to a reduction of the mean cell elastic stiffness k (red in Fig. 5). Note that myosin II inhibition can also affect the speed of the actin retrograde flow (our parameter β), but in these experiments this effect seems to be less important.
a–c, Cell length (a), speed (b) and escape time (c) were measured in different conditions: control (black), Lat. A (blue), Cyto. D (green) and Blebb. (red) at the indicated concentrations (μM). a, Cell length (μm) during one-dimensional migration measured on the linear part of the patterns for hGPCs (n = 18, 14, 26, 18, 22, 21 and 16 from left to right) (i) and HUVECs (n = 27, 16, 23 and 19) (ii). β–k simulation heat map (iii). Blue, green and red arrows display qualitatively the effect of each drug treatment in terms of the model’s β–k parameter space. b, Velocity (μm min−1) during one-dimensional migration measured for hGPCs (n = 18, 13, 26, 16, 21, 19 and 23) (i) and HUVECs (n = 27, 16, 23 and 19) (ii). β–k simulation heat map (iii). c, Junction escape time (hours) for a symmetric junction measured for hGPCs (n = 16, 19, 8, 14, 10, 27 and 25) (i) and HUVECs (n = 27, 16, 23 and 19) (ii). β–k simulation heat map (iii). Asterisks ****, ***, ** and * and NS indicate the results of a one-way ANOVA test with a Tukey multiple comparison test: P < 0.0001, 0.001, 0.01 and 0.05 and P ≥ 0.05 (not significant), respectively. csv files with the statistical tests are included in the Supplementary Information.
The results show that for both hGPC and HUVEC, there are similar qualitative changes to the cell’s average length and speed on the linear tracks and escape time across the junction, due to the drugs. Lat. A induces a decrease in the length and speed of the cells, while the escape time increases. Cyto. D treatment decreases the length and speed (by a lesser amount than for Lat. A) and increases the escape time substantially. Blebb. treatment increases the cell length and the escape time, while the velocity remains roughly unchanged. Note that the Lat. A and Cyto. D treatments for the HUVECs did not induce notable changes in the cell length in comparison to hGPCs (Fig. 5a(i,ii)), and that although the Blebb. treatment did not induce a notable increase in cell length and escape time for the hGPCs, there are larger variations of cell length and escape time compared to the controls (Fig. 5a,c(i)). Comparing these experimental observations to the model, we indicate by arrows in Fig. 5a(iii)–c(iii) the qualitative changes to the parameters β and k that each drug is expected to induce and how these changes affect the cell length and speed on a linear track and the escape time across a junction. We see that the model predicts the same qualitative changes as observed in the experiments, for all three dynamical properties of the cells.
Note that as the concentration of the drugs was increased, more cells became trapped in the junction during the entire duration of the experiment (Supplementary Information section 12). HUVECs exposed to high concentrations of all three types of drugs exhibited mostly non-motile phenotype or extensive cell death.
Effect of noise on cell escape and cell trap**
In our analysis we also found that the escape time highly depends on the amplitude of the noise in the local actin-polymerization activity σ (equation (2)). This is demonstrated in Fig. 6a for a moving cell in the stick–slip migration regime (β = 8). For a low noise level (Fig. 6a(i)), the escape time is sharply defined and the cell always leaves the junction along either of the new arms.
a, Escape time probability density function (left y axis in blue) and the probability that the cell will leave the junction along one of the arms (right y axis in orange) for β = 8 and noise amplitudes (equation (2)): σ = 0.4 (i), σ = 1.2 (ii) and σ = 1.6 (iii). Red indicates the probability to turn back (arm 1). Blue and green indicate the probability to take a new path (arms 2 and 3 respectively) b, Example of a simulation length time series with σ = 1.6. Red, green and blue indicate arms 1, 2 and 3. c–h, hGPC and HUVEC experiments showing cells that are trapped at the junction. c,f, Time-lapse images of hGPCs (c, Supplementary Video 7) and HUVECs (f, Supplementary Video 8). d,g, Arm kymographs for hGPCs (d) and HUVECs (g). Red, green and blue arrows indicate the timestamps that correspond to the time-lapse images. e,h, Experimental time series of the area of the cellular arms for hGPCs (e) and HUVECs (h). Grey dashed lines correspond to the time-lapse images. Red, green and blue indicate arms 1, 2 and 3. Black indicate the total cell length. Parameters: c = 3.85, D = 3.85, k = 0.8, fs = 5, r = 5, κ = 20 and δ = 250. Conversion of simulation units (Supplementary Table 1): 1 time unit = 5–30 min, 1 length unit = 10–100 μm.
As the noise amplitude increases, the escape time increases (Fig. 6a(ii)). In particular, the noise gives rise to additional oscillations during the migration past the junction. Each additional oscillation increases the escape time, which has several peaks. A single additional oscillation reverses the direction in which the cell leaves the junction, increasing the probability that it will ‘reflect’ (leave along the direction from which it arrived). For a large number of oscillations, the cell has equal probabilities of leaving along any of the three arms. This is also observed for a high noise level (Fig. 6a(iii)), for which any memory of the original direction of motion is lost and the escape time distribution becomes exponential. In this limit, we often observe cells that are trapped at the junction for long periods of time (Fig. 6b).
The increased incidence of cells being trapped at a junction at higher noise is unusual at first sight, as noise is normally attributed with triggering the escape of particles from traps41,42. However, for the cell to leave the junction, it needs to maintain its polarity for the duration of the escape time, and the high noise diminishes the persistence time of the polarization, inhibiting the escape probability. Nevertheless, the noise also plays a role in aiding the cell’s escape, as during the slow process the metastable configuration is destabilized (Fig. 4).
Figure 6 has examples of hGPCs and HUVECs experiencing many cycles of elongation and retraction of the arms. They are unable to maintain a persistent polarization and, therefore, remain trapped at the junction. Both examples correspond to the large noise limit of our model (Fig. 6b). Further analysis of the effects of noise on the dynamics at the junction is given in Supplementary Information section 13. Extended Data Fig. 3 shows an example of a cell that reflects at the junction, as occurs in our model when the actin flow is subjected to intermediate values of noise (Fig. 6a(ii)). Further quantification of the number of seesaw oscillations in the two cell types as they cross the junction and a comparison with the model are given in Supplementary Information section 9.
Discussion
We have presented here a new theoretical model for the directional decision-making process at the level of a single migrating cell as it moves past a symmetric Y junction. Our model describes the spontaneous symmetry breaking during cellular spreading over a junction and predicts deterministic oscillations during the migration of a cell past the junction. These oscillations arise due to the competition for the finite resource, the polarity cue, which is redistributed over the arms as their lengths evolve. As a result, the actin-polymerization activity oscillates between the competing leading edges. These out-of-phase oscillations are most prominent between the two ‘losing’ arms. Oscillations are also induced in the length dynamics of the cellular protrusions. This is a new form of deterministic oscillation during cell migration, being distinct from oscillations induced by stick–slip adhesion20,43 or periodic macropinocytosis44.
These surprising theoretical predictions were validated by experimental data acquired from two cell types, cancerous hGPCs and non-cancerous HUVECs. The predicted out-of-phase oscillations in the lamellipodia activity at the leading edges of the cellular protrusions and in the protrusion lengths were observed in these experiments when these cells migrate past a junction. These comparisons show that non-cancerous (HUVECs) and cancerous cells (hGPCs) have low or high levels of motility regimes, respectively.
The model predicts that cells with low actin-polymerization activity will be prone to remaining trapped at junctions, even if they are well polarized and motile when moving along simple linear tracks. As expected, the mean trap** time at a junction decreases as the cells become more motile (higher actin-polymerization activity). Higher noise in the actin-polymerization activity acts to increase the number of polarization oscillations and the overall trap** time at a junction. Interestingly, when the polymerization activity is high, the cells may be transiently trapped at the junction, due to the competition between two symmetric and highly elongated protrusions. The latter suggest that efficient migration across a complex geometry and the resolution of this directional dilemma occurs within a range of actin polymerization that is sufficiently strong to allow motility but weak enough to prevent the non-productive consequence of excessive acto-myosin interactions. In further agreement with the predictions of our model, treatment with Lat. A, Cyto. D or Blebb. decreased the cell speed and consequently increased the escape time from the junction. These predictions will be the subject of future experimental studies. Future extensions of the model might include asymmetric junctions8, different geometries and the interaction with other cells to evaluate the effect of cell-to-cell interactions45,46. In addition, this model could be used to explore the effects of chemotaxis, haptotaxis and barotaxis on cellular decision-making and migration. Moreover, it will be interesting to explore how these external gradients affect the seesaw-oscillation mechanism that emerges in our model.
To conclude, we have presented a detailed, yet simple, model for the cellular mechanism of directional decision-making at a Y junction. In this model the decision is performed due to spontaneous symmetry breaking involving deterministic oscillations. The model explains how the duration of the directional decision-making depends on the internal motility parameters of the cell, offering a natural explanation for the puzzling slowness of this process in many cells18.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Statistical data are available from the corresponding author upon request.
Code availability
A minimal solver code for the model is included in the Supplementary Information. Custom-built ImageJ macros and R scripts are available from the corresponding author upon request.
References
Yamada, K. M. & Sixt, M. Mechanisms of 3D cell migration. Nat. Rev. Mol. Cell Biol. 20, 738–752 (2019).
SenGupta, S., Parent, C. A. & Bear, J. E. The principles of directed cell migration. Nat. Rev. Mol. Cell Biol. 22, 529–547 (2021).
Phillipson, M. & Kubes, P. The neutrophil in vascular inflammation. Nat. Med. 17, 1381–1390 (2011).
Moreau, H. D. et al. Macropinocytosis overcomes directional bias in dendritic cells due to hydraulic resistance and facilitates space exploration. Dev. Cell 49, 171–188 (2019).
Renkawitz, J. et al. Nuclear positioning facilitates amoeboid migration along the path of least resistance. Nature 568, 546–550 (2019).
Wang, X., Hossain, M., Bogoslowski, A., Kubes, P. & Irimia, D. Chemotaxing neutrophils enter alternate branches at capillary bifurcations. Nat. Commun. 11, 2385 (2020).
Zhao, R. et al. Cell sensing and decision-making in confinement: the role of trpm7 in a tug of war between hydraulic pressure and cross-sectional area. Sci. Adv. 5, eaaw7243 (2019).
Monzo, P. et al. Adaptive mechanoproperties mediated by the formin fmn1 characterize glioblastoma fitness for invasion. Dev. Cell 56, 2841–2855 (2021).
Paul, C. D., Mistriotis, P. & Konstantopoulos, K. Cancer cell motility: lessons from migration in confined spaces. Nat. Rev. Cancer 17, 131 (2017).
Georgantzoglou, A., Poplimont, H., Walker, H. A., Lämmermann, T. & Sarris, M. A two-step search and run response to gradients shapes leukocyte navigation in vivo. J. Cell Biol. 221, e202103207 (2022).
Dai, W. et al. Tissue topography steers migrating Drosophila border cells. Science 370, 987–990 (2020).
Kameritsch, P. & Renkawitz, J. Principles of leukocyte migration strategies. Trends Cell Biol. 30, 818–832 (2020).
Farin, A. et al. Transplanted glioma cells migrate and proliferate on host brain vasculature: a dynamic analysis. Glia 53, 799–808 (2006).
Ambravaneswaran, V., Wong, I. Y., Aranyosi, A. J., Toner, M. & Irimia, D. Directional decisions during neutrophil chemotaxis inside bifurcating channels. Integr. Biol. 2, 639–647 (2010).
Pham, Q. L. et al. Cell sequence and mitosis affect fibroblast directional decision-making during chemotaxis in microfluidic mazes. Cell. Mol. Bioengin. 11, 483–494 (2018).
Um, E. et al. Immature dendritic cells navigate microscopic mazes to find tumor cells. Lab Chip 19, 1665–1675 (2019).
Belotti, Y., McGloin, D. & Weijer, C. J. Analysis of barotactic and chemotactic guidance cues on directional decision-making of dictyostelium discoideum cells in confined environments. Proc. Natl Acad. Sci. USA 117, 25553–25559 (2020).
Hadjitheodorou, A. et al. Leading edge competition promotes context-dependent responses to receptor inputs to resolve directional dilemmas in neutrophil migration. Cell Syst. 14, 196–209 (2023).
Maiuri, P. et al. Actin flows mediate a universal coupling between cell speed and cell persistence. Cell 161, 374–386 (2015).
Ron, J. E., Monzo, P., Gauthier, N. C., Voituriez, R. & Gov, N. S. One-dimensional cell motility patterns. Phys. Rev. Res. 2, 033237 (2020).
Wortel, I. M. et al. Local actin dynamics couple speed and persistence in a cellular Potts model of cell migration. Biophys. J. 120, 2609–2622 (2021).
Fonseca, C. G., Barbacena, P. & Franco, C. A. Endothelial cells on the move: dynamics in vascular morphogenesis and disease. Vascul. Biol. 2, H29–H43 (2020).
Winkler, F. et al. Imaging glioma cell invasion in vivo reveals mechanisms of dissemination and peritumoral angiogenesis. Glia 57, 1306–1315 (2009).
Stramer, B. & Mayor, R. Mechanisms and in vivo functions of contact inhibition of locomotion. Nat. Revi. Mol. Cell Biol. 18, 43–55 (2017).
Chung, J., Goode, B. L. & Gelles, J. Single-molecule analysis of actin filament debranching by cofilin and gmf. Proc. Natl Acad. Sci. USA 119, e2115129119 (2022).
Cai, L., Marshall, T. W., Uetrecht, A. C., Schafer, D. A. & Bear, J. E. Coronin 1b coordinates arp2/3 complex and cofilin activities at the leading edge. Cell 128, 915–929 (2007).
Law, A.-L. et al. Nance-Horan syndrome-like 1 protein negatively regulates scar/wave-arp2/3 activity and inhibits lamellipodia stability and cell migration. Nat. Commun. 12, 5687 (2021).
Olguin-Olguin, A. et al. Chemokine-biased robust self-organizing polarization of migrating cells in vivo. Proc. Natl Acad. Sci. USA 118, e2018480118 (2021).
Ivaska, J. Unanchoring integrins in focal adhesions. Nat. Cell Biol. 14, 981–983 (2012).
Ren, X.-D. et al. Focal adhesion kinase suppresses rho activity to promote focal adhesion turnover. J. Cell Sci. 113, 3673–3678 (2000).
Katz, Z. B. et al. β-actin mRNA compartmentalization enhances focal adhesion stability and directs cell migration. Genes Dev. 26, 1885–1890 (2012).
Berginski, M. E., Vitriol, E. A., Hahn, K. M. & Gomez, S. M. High-resolution quantification of focal adhesion spatiotemporal dynamics in living cells. PLoS ONE 6, e22025 (2011).
Gupton, S. L. & Waterman-Storer, C. M. Spatiotemporal feedback between actomyosin and focal-adhesion systems optimizes rapid cell migration. Cell 125, 1361–1374 (2006).
Rossier, O. et al. Integrins β1 and β3 exhibit distinct dynamic nanoscale organizations inside focal adhesions. Nat. Dell Biol. 14, 1057–1067 (2012).
Doedel, E. J. et al. AUTO97: Continuation and Bifurcation Software for Ordinary Differential Equations (with HomCont). Technical Report, Concordia University (1997).
Gronewold, T. M., Sasse, F., Lünsdorf, H. & Reichenbach, H. Effects of rhizopodin and latrunculin B on the morphology and on the actin cytoskeleton of mammalian cells. Cell Tissue Res. 295, 121–129 (1999).
Spector, I., Shochet, N. R., Kashman, Y. & Groweiss, A. Latrunculins: novel marine toxins that disrupt microfilament organization in cultured cells. Science 219, 493–495 (1983).
Efremov, Y. M. et al. Distinct impact of targeted actin cytoskeleton reorganization on mechanical properties of normal and malignant cells. Biochim. Biophys. Acta Mol. Cell Res. 1853, 3117–3125 (2015).
Straight, A. F. et al. Dissecting temporal and spatial control of cytokinesis with a myosin II inhibitor. Science 299, 1743–1747 (2003).
Limouze, J., Straight, A. F., Mitchison, T. & Sellers, J. R. Specificity of blebbistatin, an inhibitor of myosin II. J. Muscle Res. Cell Motil. 25, 337–341 (2004).
Wexler, D., Gov, N., Rasmussen, K. Ø. & Bel, G. Dynamics and escape of active particles in a harmonic trap. Phys. Rev. Res. 2, 013003 (2020).
Woillez, E., Kafri, Y. & Gov, N. S. Active trap model. Phys. Rev. Lett. 124, 118002 (2020).
Mukherjee, A. et al. Actin filaments couple the protrusive tips to the nucleus through the I-BAR domain protein IRSp53 for migration of elongated cells on 1D fibers. Adv. Sci. 10, 2207368 (2022).
Lavi, I., Piel, M., Lennon-Duménil, A.-M., Voituriez, R. & Gov, N. S. Deterministic patterns in cell motility. Nat. Phys. 12, 1146–1152 (2016).
Carmona-Fontaine, C. et al. Contact inhibition of locomotion in vivo controls neural crest directional migration. Nature 456, 957 (2008).
Scarpa, E. et al. A novel method to study contact inhibition of locomotion using micropatterned substrates. Biolo. Open 2, 901–906 (2013).
Polleux, F. & Ghosh, A. The slice overlay assay: a versatile tool to study the influence of extracellular signals on neuronal development. Sci. STKE 2002, pl9 (2002).
Acknowledgements
J.E.R. and N.S.G. acknowledge useful discussions with R. Voituriéz and A. Yochelis. N.S.G. is the incumbent of the Lee and William Abramowitz Professorial Chair of Biophysics and acknowledges support from the Israel Science Foundation (Grant No. 1459/17). This research is made possible in part by the historic generosity of the Harold Perlman Family. This work was supported by the Human Frontier Science Program (Grant No. RGP0032/2022 to P.J.S. and N.S.G.), the German Research Foundation (Grant No. 335447717-SFB 1328 under Project A20) and Forschungszentrum Medizintechnik Hamburg (Grant No. 04fmthh2021 to P.J.S.). This work was supported by IFOM (starting package), the Mechanobiology Institute of Singapore (Grant No. WBSR-714-016-007 -271) and the Italian Association for Cancer Research (Investigator Grant Nos. 20716 and 27101 to N.C.G. and a 3-yr doctoral fellowship MilanoMarathon-oggicorroperAIRC, ref. 22461, to M.C.). M.C. is a PhD student within the European School of Molecular Medicine.
Funding
Open access funding provided by Universitätsklinikum Hamburg-Eppendorf (UKE).
Author information
Authors and Affiliations
Contributions
J.E.R. and N.S.G. designed the analytical model. J.E.R and J.L. performed the model analysis and simulations. M.C. prepared the data analysis automation macro. J.E.R., M.C., J.M.K. and J.L. performed the data analysis. M.C. and P.M. prepared the micro-patterned tracks and performed the hGPC experiments. N.C.G. supervised the hGPC experiments. J.M.K. also prepared the micro-patterned tracks. J.M.K. and N.A.D. performed the HUVEC experiments. P.J.S. supervised the HUVEC experiments. J.E.R. prepared the manuscript. N.S.G, N.C.G. and P.J.S. supervised the research and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Physics thanks Rhoda Hawkins and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Seesaw oscillation analysis for the symmetric case.
A) Time series of the (i) length, (ii) the local polymerization speed (local flow) and (iii) the arm ratio. Red/Green/Blue colors in (i,ii) correspond to arms 1/2/3 respectively. Blue and Orange curves in (iii) correspond to the arm ratios l1/l3 and l2/l3 respectively. Black dashed lines indicate the examined cross-sections of the first and second cycles of oscillations. B-C) The solution branches of the local flows as a function of the winning arm l3 in the cross-sections of the first and second cycle respectively. (i)/(ii)/(iii) correspond to v1/v2/v3. Left/Right panels correspond to the solutions branches before/after the transition. Points indicate the values from the simulations in (A,iii). Parameters: δ = 250, β = 6.5, c = 3.85, D = 3.85, k = 0.8, fs = 5, r = 5, κ = 20, σ = 10−7 (noise amplitude).
Extended Data Fig. 2 hGPC branching along blood vessels on a mouse brain slice explant.
A) Time-lapse images of an hPGC cell migrating in a mouse brain slice and chooses a new path when encountering a bifurcation along a blood vessel (movie E1). White dashed lines indicate the blood vessels over which the cell migrates upon. B) Comparison between the time-series of the arms in (i) the experiment and (ii) a simulation. C) Kymographs of the arms of the cell in the blood vessel bifurcation. D) Time-lapse images of an hPGC cell migrating in a mouse brain slice and reflects back when encountering a bifurcation along a blood vessel (movie E2). White dashed lines indicate the blood vessels over which the cell migrates upon. E) B) Comparison between the time-series of the arms in (i) the experiment and (ii) a simulation. F) Kymographs of the arms of the cell in the blood vessel bifurcation. Scale bars in (A,D) are 50 [μm]. Simulation parameters: β = 8.5,σ = 1.2,c = 3.85, D = 3.85, k = 0.8, fs = 5, r = 5, κ = 20, δ = 250. Simulation units conversion (Supplementary Table 1: 1 time unit = 5–30 [min], 1 length unit = 10-100 [μm].
Extended Data Fig. 3 Comparison between a HUVEC experiment and simulation of a cell reflecting back from the junction.
A) Time-lapse images (movie E3). B) Arm kymograph. Red/Green/Blue arrows indicate the time stamps which correspond to the images in A. C) Time series of the areas of the arms. vertical dashed lines correspond to the images in A. D) Simulation time series of the arm lengths with (β, σ) = (6.9, 0.8). Red/Green/Blue indicate arm 1/2/3. Black indicates the total length of the cell. Time-lapse images correspond to movie S7. Other simulation parameters: c = 3.85, D = 3.85, k = 0.8, fs = 5, r = 5, κ = 20, δ = 250. Simulation units conversion (see SI Table S-1): 1 time unit = 5-30 [min], 1 length unit = 10-100 [μm].
Supplementary information
Supplementary Information
Supplementary information pdf file, which includes 15 sections, 20 figures and two tables.
Supplementary Video 1
Human glioma propagating cells (hGPCs) migrating in a mouse brain slice (Fig. 1a).
Supplementary Video 2
Example of an hGPC migrating along a laminin-coated Y junction (Fig. 1b).
Supplementary Video 3
Example of a HUVEC migrating along a fibronectin-coated Y junction (Fig. 1b).
Supplementary Video 4
HUVEC experiment showing out-of-phase oscillations in arm length and actin activity (Fig. 3d).
Supplementary Video 5
hGPC experiment showing out-of-phase oscillations in arm length and actin activity (Fig. 3i).
Supplementary Video 6
hGPC experiment, where two competing arms extend, as in the slow process (Fig. 4i).
Supplementary Video 7
hGPC experiment, where the cell is not deciding on a new direction of motion (Fig. 6c).
Supplementary Video 8
HUVEC experiment in which the cell is not deciding on a new direction of motion (Fig. 6f).
Supplementary Video 9
hGPC migrating over a blood vessel bifurcation and deciding on a new path (Extended Data Fig. 2a).
Supplementary Video 10
hGPC migrating over a blood vessel bifurcation and reflecting back (Extended Data Fig. 2d).
Supplementary Video 11
HUVEC experiment showing a cell that reflects back in the direction from which it has entered the junction (Extended Data Fig. 3a).
Supplementary Video 12
HUVEC experiment showing out-of-phase oscillations in arm length and actin activity (Supplementary Fig. 7d).
Supplementary Video 13
HUVEC experiment showing out-of-phase oscillations in arm length and actin activity (Supplementary Fig. 7h).
Supplementary Video 14
HUVEC experiment showing out-of-phase oscillations in arm length and actin activity (Supplementary Fig. 7l).
Supplementary Video 15
hGPC experiment showing an unpolarized cell that resides in the centre of the junction (after a stick–slip event). All protruding arms spread symmetrically (Supplementary Fig. 8d).
Supplementary Video 16
hGPC experiment showing out-of-phase oscillations in arm length and actin activity (Supplementary Fig. 8h).
Supplementary Video 17
HUVEC experiment showing a normal escape pattern (Supplementary Fig. 9a).
Supplementary Dataset 1
ANOVA test data for the drug experiments shown in Supplementary Information section 12.
Supplementary Code File 1
Matlab three-way junction simulation numerical solver.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ron, J.E., Crestani, M., Kux, J.M. et al. Emergent seesaw oscillations during cellular directional decision-making. Nat. Phys. 20, 501–511 (2024). https://doi.org/10.1038/s41567-023-02335-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41567-023-02335-6
- Springer Nature Limited