Abstract
The applications of self-assembled InAs/GaAs quantum dots (QDs) for lasers and single photon sources strongly rely on their density and quality. Establishing the process parameters in molecular beam epitaxy (MBE) for a specific density of QDs is a multidimensional optimization challenge, usually addressed through time-consuming and iterative trial-and-error. Here, we report a real-time feedback control method to realize the growth of QDs with arbitrary density, which is fully automated and intelligent. We develop a machine learning (ML) model named 3D ResNet 50 trained using reflection high-energy electron diffraction (RHEED) videos as input instead of static images and providing real-time feedback on surface morphologies for process control. As a result, we demonstrate that ML from previous growth could predict the post-growth density of QDs, by successfully tuning the QD densities in near-real time from 1.5 × 1010 cm−2 down to 3.8 × 108 cm−2 or up to 1.4 × 1011 cm−2. Compared to traditional methods, our approach can dramatically expedite the optimization process and improve the reproducibility of MBE. The concepts and methodologies proved feasible in this work are promising to be applied to a variety of material growth processes, which will revolutionize semiconductor manufacturing for optoelectronic and microelectronic industries.
Similar content being viewed by others
Introduction
Self-assembled quantum dots (QDs) have attracted great interest due to their applications in various optoelectronic devices1. The so-called Stranski-Krastanow (SK) mode in molecular beam epitaxy (MBE) is widely used for growing these high-quality QDs2. For specific applications, such as QD lasers, high QD densities are required; while low-density QDs are necessary for other applications, such as single photon sources1. However, the outcomes of any QD growth process are a complex function of a large number of variables including the substrate temperature, III/V ratio, and growth rate, etc. Building a comprehensive analytical model that describes complex physical processes occurring during growth is an intractable problem3. The optimization of material growth largely depends on the skills and experience of MBE researchers. Time-consuming trial-and-error testing is inevitably required to establish optimal process parameters for the intended material specification. It is needed to build a real-time connection between in situ characterization and material growth status.
Machine learning (ML) is revolutionary due to its exceptional capability for pattern recognition and its potential in approximating the empirical functions of complex systems. It enables researchers to extract valuable insights and identify hidden patterns from large datasets, leading to a better understanding of complex phenomena required to build predictive models, generate new hypotheses, and determine optimal growth conditions for MBE4,5,6,7,8. It offers an alternative approach where the growth outcomes for an arbitrary set of parameters can be predicted via a trained neural network, which has been applied to extract film thickness and growth rate information9,10. Moreover, by enabling the direct adjustment of parameters during material growth, ML-based in situ control can detect and correct any deviation from expected values in a timely manner11,12,13. Meng et al. have utilized a feedback control to adjust cell temperatures after an amount of film being deposited, thereby capable of regulating AlxGa1-xAs compositions in situ13. However, these remain posteriori ML-based approaches, since they require the completion of the growth10. Therefore, once the sample characterization results deviate from expectation, it becomes challenging and time-consuming to identify reasons11.
To fully exploit the effectiveness of real-time connection based on ML, an in situ characterization with continuous and transient working mode is highly desirable to unveil the material status. Reflection high-energy electron diffraction (RHEED) has been widely used to capture a wealth of growth information in situ14. However, it still faces the challenges of extracting information from noisy and overlap** images. Traditionally, identifying RHEED patterns during material growth was mainly depended on the experience of growers. Progress has been made in the automatic classification of RHEED patterns through ML that incorporates principal component analysis (PCA) and clustering algorithms9,14,15,16,17. It surpasses human analysis when a single static RHEED image is gathered with the substrate held at a fixed angle16,18. However, this may result in an incomplete utilization of the temporal information available in RHEED videos taken with the substrate rotating continuously16,19,20. Although few researchers have highlighted the importance of analyzing RHEED videos, their ML models were still based on PCA and encoder-decoder architecture with a single image input16,21. To date, it is still hard to build a real-time connection between in situ characterization and material growth status, which is a prospective way to realize the precise control on material specifications.
In this work, we proposed a real-time feedback control based on ML to connect in situ RHEED videos and QD density, automatically and intelligently achieving a low-cost, efficient, reliable, and reproduceable growth of QDs with required density. By investigating the temporal evolution of RHEED information during the growth of InAs QDs on GaAs substrates, the real-time status of the sample was better reflected. We applied two structurally identical 3D ResNet 50 models in ML deployment, which takes three-dimensional variables as inputs to determine the QD formation and classify density22,23. We showed that ML can simulate and predict the post-growth material specifications, and based on which, growth parameters are continuously monitored and adjusted. As a result, the QD densities were successfully tuned from 1.5 × 1010 cm−2 down to 3.8 × 108 cm−2, and up to 1.4 × 1011 cm−2, respectively. The methodology can also be adapted to other QD systems, such as the droplet QDs, the local droplet etching, and manhole infilling QDs with distinct RHEED features24,25,26,27. The effectiveness of our approach by taking full advantages of in situ characterization marks a significant achievement of establishing a precise growth control scheme, with the capabilities and potentials of being extended to large-scale material growth, reducing the impact on material growth due to instability and uncertainty of MBE operation, shortening the parameter optimization cycle, and improving the final yield of material growth.
Results
Sample structure and data labeling
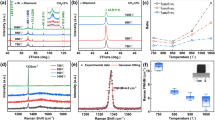
Our method relied on an existing database of growth parameters with corresponding QD characteristics, which helped us decide the growth parameters needed to achieve a target density for QDs. The sample structure is shown in Fig. 1a, with density results of 30 QD samples summarized in Table 1, which are correlated with the RHEED videos. Each QD growth was repeated on average 4 times, leading to the generation of 120 RHEED videos. The samples grown without QDs have flat surfaces determined by an atomic force microscope (AFM) as shown in Fig. 1b, corresponding to streaky RHEED patterns in Fig. 1f, which was artificially labeled with “zero” referring to the density of QDs on the surface27. With the increase of QD density, it was found that the RHEED gradually converted from streaky to spotty patterns. The QDs density of ~2.9 × 109 cm−2 from Fig. 1c corresponds to the RHEED pattern consisted of streaks and spots in Fig. 1g27. As shown in Fig. 1d, h, with the further increase of QDs density reaching 1.9 × 1010 cm−2, the arrangement of RHEED pattern showed typical spot features28,29,30. Upon conducting a more thorough analysis of the RHEED patterns, we identified the patterns transition from having both streaks and spots to only spots, corresponding to a density of 1 × 1010 cm−2 in AFM images. So, we assigned the “low” label to densities below 1 × 1010 cm−2, the “middle” label to densities ranging from 1 × 1010 cm−2 to 4 × 1010 cm−2. As the density exceeds 4 × 1010 cm−2, the spot features become more rounded and show higher brightness, we labeled densities exceeding 4 × 1010 cm−2 as “high” (see Supplementary Information for RHEED characteristics of QDs with different labels, S1)31,32,33. Fig. 1e, i show the typical AFM and RHEED images after the QD formation, respectively, with the QD density of ~1.0 × 1011 cm−2. It is worth noting that we keep the labels to the RHEED videos the same before and after the QD formation. The RHEED information before the QD formation can be used to determine the resulting QD density grown under the same conditions, enabling us to adjust the material growth parameters before the QD formation.
a The schematic of the sample structure. The thick GaAs buffer layer is grown on an n-type GaAs substrate at 600 °C. Subsequently, the substrate is cooled to 450–500 °C for the growth of InAs QDs. b–e The 2 μm × 2 μm atomic force microscope (AFM) images of QDs with varied labels. “zero”: there are no QDs on the surface. “low”: the InAs QDs sample with density less than 1 × 1010 cm−2. “middle”: the InAs QDs sample with density range from 1 × 1010 cm−2 to 4 × 1010 cm−2. “high”: the InAs QDs sample with density higher than 4 × 1010 cm−2. f–i Corresponding reflection high-energy electron diffraction (RHEED) images with varied labels.
Program framework and video processing
The design framework of our program is illustrated in Fig. 2a. An appropriate scheme was designed to pre-process the RHEED video data. Then, a ML model was selected based on the pre-processing results. Furthermore, we also determined the way to adjust parameters according to output results of the model.
a The framework of the program. An electron beam generated by an electron gun is continuously applied to the rotating substrate, creating a diffraction image frame by frame on the fluorescent screen. Multiple images are then converted to a new matrix, and the reshaped matrix is then transferred to the model for generation of results to guide the adjustment of material growth parameters. b A typical reflection high-energy electron diffraction (RHEED) image taken from the camera and the crop** area. The area marked by the blue square is an effective selection area that the software must handle and subsequently provide as input to the model. c Continuous sampling method for RHEED images. The software processes images with data at position T during each iteration. T is comprised of several sub-images taken sequentially from time t, including t-1, t-2, and so forth. Similarly, the data positions processed by the software in the previous iteration are T-1, T-2, and so on. d Processing method for sampled images. The data at a specific position T is initially acquired. Subsequently, each data point within T, denoted as t, t-1, t-2, and so forth, is transformed into a 2D matrix containing only black and white channels. Finally, these individual 2D matrices are stitched into a 3D matrix.
RHEED videos taken during growth were first deconstructed into multiple temporal images and utilized as input for our model, which is a technique for breaking down video processing, offering a more versatile and efficient approach to analyze and process information in dynamic scenes34. The original image collected has uncompressed 4 channels of 8-bit depth color with a resolution of 1920 × 1200. The software cropped a square area from the image, as shown in Fig. 2b, immediately compressed to 300 × 300 by zero-order sampling and converted it to a single 4 × 300 × 300 matrix (see Supplementary Information for the crop** area of RHEED images, S2). To efficiently utilize the temporal information within RHEED data, we used the latest RHEED image as a starting point, acquired an additional several consecutive images before, and bundled as the original RHEED dataset for the model, as shown in Fig. 2c35. Subsequently, each image was converted to a single-channel 8-bit grayscale image based on the luminance information16,21, and stitched into an N × 300 × 300 3D matrix through shift registers, as shown in Fig. 2d. Additionally, we modified the convolutional filtering order. Traditional Convolutional Neural Networks (CNNs) tend to uniformly sample color channels, which is effective for images with rich color information, but not suitable for our data36,37. To address this issue, we designed an image processing approach that uses longitudinal size information instead of color channel information, which can improve the correlation between channels, reduce unnecessary calculations, and improve data processing efficiency38,39,40. Therefore, the size of the preprocessed data was N × 300 × 1 × 300, representing the image number, image width, image color channel, and image length. We then compared the impact of choosing different numbers of images on model training accuracy and speed, ultimately determined that selecting 8 images was the appropriate choice (see Supplementary Information for the number of images selected and speed management, S3 and S4). Finally, we conducted preprocessing for all data before model training to reduce the time. We have also applied data augmentation techniques to the training data, including adjusting image curves, crop** images, and scaling images, which effectively increase data diversity and improve the model’s generalization ability. This process resulted in the acquisition of ~360,000 NumPy arrays with each sample ~704 KB in size.
Model construction and evaluation
The ResNet 50 model has emerged as our preferred choice for several reasons, including its capability to address gradient vanishing issues and automatically learn methods for extracting features from raw data without manual feature engineering or dimensionality reduction, rapid convergence, and suitability for deep training. When processing video frame data, 3D convolution proves more effective in extracting spatial and temporal dimension information from videos than the 2D convolution used in the standard ResNet model. The difference between our network and the original ResNet lies in the dimensionality of the convolutional kernel and batch normalization operations23. Our 3D ResNet 50 model employs 3D convolutions and 3D batch normalization. This spatiotemporal-feature-learning method enables the 3D ResNet 50 model to better capture the dynamic information in videos compared to 2D CNN, resulting in an enhanced ability to identify and differentiate the categories of videos53. A C-type thermocouple measured the substrate temperature, and the growth rates were calibrated through the RHEED oscillations of extra layers grown on GaAs substrates. The substrate was outgassed at 350 °C in the buffer chamber, then heated to 620 °C in the growth chamber for deoxidation. The substrate was then cooled to 600 °C for the growth of the GaAs buffer layer. The sample structure is shown in Fig. 1a. The BEP of In was 1.4–1.5 × 10-8 Torr, while the BEP of As was 0.95–1.0 × 10-7 Torr. The QDs were formed in the SK growth mode after depositing 1.7 ML of InAs at a temperature of 450–500 °C and a rate of 0.01 ML/s. The initial growth showed a relatively flat planar structure, pertaining to the wetting layer growth54,55,56,57,58,59. The formation of InAs QDs was verified by the streaky-spotty RHEED pattern transition60.
Material characterization
RHEED in the MBE growth chamber enabled us to analyze and monitor the surface of the epilayer during the growth. RHEED patterns were recorded at an electron energy of 12 kV (RHEED 12, from STAIB Instruments). A darkroom equipped with a camera was placed outside the chamber to continuously collect RHEED videos with the substrate rotating at 20 revolutions per minute. The exposure time was 100 ms, with a sampling rate of 8 frames per second. To achieve a clear correlation of RHEED with QD density, we also characterized the surface morphology of the InAs QDs using AFM (Dimension Icon, from Burker).
Hardware wiring scheme
Our model is deployed on a Windows 10 computer system with an AMD R9 7950X CPU, 64GB of memory, an NVIDIA 3090 graphics card, and a 2TB solid-state drive. The program primarily obtains information from the temperature controllers, shutter controllers, and the camera (Fig. S11). We use a USB 3.0 interface to connect the camera to the computer. Temperature controllers are connected in series using the Modbus protocol to simplify the wiring. To reduce the delay in substrate temperature control, our program solely looks up information by address from the temperature controllers connected to the substrate.
Model construction environment
The model development occurred in the Jupyter Notebook environment on the Ubuntu system, utilizing Python version 3.9. We installed PyTorch based on CUDA 11.8 in this environment to leverage graphics card operations. The original video data was rapidly converted into the NumPy arrays format using code with random data augmentation within the Ubuntu system. Subsequently, the processed data was stored on our computer’s hard disk.
Program interface and deployment environment
The program was developed using LabVIEW, the built-in NI VISA, NI VISION, and Python libraries for data acquisition and processing (see Supplementary Information for program interface and deployment environment, S12). The program also employs ONNX for model deployment and TensorRT for inference acceleration and allows users to set targets for the desired QD density. Once the substrate and In cell temperatures stabilize, the program can be initiated. Firstly, the program checks the shutter status and controls the substrate and cell temperatures. Once the In shutter is open, the model starts to analyze RHEED data in real-time. The model outputs are displayed as numerical values at the top of the interface and are converted to corresponding label characters on the right side. The growth status can be displayed in the “Reminder Information”. Before and after the QDs formation, the “Reminder Information” will show up as “Stage 1” and “Stage 2”, respectively. It will display “Finished” when the model results meet the targets several times after the QD formation.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The datasets generated during and/or analyzed during the current study are available in the Figshare repository, https://doi.org/10.6084/m9.figshare.2434705361. Source data are provided with this paper.
Code availability
The codes supporting the findings of this study are available from the corresponding authors upon request.
References
Verma, A. K. et al. Low areal densities of InAs quantum dots on GaAs(1 0 0) prepared by molecular beam epitaxy. J. Cryst. Growth 592, 126715 (2022).
Sun, J., **, P. & Wang, Z.-G. Extremely low density InAs quantum dots realized in situ on (100) GaAs. Nanotechnology 15, 1763–1766 (2004).
Chu, L., Arzberger, M., Böhm, G. & Abstreiter, G. Influence of growth conditions on the photoluminescence of self-assembled InAs/GaAs quantum dots. J. Appl. Phys. 85, 2355–2362 (1999).
LeCun, Y., Bengio, Y. & Hinton, G. Deep learning. Nature 521, 436–444 (2015).
Jordan, M. I. & Mitchell, T. M. Machine learning: trends, perspectives, and prospects. Science 349, 255–260 (2015).
Wakabayashi, Y. K. et al. Machine-learning-assisted thin-film growth: Bayesian optimization in molecular beam epitaxy of SrRuO3 thin films. APL Mater. 7, 101114 (2019).
Wakabayashi, Y. K. et al. Bayesian optimization with experimental failure for high-throughput materials growth. npj Comput. Mater. 8, 180 (2022).
Peng, J., Muhammad, R., Wang, S. L. & Zhong, H. Z. How machine learning accelerates the development of quantum dots?†. Chin. J. Chem. 39, 181–188 (2020).
Uddin, G. M., Ziemer, K. S., Zeid, A., Lee, Y.-T. T. & Kamarthi, S. Process control model for growth rate of molecular beam epitaxy of MgO (111) nanoscale thin films on 6H-SiC (0001) substrates. Int. J. Adv. Manuf. Technol. 91, 907–916 (2016).
Kim, H. J. et al. Machine-learning-assisted analysis of transition metal dichalcogenide thin-film growth. Nano Converg. 10, 10 (2023).
Currie, K. R., LeClair, S. R. & Patterson, O. D. Self-directed, self-improving control of a molecular beam epitaxy process. IFAC Proc. Volumes 25, 83–87 (1992).
Currie, K. R. & LeClair, S. R. Self-improving process control for molecular beam epitaxy. Int. J. Adv. Manuf. Technol. 8, 244–251 (1993).
Meng, Z. et al. Combined use of computational intelligence and materials data for on-line monitoring and control of MBE experiments. Eng. Appl. Artif. Intell. 11, 587–595 (1998).
Liang, H. et al. Application of machine learning to reflection high-energy electron diffraction images for automated structural phase map**. Phys. Rev. Mater. 6, 063805 (2022).
Kwoen, J., Arakawa, Y. Classification of in situ reflection high energy electron diffraction images by principal component analysis. Jpn. J. Appl. Phys. 60, SBBK03 (2021).
Provence, S. R. et al. Machine learning analysis of perovskite oxides grown by molecular beam epitaxy. Phys. Rev. Mater. 4, 083807 (2020).
Gliebe, K. & Sehirlioglu, A. Distinct thin film growth characteristics determined through comparative dimension reduction techniques. J. Appl. Phys. 130, 125301 (2021).
Kwoen, J. & Arakawa, Y. Multiclass classification of reflection high-energy electron diffraction patterns using deep learning. J. Cryst. Growth 593, 126780 (2022).
Lee, K. K. et al. Using neural networks to construct models of the molecular beam epitaxy process. IEEE Trans. Semicond. Manuf. 13, 34–45 (2000).
Zhang, L. & Shao, S. Image-based machine learning for materials science. J. Appl. Phys. 132, 100701 (2022).
Vasudevan, R. K., Tselev, A., Baddorf, A. P. & Kalinin, S. V. Big-data reflection high energy electron diffraction analysis for understanding epitaxial film growth processes. ACS Nano 8, 10899–10908 (2014).
Xue, S. & Abhayaratne, C. Region-of-interest aware 3D ResNet for classification of COVID-19 chest computerised tomography scans. IEEE Access 11, 28856–28872 (2023).
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition. Proc. IEEE Conference on Computer Vision and Pattern Recognition (IEEE, 2016).
Watanabe, K., Koguchi, N. & Gotoh, Y. Fabrication of GaAs quantum dots by modified droplet epitaxy. Jpn. J. Appl. Phys. 39, L79 (2000).
Wang, Z. M., Liang, B. L., Sablon, K. A., Salamo, G. J. Nanoholes fabricated by self-assembled gallium nanodrill on GaAs(100). Appl. Phys. Lett. 90, 113120 (2007).
Heyn, C. et al. Highly uniform and strain-free GaAs quantum dots fabricated by filling of self-assembled nanoholes. Appl. Phys. Lett. 94, 183113 (2009).
Nemcsics, Á. et al. The RHEED tracking of the droplet epitaxial grown quantum dot and ring structures. Mater. Sci. Eng.: B 165, 118–121 (2009).
Ozaki, N. et al. Selective-area growth of self-assembled InAs-QDs by metal mask method for optical integrated circuit applications. MRS Proc. 959, 1703 (2011).
Oikawa, S., Makaino, A., Sogabe, T. & Yamaguchi, K. Growth Process and Photoluminescence Properties of In‐plane Ultrahigh‐density InAs Quantum Dots on InAsSb/GaAs(001). Phys. Status Solidi (b) 255, 1700307 (2017).
Yamaguchi, K. & Kanto, T. Self-assembled InAs quantum dots on GaSb/GaAs(0 0 1) layers by molecular beam epitaxy. J. Cryst. Growth 275, e2269–e2273 (2005).
Shimomura, K., Shirasaka, T., Tex, D. M., Yamada, F. & Kamiya, I. RHEED transients during InAs quantum dot growth by MBE. J. Vac. Sci. Technol. B 30, 02B128 (2012).
Gunasekera, M., Freundlich, A. Real time during growth metrology and assessment of kinetics of epitaxial quantum dots by RHEED. In Proc. 38th IEEE Photovoltaic Specialists Conference (IEEE, 2012).
Patella, F. et al. Reflection high energy electron diffraction observation of surface mass transport at the two- to three-dimensional growth transition of InAs on GaAs(001). Appl. Phys. Lett. 87, 252101 (2005).
Wang, Y. et al. End-to-end video instance segmentation with transformers. I: Proc. IEEE/CVF conference on computer vision and pattern recognition (IEEE, 2021).
Chen, Y.-H., Krishna, T., Emer, J. S. & Sze, V. Eyeriss: an energy-efficient reconfigurable accelerator for deep convolutional neural networks. IEEE J. Solid-State Circuits 52, 127–138 (2017).
Jiang, J. et al. Multi-spectral RGB-NIR image classification using double-channel CNN. IEEE Access 7, 20607–20613 (2019).
Xu, W., Gao, F., Zhang, J., Tao, X. & Alkhateeb, A. Deep learning based channel covariance matrix estimation with user location and scene images. IEEE Trans. Commun. 69, 8145–8158 (2021).
Codella, N. C. F. et al. Deep learning ensembles for melanoma recognition in dermoscopy images. IBM J. Res. Dev. 61, 5:1–5:15 (2017).
Komiske, P. T., Metodiev, E. M. & Schwartz, M. D. Deep learning in color: towards automated quark/gluon jet discrimination. J. High. Energy Phys. 2017, 110 (2017).
Reith, F., Koran, M. E., Davidzon, G. & Zaharchuk, G. Application of deep learning to predict standardized uptake value ratio and amyloid status on 18F-Florbetapir PET using ADNI data. Am. J. Neuroradiol. 41, 980–986 (2020).
Liao, Y., **ong, P., Min, W., Min, W. & Lu, J. Dynamic sign language recognition based on video sequence with BLSTM-3D residual networks. IEEE Access 7, 38044–38054 (2019).
Xue, F., Ji, H. & Zhang, W. Mutual information guided 3D ResNet for self‐supervised video representation learning. IET Image Process. 14, 3066–3075 (2020).
Tatebayashi, J., Nishioka, M., Someya, T. & Arakawa, Y. Area-controlled growth of InAs quantum dots and improvement of density and size distribution. Appl. Phys. Lett. 77, 3382–3384 (2000).
Frigeri, P. et al. Effects of the quantum dot ripening in high-coverage InAs∕GaAs nanostructures. J. Appl. Phys. 102, 083506 (2007).
Kuncheva, L.I. Combining Pattern Classifiers: Methods and Algorithms (John Wiley & Sons, Hoboken, 2014).
Wolpert, D. H. Stacked generalization. Neural Netw. 5, 241–259 (1992).
Dietterich, T. G. Ensemble methods in machine learning. in Multiple Classifier Systems (Springer Berlin Heidelberg, 2000).
Placidi, E. et al. InAs/GaAs(001) epitaxy: kinetic effects in the two-dimensional to three-dimensional transition. J. Phys.: Condens. Matter 19, 225006 (2007).
Kobayashi, N. P., Ramachandran, T. R., Chen, P. & Madhukar, A. In situ, atomic force microscope studies of the evolution of InAs three-dimensional islands on GaAs(001). Appl. Phys. Lett. 68, 3299–3301 (1996).
Cheng, Y., Wang, D., Zhou, P., Zhang, T. A survey of model compression and acceleration for deep neural networks. Preprint at https://arxiv.org/abs/1710.09282.
Sutton, R. S. Learning to predict by the methods of temporal differences. Mach. Learn. 3, 9–44 (1988).
Vamathevan, J. et al. Applications of machine learning in drug discovery and development. Nat. Rev. Drug Discov. 18, 463–477 (2019).
Joyce, T. B. & Bullough, T. J. Beam equivalent pressure measurements in chemical beam epitaxy. J. Cryst. Growth 127, 265–269 (1993).
Sfaxi, L., Bouzaiene, L., Sghaier, H. & Maaref, H. Effect of growth temperature on InAs wetting layer grown on (113)A GaAs by molecular beam epitaxy. J. Cryst. Growth 293, 330–334 (2006).
Song, H. Z. et al. Formation ofInAs∕GaAsquantum dots from a subcritical InAs wetting layer: a reflection high-energy electron diffraction and theoretical study. Phys. Rev. B 73, 115327 (2006).
Guo, S. P., Ohno, H., Shen, A., Matsukura, F. & Ohno, Y. InAs self-organized quantum dashes grown on GaAs (211)B. Appl. Phys. Lett. 70, 2738–2740 (1997).
Okumura, S. et al. Impact of low-temperature cover layer growth of InAs/GaAs quantum dots on their optical properties. Jpn. J. Appl. Phys. 61, 085503 (2022).
Lee, S., Lazarenkova, O. L., von Allmen, P., Oyafuso, F. & Klimeck, G. Effect of wetting layers on the strain and electronic structure of InAs self-assembled quantum dots. Phys. Rev. B 70, 125307 (2004).
Offermans, P., Koenraad, P. M., Nötzel, R., Wolter, J. H. & Pierz, K. Formation of InAs wetting layers studied by cross-sectional scanning tunneling microscopy. Appl. Phys. Lett. 87, 111903 (2005).
Ruiz-Marín, N. et al. Effect of the AlAs cap** layer thickness on the structure of InAs/GaAs QD. Appl. Surf. Sci. 573, 151572 (2022).
Shen, C. et al. Machine-Learning-Assisted and Real-Time-Feedback-Controlled Growth of InAs/GaAs Quantum Dots, Figshare, https://doi.org/10.6084/m9.figshare.24347053, 2024.
Acknowledgements
This work was supported by the National Key R&D Program of China (Grant No. 2021YFB2206503, C.Z.), National Natural Science Foundation of China (Grant No. 62274159, C.Z.), the “Strategic Priority Research Program” of the Chinese Academy of Sciences (Grant No. XDB43010102, C.Z.), and CAS Project for Young Scientists in Basic Research (Grant No. YSBR-056, C.Z.).
Author information
Authors and Affiliations
Contributions
C.S. and W.K.Z. contributed equally. C.Z. conceived of the idea, designed the investigations and the growth experiments. C.S., W.K.Z. and M.Y.L. performed the molecular beam epitaxial growth. C.S., K.Y.X., H.C. and C.X. did the sample characterization. C.S., C.Z., Z.Y.S., J.T., Z.F.W, Z.M.W. and C.L.X. wrote the manuscript. C.Z. led the molecular beam epitaxy program. B.X. and Z.G.W. supervised the team. All authors have read, contributed to, and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Shen, C., Zhan, W., **n, K. et al. Machine-learning-assisted and real-time-feedback-controlled growth of InAs/GaAs quantum dots. Nat Commun 15, 2724 (2024). https://doi.org/10.1038/s41467-024-47087-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-024-47087-w
- Springer Nature Limited