Abstract
Interactions of tumor cells with immune cells in the tumor microenvironment play an important role during malignancy progression. We previously identified that GAS5 inhibited tumor development by suppressing proliferation of tumor cells in non-small cell lung cancer (NSCLC). Herein, we discovered a tumor-suppressing role for tumor cell-derived GAS5 in regulating tumor microenvironment. GAS5 positively coordinated with the infiltration of macrophages and T cells in NSCLC clinically, and overexpression of GAS5 promoted macrophages and T cells recruitment both in vitro and in vivo. Mechanistically, GAS5 stabilized p53 by directly binding to MYBBP1A and facilitating MYBBP1A-p53 interaction, and enhanced p53-mediated transcription of IRF1, which activated type I interferon signaling and increased the production of downstream CXCL10 and CCL5. We also found that activation of type I interferon signaling was associated with better immunotherapy efficacy in NSCLC. Furthermore, the stability of GAS5 was regulated by NAT10, the key enzyme responsible for N4-acetylcytidine (ac4C) modification, which bound to GAS5 and mediated its ac4C modification. Collectively, tumor cell-derived GAS5 could activate type I interferon signaling via the MYBBP1A-p53/IRF1 axis, promoting immune cell infiltration and potentially correlating with immunotherapy efficacy, which suppressed NSCLC progression. Our results suggested GAS5 as a promising predictive marker and potential therapeutic target for combination therapy in NSCLC.

A schematic diagram demonstrating the regulatory effect of GAS5 on immune cell infiltration by activating type I interferon signaling via MYBBP1A-p53/IRF1 axis in non-small cell lung cancer. IFN, interferon.
Similar content being viewed by others
Introduction
Lung cancer is the most common cause of cancer-related death and non-small cell lung cancer (NSCLC) accounts for the majority of lung cancer [1]. The initiation, growth, and metastasis of lung cancer are greatly influenced by the tumor microenvironment (TME), which is composed of tumor cells, infiltrating immune cells, cancer-associated fibroblasts, extracellular matrix, and other components [2]. Treatment strategy of lung cancer has been changed by immunotherapy in recent years, which targets TME and has improved survival of NSCLC patients [2]. Tumors exhibiting ‘hot’ immune signature are infiltrated with more immune cells and patients with these tumors are more likely to benefit from immunotherapy [3]. The TME can be modulated by tumor cells via the secretion of cytokines, chemokines, and exosomes [4], and a better understanding of this interaction is crucial for the development of novel diagnosis, prognosis, and treatment strategy of NSCLC.
Type I interferon signaling was initially discovered for its antiviral effects [5], and its role in tumor biology has been explored. Generally, activation of the type I interferon signaling exhibits anti-tumor effects, such as inhibiting the growth and metastasis of tumor cells and regulating the infiltration of immune cells [6]. These effects are achieved by inducing the expression of interferon-stimulated genes (ISGs), including genes encoding chemokines and major histocompatibility complex (MHC) molecules [7, 8]. Type I interferon signaling also mediates tumor response to immunotherapy [9, 10]. Novel anticancer therapeutic approaches involving type I interferon signaling are being explored, some of which are already in phases of clinical trials [6].
Long non-coding RNAs (lncRNAs) are RNAs that contain more than 200 nucleotides in length and cannot be translated into functional proteins [11]. LncRNA growth arrest-specific 5 (GAS5) was originally discovered in growth-arrested cells [12] and has been reported to be involved in multiple diseases by regulating NF-κB pathway [13], cAMP/CREB pathway [14], Wnt/β-catenin pathway [15] and etc. We previously found that GAS5 was downregulated and associated with worse clinicopathological characteristics in NSCLC, and suppressed growth and migration of tumor cells by increasing p53 expression at post-transcription level [16]. In the current study, we further explored the influence of tumor cell-derived GAS5 on immune cells in the TME and the underlying mechanism. We found that GAS5 modulated the infiltration of macrophages and T cells by activating the type I interferon signaling via MYBBP1A-p53/IRF1 axis in NSCLC, which might enhance immunotherapy efficacy, indicating a promising predictive and therapeutic target for NSCLC.
Results
GAS5 inhibits NSCLC development and promotes immune cell infiltration
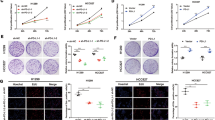
We had reported that GAS5 expression was negatively correlated with worse clinical characteristics in NSCLC specimens previously, including tumor size and TNM stage [16], and we analyzed its association with immune cell infiltration in the current study. Firstly, we explored this association using clinical specimens of NSCLC. The expression of GAS5 was analyzed by qPCR and patients were categorized into GAS5-high and GAS5-low groups according to the median value. Results of immunohistochemical (IHC) staining showed that higher staining signals of CD68, CD3, CD4, and CD8 were detected in the GAS5-high group than in the GAS5-low group (Fig. 1A, B), suggesting NSCLC tissues with higher GAS5 expression levels were infiltrated with more immune cells. Then we adopted transwell assay to evaluate the influence of tumor cell-derived GAS5 on immune cell recruitment (Fig. 1C). Macrophages and T cells are important parts of TME and can influence immunotherapeutic responsiveness [4, 17, 18]; therefore, THP-1-derived macrophages and PBMCs were chosen. A significant increase in the number of migrated macrophages and PBMCs attracted by conditioned medium from GAS5-overexpressing cells was observed (Fig. 1D; Supplementary Fig. 1A). Consistent with this result, fewer macrophages and PBMCs migrated when the lower chamber was filled with conditioned medium from GAS5-knockdown cells (Fig. 1E; Supplementary Fig. 1B). To validate the above results in vivo, a xenograft mouse model was established with A549 cells transfected with lentivirus encoding GAS5 or control lentivirus. Tumor growth was significantly inhibited in mice implanted with GAS5-overexpressing cells compared to that in mice implanted with control cells (Fig. 1F–H). The IHC analyses indicated that overexpression of GAS5 inhibited tumor cell proliferation (Ki-67 staining) and facilitated macrophage infiltration (F4/80 staining) (Fig. 1I; Supplementary Fig. 1C). In addition, we found that the expression of Il12a, Il23a and Tnf was upregulated, while Il10 was downregulated in GAS5-overexpressing subcutaneous tumors (Fig. 1J), indicating a proinflammatory TME. Taken together, our results indicated that tumor cell-derived GAS5 could enhance immune cell recruitment and suppress tumor development in NSCLC.
Representative images (A) and statistical results (B) of CD68, CD3, CD4, and CD8 staining in GAS5-high and GAS5-low NSCLC tissue samples. Scale bar = 50 μm. GAS5-low n = 23, GAS5-high n = 22. C Graphic illustration of a transwell assay. Immune cells were placed into the upper chamber initially, and the conditioned medium from tumor cells was placed in the lower chamber. D More macrophages and PBMCs migrated when the lower chamber was filled with conditioned medium collected from GAS5-overexpressing cells. E Less macrophages and PBMCs migrated when the lower chamber was filled with conditioned medium collected from GAS5-knockdown cells. F–H GAS5-overexpressing cells and control cells were injected subcutaneously into the right flank of each BALB/c nude mouse. Tumor volume and weight were significantly suppressed in the GAS5-overexpressing group. n = 5 in each group. I Representative images of Ki-67 and F4/80 staining in subcutaneous tumors. Scale bar = 50 μm. J The expression of Il12a, Il23a, Tnf, and Il10 analyzed by qPCR in subcutaneous tumors. n = 5 in each group. Data are represented as mean ± SD. *P < 0.05.
GAS5 facilitates immune cell infiltration potentially by activating type I interferon signaling pathway
To explore the underlying mechanism, RNA sequencing was performed in GAS5-overexpressing A549 cells and control cells, with 659 genes upregulated and 541 genes downregulated (Fig. 2A). Gene Ontology (GO) enrichment analysis showed that the most upregulated pathways are type I interferon signaling-related (Fig. 2B). The most differentially expressed genes in type I interferon signaling included IRF1, HLA-B, CXCL10, and CCL5 (Fig. 2C). To explore the association between GAS5 and the type I interferon signaling pathway, IHC staining was conducted on formalin-fixed, paraffin-embedded (FFPE) specimens of NSCLC. Since the upregulation of signal transducer and activator of transcription 1 (STAT1) phosphorylation is one of the hallmark events during activation of the canonical type I interferon signaling pathway [7], we also examined the level of phosphorylated STAT1 (p-STAT1). The results suggested a higher activation level of the type I interferon signaling in patients in the GAS5-high group than in patients in the GAS5-low group (Fig. 2D, E). The analyzation of specimens from the xenograft mouse model also supported that GAS5 could activate type I interferon signaling pathway at both mRNA level (Fig. 2F) and protein level (Fig. 2G, H; Supplementary Fig. 1D). In addition to IRF1, HLA-B, CXCL10, and CCL5, several other molecules in type I interferon signaling, including IFIT2, RSAD2 and IFITM3, were also detected upregulated in GAS5-overexpressing subcutaneous tumors (Supplementary Fig. 1E–G).
The volcano plot (A) and the GO analysis (B) of the RNA sequencing of GAS5-overexpressing cells and control cells (n = 3). C The expression profile of IRF1, HLA-B, CXCL10, and CCL5 in RNA sequencing of GAS5-overexpressing cells and control cells. Representative images (D) and statistical analyses (E) of p-STAT1, IRF1, HLA-B, CXCL10, and CCL5 staining in human NSCLC tissue samples. Scale bar = 50 μm. GAS5-low n = 23, GAS5-high n = 22. F The mRNA expression of IRF1, HLA-B, CXCL10, and CCL5 was analyzed by qPCR in subcutaneous tumors. n = 5 in each group. G The protein level of IRF1, HLA-B, CXCL10, and CCL5 was analyzed by western blot in subcutaneous tumors. n = 5 in each group. H Representative images of p-STAT1, IRF1, HLA-B, CXCL10, and CCL5 staining in subcutaneous tumors. Scale bar = 50 μm. Representative images (I) and statistical results (J) of GAS5-downstream molecules staining in FFPE specimens of NSCLC patients receiving anti-PD-1/anti-PD-L1 treatment. Scale bar = 50 μm. NDB n = 11, DCB n = 11. DCB: Durable clinical benefit; NDB no durable benefit. Data are represented as mean ± SD. *P < 0.05.
Immunotherapy has achieved great clinical success and studies have reported tumor immune infiltrates are associated with patients’ response to immunotherapy [18]. We also found that in lung cancer patients receiving neoadjuvant immunotherapy, more infiltrated CD3+, CD4+, CD8+, or CD68+ cells were detected in pre-treatment tumor tissues from patients achieving pathologic complete response (pCR) (Supplementary Fig. 1H). Since GAS5 was identified to influence the infiltration of immune cells and activate type I interferon signaling pathway, we tried to demonstrate the association of the type I interferon signaling and the response of NSCLC patients to immunotherapy. We investigated the protein levels of p-STAT1, IRF1, and HLA-B in these patients by performing IHC staining on the most recent FFPE samples collected before they received anti-PD-1/anti-PD-L1 treatment. Patients who achieved durable clinical benefit (DCB) tended to express higher levels of p-STAT1, IRF1, and HLA-B than patients who received no durable benefit (NDB) (Fig. 2I, J). Taken together, these data suggested that the type I interferon signaling pathway could be activated by GAS5 in vivo and was correlated with immunotherapy efficacy in NSCLC.
GAS5 facilitates immune cell infiltration by activating type I interferon signaling pathway
The regulation of type I interferon signaling pathway by GAS5 was validated in vitro using A549 and HCC827 cells. Results of qPCR showed increased expression of IRF1, HLA-B, CXCL10, and CCL5 in GAS5-overexpressing cells (Fig. 3A, B) and decreased expression of these genes in GAS5-knockdown cells (Fig. 3C, D) compared to control cells. The western blot analysis results revealed that the protein levels of p-STAT1, IRF1, HLA-B, CXCL10, and CCL5 were significantly increased by overexpression of GAS5 (Fig. 3E) and decreased by knockdown of GAS5 (Fig. 3F). To investigate the role of type I interferon signaling in the recruitment of immune cells by GAS5, we added neutralizing antibodies targeting CXCL10 or CCL5, or isotype control into the medium of lower chamber in the transwell assay. Results suggested that the influence of GAS5 on the migration of macrophages and PBMCs (Fig. 3G, Supplementary Fig. 1I) was partly reversed with the addition of these neutralizing antibodies, indicating that type I interferon signaling mediated tumor cell-derived GAS5’s regulatory effect on immune cell infiltration.
A, B The mRNA expression of IRF1, HLA-B, CXCL10, and CCL5 in GAS5-overexpressing cells and control cells. C, D The mRNA expression of IRF1, HLA-B, CXCL10, and CCL5 in GAS5-knockdown and control cells. E The protein level of molecules in type I signaling in GAS5-overexpressing cells and control cells. F The protein level of molecules in type I signaling in GAS5-knockdown and control cells. G Decreased number of migrated macrophages and PBMCs with the presence of anti-CXCL10 antibody (2.0 μg/ml) or anti-CCL5 antibody (2.0 μg/ml) was observed. Data are represented as mean ± SD. *P < 0.05.
GAS5 regulates type I interferon signaling by modulating IRF1
Since the type I interferon signaling was regulated at the mRNA level and the gene expression can be modulated by transcription factors [19], we tried to identify the transcription factor responsible for the activation of this pathway. The results of transcription factor enrichment analysis performed on ChEA3 [20] showed that IRF1, which is one of the ISGs [21] and can mediate the activation of type I interferon signaling [22, 23] was among the top-ranked potential transcription factors (Fig. 4A). In addition, the level of IRF1 was proven to be modulated by GAS5 (Figs. 2, 3). Thus, we used a siRNA to reduce the level of IRF1 while overexpressing GAS5, and it turned out that the type I interferon signaling activation triggered by overexpression of GAS5 was attenuated at both the mRNA (Fig. 4B) and protein levels (Fig. 4C). We also increased the level of IRF1 while reducing GAS5 expression with a siRNA, and the results indicated that the elevation of IRF1 level restored the inhibition of the type I interferon signaling caused by downregulation of GAS5 (Fig. 4D, E). Moreover, we found that the knockdown of IRF1 counterbalanced the enhancement of macrophage migration by GAS5-overexpression, and consistently, overexpression of IRF1 restored the GAS5-knockdown mediated decreased migration of macrophages (Fig. 4F). These data suggested that GAS5 regulates the type I interferon signaling pathway possibly by modulating the level of IRF1.
A The result of the transcription factor enrichment analysis of changed genes in type I interferon signaling using ChEA3. Knockdown of IRF1 downregulated the GAS5-overexpression-induced increased mRNA (B) and protein (C) level of molecules in type I interferon signaling. Overexpression of IRF1 upregulated the GAS5-knockdown-induced decreased mRNA (D) and protein (E) level of molecules in type I interferon signaling. F Knockdown or overexpression of IRF1 in tumor cells rescued the GAS5-induced changed migration of macrophages. Data are represented as mean ± SD. *P < 0.05.
MYBBP1A/p53 mediates the regulatory effect of GAS5 on IRF1
To further demonstrate the mechanism by which GAS5 regulates the expression of IRF1, we predicted potential transcription factors of IRF1 with PROMO [24] after obtaining the promoter sequence of IRF1 from the UCSC Genome Browser. The results showed that p53 was one of the possible transcription factors of IRF1 (Fig. 5A), and our previous study revealed that p53-mediated GAS5’s function [16]. Therefore, we chose p53 as the candidate transcription factor. Our previous results had shown GAS5 regulated p53 at post-transcription level [16], and we confirmed the positive regulatory effect of GAS5 on p53 protein in both GAS5-overexpressing cells and GAS5-knockdown cells (Supplementary Fig. 2A). Then, we inhibited protein synthesis using cycloheximide (CHX) and found that degradation of the p53 protein was faster in GAS5-knockdown cells (Fig. 5B), indicating that the positive regulatory effect of GAS5 on p53 was at least partly achieved by an increase in the stability of the p53 protein. After validating the relationship between GAS5 and p53, we explored whether the effect of GAS5 on IRF1 is p53-dependent. Firstly, we found p53 could positively regulate the expression of IRF1 at both mRNA level (Fig. 5C) and protein level (Fig. 5D). Second, we confirmed that the decrease of p53 could alleviate GAS5’s upregulation effect on IRF1 (Fig. 5E, F), and consistently, the overexpression of p53 could rescue the decreased level of IRF1 by the GAS5-specific siRNA (Fig. 5G, H). Next, the interaction of p53 and IRF1 promoter region was validated by the analysis of chromatin immunoprecipitation (ChIP) products, and the CDKN1A promoter fragment was used as a positive reference (Fig. 5I, Supplementary Fig. 2B). Moreover, the dual-luciferase reporter assays demonstrated that the mutation of binding sites (Fig. 5J) decreased the transcriptional activity of IRF1 promoter (Fig. 5K). These results indicated that GAS5 depended on p53 to influence IRF1.
A p53 was predicted as a potential transcription factor of IRF1 by PROMO. B GAS5-knockdown cells or control cells were treated with cycloheximide (CHX) for the indicated times and western blot analyses of p53 and β-actin were performed. The expression of IRF1 was regulated by p53 at the mRNA level (C) and protein level (D). Knockdown of p53 rescued the GAS5-overexpression-induced increased mRNA (E) and protein (F) level of IRF1. Overexpression of p53 rescued the GAS5-knockdown-induced decreased mRNA (G) and protein (H) level of IRF1. I ChIP-qPCR was performed to determine the binding of p53 to the IRF1 promoter region and CDKN1A promoter region. J Graphic illustration of the potential p53-binding sequence in the IRF1 promoter region. K The wild-type and mutated luciferase reporter plasmids, p53 plasmid, and Renilla luciferase control plasmid were transfected into HEK293 cells, followed by dual-luciferase reporter assays. Data are represented as mean ± SD. *P < 0.05.
Since lncRNAs can mediate signal transduction by interacting with proteins [25], we performed RNA pull-down followed by mass spectrometry to identify GAS5-binding proteins that might be involved in the effect of GAS5 on p53 level, and MYBBP1A was one of the proteins specifically bound to sense GAS5 but not antisense GAS5 (Fig. 6A). MYBBP1A is a nucleolar protein that has been reported to bind to transcription factors and regulate their activity [26, 27]. It has been reported that MYBBP1A is able to stabilize the p53 protein and promote p53 activity by directly binding to p53, facilitating the p53-p300 interaction and thus enhancing p53 acetylation [28, 29]. Consistent with previous studies, our results supported that MYBBP1A could stabilize p53 protein by using CHX (Fig. 6B) and reduced level of p53 with knockdown of MYBPP1A was observed (Fig. 6C). To confirm the interaction between GAS5 and MYBBP1A, an RNA immunoprecipitation (RIP) assay was performed using anti-MYBBP1A antibodies to immunoprecipitate endogenous MYBBP1A together with its bound RNAs from A549 cell lysates. The qPCR results revealed that GAS5 directly bound to MYBBP1A in A549 cells (Fig. 6D). To evaluate the influence of GAS5 on the interaction between MYBBP1A and p53, co-immunoprecipitation (co-IP) was conducted using A549 and HCC827 cell lysates with anti-MYBBP1A antibodies or anti-p53 antibodies. Less p53 was detected in GAS5-knockown cells than in control cells in the MYBBP1A IP experiment (Fig. 6E, F; Supplementary Fig. 2C, D) and consistently, less MYBBP1A was detected in GAS5-knockdown cells than in control cells in the p53 IP experiment (Fig. 6G, H; Supplementary Fig. 2E, F). Taken together, our results suggested that GAS5 affects the stability of the IRF1 transcription factor p53 by binding to MYBBP1A and therefore regulates the level of IRF1.
A An RNA pull-down was performed to identify GAS5-binding proteins. B MYBBBP1A-knockdown cells or control cells were treated with cycloheximide (CHX) for the indicated times and western blot analyses of p53 and β-actin were performed. C Level of p53 protein with the knockdown of MYBBP1A. D RIP analyses were performed to determine the binding of MYBBP1A protein to GAS5. E GAS5-knockdown or control A549 cells were subjected to co-IP with anti-MYBBP1A antibody and immunoprecipitated proteins were analyzed by western blotting. F After quantification of western blotting bands by ImageJ, relative integrated density was calculated using the following formula: relative integrated density = integrated density of p53 in co-IP sample/integrated density of p53 in corresponding group of WCL. G GAS5-knockdown or control A549 cells were subjected to co-IP with anti-p53 antibody and immunoprecipitated proteins were analyzed by western blotting. H After quantification of western blotting bands by ImageJ, relative integrated density was calculated using the following formula: relative integrated density = integrated density of MYBBP1A in co-IP sample/integrated density of MYBBP1A in corresponding group of WCL. WCL whole cell lysate. Data are represented as mean ± SD. *P < 0.05.
NAT10 regulates GAS5 expression and mediates its ac4C modification
The expression of lncRNAs can be regulated by their binding proteins [30]. The results of mass spectrometry performed after RNA pull-down showed that NAT10 bound to GAS5 (Fig. 6A). NAT10 had been reported to influence the RNA expression by mediating ac4C modification [31, 32]; thus, we tried to determine the association of NAT10 and GAS5. Clinically, the analysis of CCLE dataset showed that the RNA expression of NAT10 and GAS5 was significantly positively correlated in tumor cells (Fig. 7A). A positive association of NAT10 and GAS5 was also found in clinical specimens of NSCLC (Fig. 7B). Then, we explored whether NAT10 could regulate the expression of GAS5, and the results showed that the expression of GAS5 was decreased with the knockdown of NAT10 in NSCLC cells (Fig. 7C). After confirming the correlation of GAS5 and NAT10, we validated the interaction of NAT10 and GAS5 by western blot analysis (Fig. 7D). Consistently, the qPCR results of the RIP assay products precipitated with the anti-NAT10 antibody showed significant enrichment of GAS5 in the anti-NAT10 antibody group compared to the IgG group (Fig. 7E). Previous studies suggested that ac4C modification plays an important role in NAT10’s function [ The human non-small cell lung cancer cell lines A549 and HCC827, human embryonic kidney 293 (HEK293) cells, and the human monocyte cell line THP-1 were purchased from the Institute of Biochemistry and Cell Biology, Chinese Academy of Sciences (Shanghai, China). The purchased cell lines were validated by short tandem repeat analysis and tested for mycoplasma contamination within the past 6 months. Peripheral blood mononuclear cells (PBMCs) were isolated from the volunteer’s blood using a lymphocyte separation solution (FCMACS, Nan**g, China). The A549 cell line, HCC827 cell line, and HEK293 cell line were maintained in F-12K (Meilun Bio, Dalian, China), RPMI-1640 (Gibco, Carlsbad, California), and DMEM (Gibco, Carlsbad, California) medium, respectively, supplemented with 10% FBS (HyClone, Carlsbad, California). The THP-1 cell line was cultured in RPMI-1640 with 10% FBS and 0.05 mM β-mercaptoethanol (Procell, Wuhan, China). All the cell lines were maintained in a humidified environment at 37 °C in 5% CO2. For differentiation into macrophages, THP-1 cells were maintained in RPMI-1640 medium supplemented with 1% FBS and 100 ng/ml phorbol-12-myristate acetate (FCMACS, Nan**g, China) for 24 h, and the culture medium was then replaced with RPMI-1640 medium supplemented with 10% FBS for another 24 h. Plasmid and siRNA transfection was performed using X-tremeGENE HP DNA Transfection Reagent (06366546001, Roche) and jetPRIME (114-15, Polyplus-transfection, Illkirch, France), respectively, following the manufacturers’ protocols. The detailed sequences of the siRNAs are listed in Supplementary Table 1. Since si-NC was purchased from RiboBio (Guangzhou, China) directly, the sequence was not provided. To generate cells with stable GAS5-overexpression, the GAS5-overexpression and empty vector lentiviruses were purchased from HanBio (Shanghai, China). For lentiviral transduction, A549 cells were cultured in a complete medium containing lentivirus in the presence of 6 μg/mL polybrene, and after 24 h, the medium was replaced with a normal complete culture medium. After selection with 4 μg/mL puromycin (Solarbio, Bei**g, China), the stable GAS5-overexpressing A549 cells and control cells were used for the following studies. A total of 47 lung cancer tumor tissues were obtained from **ling Hospital. Some of these tissues were fixed with formalin and then embedded in paraffin, and the others were rapidly frozen in liquid nitrogen for RNA extraction and other experiments. FFPE lung cancer tissues from 24 patients who had received anti-PD-1/anti-PD-L1 therapy were also obtained from **ling Hospital. The efficacy of anti-PD-1/anti-PD-L1 therapy was defined as DCB (complete response or partial response or stable disease lasting over 6 months) or NDB (progressive disease or stable disease lasting no longer than 6 months) [63]. In addition, FFPE lung cancer tissues from 17 patients who received neoadjuvant immunotherapy were also retrospectively collected from **ling Hospital. The achievement of a pathologic complete response was evaluated by pathologists. This study was approved by the Ethics Committee of **ling Hospital, and tissue samples were collected in compliance with the informed consent policy. To extract RNA from tissues, tissues were homogenized in TRIzol reagent (Vazyme, Nan**g, China) with beads and were then placed on ice for 30 min. After that, ice-cold chloroform, isopropanol, and 75% ethanol were used sequentially for RNA extraction. To extract RNA from cultured cells, the same protocol was followed, except that the step of homogenization with beads was omitted. Reverse transcription was performed with HiScript III RT SuperMix for qPCR (Vazyme, Nan**g, China) and the expression levels of targeted genes were evaluated via qPCR with ChamQ SYBR Color qPCR Master Mix (Vazyme, Nan**g, China). β-actin was used as the internal reference gene. The primer sequences are listed in Supplementary Table 1. Cells were homogenized using RIPA buffer with protease inhibitor cocktail (Bimake, Houston, Texas) on ice for 20 min. Then, cell lysates were centrifuged at 12,000 rpm for 15 min at 4 °C, and supernatants were collected for western blot analysis or co-IP with the indicated antibodies (Supplementary Table 2). Western blot signals were detected using ECL Ultra Western HRP Substrate (Millipore, Darmstadt, Germany). For co-IP, supernatants of cell lysates were incubated with the indicated antibodies or control IgG in RIPA buffer supplemented with protease inhibitor cocktail with rotation at 4 °C overnight, and Protein A/G PLUS-Agarose beads (Santa Cruz, Dallas, Texas) were then added and incubated with rotation for 4 more hours. Then, the beads with bound protein complexes were washed with RIPA buffer 3 times and eluted with 2× SDS loading buffer at 100 °C for 10 min. The eluted protein was analyzed by western blotting. Quantifications of western blotting bands were achieved using ImageJ software (for Windows, USA). ChIP was performed using a Magna ChIP Kit (17-10086, Millipore, Darmstadt, Germany) according to the manufacturer’s protocol. Briefly, 1 × 107 cells were fixed with 1% formaldehyde, lysed, and sonicated. Then, an anti-p53 antibody or control rabbit IgG was added to the sheared cross-linked chromatin with protein A/G magnetic beads and rotated overnight at 4 °C. After elution and reverse cross-linking and purification, the immunoprecipitated chromatin DNA was quantified by qPCR or DNA agarose electrophoresis. CDKN1A was used as a positive control. The sequences of primers targeting p53-binding sites for IRF1 promoter and CDKN1A promoter are listed in Supplementary Table 1. RIP was performed following the instructions of the EZ-Magna RIP Kit (17-701, Millipore, Darmstadt, Germany). Briefly, cells were harvested and were then lysed in a complete RIP Lysis Buffer, and magnetic beads were prepared. Then, the antibody of interest or negative control IgG was incubated with the magnetic beads for 30 min with rotation at room temperature, and cell lysate supernatants were then added and incubated with rotation overnight at 4 °C. The magnetic beads with bound protein-RNA complexes were washed 6 times, and the captured RNA was then freed from beads by proteinase K digestion and purified by phenol: chloroform: isoamyl alcohol. After being precipitated overnight at −80 °C, RNA was extracted and analyzed via qPCR. The pcDNA3.1-GAS5-sense and pcDNA3.1-GAS5-antisense constructs were linearized by digestion with the BamHI restriction enzyme (FD0054, Thermo Scientific, Waltham, Massachusetts), and in vitro transcription was then performed using a mMESSAGE mMACHINE™ T7 Transcription Kit (AM1344, Invitrogen, Waltham, Massachusetts). Transcribed RNAs were purified by a MEGAclear™ Transcription Clean-Up Kit (AM1908, Invitrogen, Waltham, Massachusetts) and were then biotin-labeled with a Pierce™ RNA 3’ End Desthiobiotinylation Kit (20163, Thermo Scientific, Waltham, Massachusetts). RNA pull-down was performed using a Pierce™ Magnetic RNA-Protein Pull-Down Kit (20164, Thermo Scientific, Waltham, Massachusetts). Briefly, labeled RNAs were captured by magnetic beads in a capture buffer with rotation at room temperature for 30 min. Then, prepared cell lysates were rotated with RNA-magnetic bead complexes in RNA-protein binding buffer at 4 °C for 60 min. After washing 3 times, the precipitated products were eluted in an elution buffer and subjected to mass spectrometry analysis or western blot analysis. For RNA sequencing, stable GAS5-overexpressing A549 cells and control cells were collected for RNA extraction. After checking the purity and integrity of RNA, a total amount of 1 µg RNA per sample was used as input material for the generation of sequencing libraries using NEBNext UltraTM RNA Library Prep Kit for Illumina (NEB, USA) following the manufacturer’s recommendations. The libraries were sequenced on an Illumina Novaseq platform and 150 bp paired-end reads were generated at Novogene Co., Ltd. Paired-end clean reads were aligned to the reference genome using Hisat2 v2.0.5. FPKM, the expected number of Fragments Per Kilobase of transcript sequence per Million base pairs sequenced, was used for estimating gene expression. Differential expression analysis was performed using the DESeq2 R package (1.16.1). The P-values were adjusted using Benjamini and Hochberg’s approach and genes with an adjusted P-value < 0.05 were considered as differentially expressed. GO enrichment analysis of differentially expressed genes was conducted via the clusterProfiler R package. Briefly, 1 × 105 THP-1 cells were seeded into the upper chamber (3422, Corning, New York, New York) and were then induced to differentiate into macrophages by PMA as described in the ‘Cell lines and transfection’ section. Then, the medium in the lower chamber was replaced with conditioned medium collected from tumor cells, and the medium in the upper chamber was replaced with RPMI-1640. After 24 h, the Transwell membrane was fixed with methanol and stained with crystal violet (C0121, Beyotime, Suzhou, China). To count the cells that had migrated, a microscope (Zeiss, Oberkochen, Germany) was used to image the fields of view. For PBMCs, 1 × 105 PBMCs in 1640 containing 1% FBS were placed in the upper chamber (MCMP24H48, Millipore, Darmstadt, Germany), and the lower chamber was filled with conditioned media from tumor cells. After 3-4 h, cells that migrated into the lower chamber were counted using a cell counting chamber. For CXCL10 and CCL5 neutralizing experiments, anti-CXCL10 antibody (2.0 μg/ml) and anti-CCL5 antibody (2.0 μg/ml) were added to the collected conditioned medium 30 min before the migration assay started. FFPE tissue samples were deparaffinized in xylene and rehydrated in ethanol. After heat-induced epitope retrieval, IHC staining was conducted using anti-CD68, anti-F4/80, anti-CD3, anti-CD4, anti-CD8, anti-IRF1, anti-p-STAT1, anti-HLA-B, anti-CXCL10, anti-CCL5, anti-IFIT2, anti-RSAD2, and anti-IFITM3 antibodies. Immunofluorescence staining was performed using anti-CD68, anti-CD163, anti-CD3, anti-CD4 and anti-CD8 antibodies. For IRF1, p-STAT1, and HLA-B, CXCL10, and CCL5 staining, the score of the percentage of positive cells (0, 0–5%; 1, 6–25%; 2, 26–50%; 3, 51–75%; 4, >75%) and the score of the intensity of positive staining (0, negative; 1, week; 2, moderate; 3, strong) were multiplied to evaluate the expression of the molecules of interest. For CD68, CD3, CD4, CD8, and F4/80 staining, the number of positive cells in the fields of view was counted to assess the infiltration level of corresponding immune cells. The wild-type and mutated promoter sequences of IRF1 were synthesized and then cloned into Pgl4.10 to construct the luciferase reporter plasmids, respectively. HEK293 cells were seeded into 24-well plates and were transiently transfected with the luciferase reporter plasmids, p53 plasmid, and Renilla luciferase control plasmids. The luciferase activities were detected using the Dual-Luciferase Assay kit (E1910, Promega) following the manufacturer’s protocol. The relative luciferase activity was obtained by normalizing the firefly luciferase activity to the corresponding Renilla luciferase activity. Four- to six-week-old female BALB/c nude mice were purchased from GemPharmatech (Nan**g, China). A total of 3 × 106 stable GAS5-overexpressing A549 cells or control A549 cells in 60 μL PBS (mixed with 40 μL Matrigel) were injected subcutaneously into the right flank of each mouse using an insulin syringe (KDL, Wenzhou, China). The tumor size was measured by a caliper every 3 days after 6 days of injection, and the tumor volume was calculated using the following formula: 1/2 (length × width2). Mice were humanely sacrificed on Day 21 and tumors were carefully isolated. The harvested tumors were weighed, photographed, and then fixed with formalin for IHC staining or rapidly frozen in liquid nitrogen for RNA and protein assays. To reduce the influence of individual differences on the experimental results, the same species, strain, sex, and age of animals were selected in this study and their weights were similar when measured before the injection of tumor cells. The operators who injected tumor cells into mice were blinded to the group allocation when they operated. All animal experiments were performed in accordance with a protocol approved by the Animal Care and Use Committee of **ling Hospital. To identify transcription factors responsible for the activation of type I interferon signaling, ChEA3 (https://maayanlab.cloud/chea3/) was used. Potential transcription factors of IRF1 were obtained via PROMO (https://alggen.lsi.upc.es/cgi-bin/promo_v3/promo/promoinit.cgi?dirDB=TF_8.3). The association of NAT10 and GAS5 expression was assessed in CCLE datasets via cbioportal (http://www.cbioportal.org/). All in vitro experiments were repeated in triplicate. Statistical analyses were performed using GraphPad Prism 8.0 (GraphPad Software, San Diego, California). Data were plotted using GraphPad Prism 8.0. Two-tailed Student’s t-test was used to compare the differences between the two groups. Pearson correlation analysis and Spearman correlation analysis were both conducted to analyze correlations between two groups. A P-value < 0.05 was considered statistically significant.Materials and methods
Cell lines and transfection
Tissue samples
RNA extraction and qPCR
Western blot analysis and co-IP
ChIP
RIP
RNA pull-down assay
RNA sequencing
Transwell assay
IHC staining and immunofluorescence staining
Dual-luciferase reporter assay
In vivo study
Bioinformatics analysis
Statistical analysis
Data availability
The data generated or analyzed during the current study are available from the corresponding authors upon reasonable request.
References
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, et al. Global Cancer Statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2021;71:209–49.
Altorki NK, Markowitz GJ, Gao D, Port JL, Saxena A, Stiles B, et al. The lung microenvironment: an important regulator of tumour growth and metastasis. Nat Rev Cancer. 2019;19:9–31.
Galon J, Bruni D. Approaches to treat immune hot, altered and cold tumours with combination immunotherapies. Nat Rev Drug Discov. 2019;18:197–218.
Quail DF, Joyce JA. Microenvironmental regulation of tumor progression and metastasis. Nat Med. 2013;19:1423–37.
Lindenmann J, Burke DC, Isaacs A. Studies on the production, mode of action and properties of interferon. Br J Exp Pathol. 1957;38:551–62.
Yu R, Zhu B, Chen D. Type I interferon-mediated tumor immunity and its role in immunotherapy. Cell Mol Life Sci. 2022;79:191.
Lazear HM, Schoggins JW, Diamond MS. Shared and distinct functions of type I and type III interferons. Immunity. 2019;50:907–23.
Fuertes MB, Woo SR, Burnett B, Fu YX, Gajewski TF. Type I interferon response and innate immune sensing of cancer. Trends Immunol. 2013;34:67–73.
Mondal I, Das O, Sun R, Gao J, Yu B, Diaz A, et al. PP2Ac deficiency enhances tumor immunogenicity by activating STING-type I interferon signaling in glioblastoma. Cancer Res. 2023;83:2527–42.
Trono P, Tocci A, Palermo B, Di Carlo A, D’Ambrosio L, D’Andrea D, et al. hMENA isoforms regulate cancer intrinsic type I IFN signaling and extrinsic mechanisms of resistance to immune checkpoint blockade in NSCLC. J Immunother Cancer. 2023. https://doi.org/10.1136/jitc-2023-006913.
Statello L, Guo CJ, Chen LL, Huarte M. Gene regulation by long non-coding RNAs and its biological functions. Nat Rev Mol Cell Biol. 2021;22:96–118.
Schneider C, King RM, Philipson L. Genes specifically expressed at growth arrest of mammalian cells. Cell. 1988;54:787–93.
Zhu X, Sun Y, Yu Q, Wang X, Wang Y, Zhao Y. Exosomal lncRNA GAS5 promotes M1 macrophage polarization in allergic rhinitis via restraining mTORC1/ULK1/ATG13-mediated autophagy and subsequently activating NF-кB signaling. Int Immunopharmacol. 2023;121:110450.
Xu S, Wang Y, Li Z, Hua Q, Jiang M, Fan X. LncRNA GAS5 knockdown mitigates hepatic lipid accumulation via regulating MiR-26a-5p/PDE4B to activate cAMP/CREB pathway. Front Endocrinol. 2022;13:889858.
Huang Y, Ma J, Fan Y, Yang L. Mechanisms of human umbilical cord mesenchymal stem cells-derived exosomal lncRNA GAS5 in alleviating EMT of HPMCs via Wnt/β-catenin signaling pathway. Aging. 2023;15:4144–58.
Shi X, Sun M, Liu H, Yao Y, Kong R, Chen F, et al. A critical role for the long non-coding RNA GAS5 in proliferation and apoptosis in non-small-cell lung cancer. Mol Carcinog. 2015;54 Suppl 1:E1–e12.
Noy R, Pollard JW. Tumor-associated macrophages: from mechanisms to therapy. Immunity. 2014;41:49–61.
Binnewies M, Roberts EW, Kersten K, Chan V, Fearon DF, Merad M, et al. Understanding the tumor immune microenvironment (TIME) for effective therapy. Nat Med. 2018;24:541–50.
Lee TI, Young RA. Transcriptional regulation and its misregulation in disease. Cell. 2013;152:1237–51.
Keenan AB, Torre D, Lachmann A, Leong AK, Wojciechowicz ML, Utti V, et al. ChEA3: transcription factor enrichment analysis by orthogonal omics integration. Nucleic Acids Res. 2019;47:W212–w24.
Forero A, Ozarkar S, Li H, Lee CH, Hemann EA, Nadjsombati MS, et al. Differential activation of the transcription factor IRF1 underlies the distinct immune responses elicited by type I and type III interferons. Immunity. 2019;51:451–64.e6.
Venkatesh D, Ernandez T, Rosetti F, Batal I, Cullere X, Luscinskas FW, et al. Endothelial TNF receptor 2 induces IRF1 transcription factor-dependent interferon-β autocrine signaling to promote monocyte recruitment. Immunity. 2013;38:1025–37.
Yarilina A, Park-Min KH, Antoniv T, Hu X, Ivashkiv LB. TNF activates an IRF1-dependent autocrine loop leading to sustained expression of chemokines and STAT1-dependent type I interferon-response genes. Nat Immunol. 2008;9:378–87.
Messeguer X, Escudero R, Farré D, Núñez O, Martínez J, Albà MM. PROMO: detection of known transcription regulatory elements using species-tailored searches. Bioinformatics. 2002;18:333–4.
Lin C, Yang L. Long noncoding RNA in cancer: wiring signaling circuitry. Trends Cell Biol. 2018;28:287–301.
Owen HR, Elser M, Cheung E, Gersbach M, Kraus WL, Hottiger MO. MYBBP1a is a novel repressor of NF-kappaB. J Mol Biol. 2007;366:725–36.
Tavner FJ, Simpson R, Tashiro S, Favier D, Jenkins NA, Gilbert DJ, et al. Molecular cloning reveals that the p160 Myb-binding protein is a novel, predominantly nucleolar protein which may play a role in transactivation by Myb. Mol Cell Biol. 1998;18:989–1002.
Kuroda T, Murayama A, Katagiri N, Ohta YM, Fujita E, Masumoto H, et al. RNA content in the nucleolus alters p53 acetylation via MYBBP1A. EMBO J. 2011;30:1054–66.
Ono W, Akaogi K, Waku T, Kuroda T, Yokoyama W, Hayashi Y, et al. Nucleolar protein, Myb-binding protein 1A, specifically binds to nonacetylated p53 and efficiently promotes transcriptional activation. Biochem Biophys Res Commun. 2013;434:659–63.
Ghisays F, Garzia A, Wang H, Canasto-Chibuque C, Hohl M, Savage SA, et al. RTEL1 influences the abundance and localization of TERRA RNA. Nat Commun. 2021;12:3016.
Feng Z, Li K, Qin K, Liang J, Shi M, Ma Y, et al. The LINC00623/NAT10 signaling axis promotes pancreatic cancer progression by remodeling ac4C modification of mRNA. J Hematol Oncol. 2022;15:112.
Zhang Y, **g Y, Wang Y, Tang J, Zhu X, ** WL, et al. NAT10 promotes gastric cancer metastasis via N4-acetylated COL5A1. Signal Transduct Target Ther. 2021;6:173.
**e L, Zhong X, Cao W, Liu J, Zu X, Chen L. Mechanisms of NAT10 as ac4C writer in diseases. Mol Ther Nucleic Acids. 2023;32:359–68.
Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144:646–74.
Sang LJ, Ju HQ, Liu GP, Tian T, Ma GL, Lu YX, et al. LncRNA CamK-A regulates Ca(2+)-signaling-mediated tumor microenvironment remodeling. Mol Cell. 2018;72:71–83.e7.
Sun CC, Zhu W, Li SJ, Hu W, Zhang J, Zhuo Y, et al. FOXC1-mediated LINC00301 facilitates tumor progression and triggers an immune-suppressing microenvironment in non-small cell lung cancer by regulating the HIF1α pathway. Genome Med. 2020;12:77.
Zitvogel L, Galluzzi L, Kepp O, Smyth MJ, Kroemer G. Type I interferons in anticancer immunity. Nat Rev Immunol. 2015;15:405–14.
Turan T, Kongpachith S, Halliwill K, McLaughlin RT, Binnewies M, Reddy D, et al. iBRIDGE: a data integration method to identify inflamed tumors from single-cell RNA-Seq data and differentiate cell type-specific markers of immune-cell infiltration. Cancer Immunol Res. 2023;11:732–46.
Mowat C, Mosley SR, Namdar A, Schiller D, Baker K. Anti-tumor immunity in mismatch repair-deficient colorectal cancers requires type I IFN-driven CCL5 and CXCL10. J Exp Med. 2021. https://doi.org/10.1084/jem.20210108.
Patin EC, Soulard D, Fleury S, Hassane M, Dombrowicz D, Faveeuw C, et al. Type I IFN receptor signaling controls IL7-dependent accumulation and activity of protumoral IL17A-producing γδT cells in breast cancer. Cancer Res. 2018;78:195–204.
Zhu Q, Dai Q, Zhao L, Zheng C, Li Q, Yuan Z, et al. Novel dual inhibitors of PARP and HDAC induce intratumoral STING-mediated antitumor immunity in triple-negative breast cancer. Cell Death Dis. 2024;15:10.
Wang H, **a L, Yao CC, Dong H, Yang Y, Li C, et al. NLRP4 negatively regulates type I interferon response and influences the outcome in anti-programmed cell death protein (PD)-1/PD-ligand 1 therapy. Cancer Sci. 2022;113:838–51.
Ao YQ, Gao J, ** C, Wang S, Zhang LC, Deng J, et al. ASCC3 promotes the immunosuppression and progression of non-small cell lung cancer by impairing the type I interferon response via CAND1-mediated ubiquitination inhibition of STAT3. J Immunother Cancer. 2023. https://doi.org/10.1136/jitc-2023-007766.
Gu C, Wang X, Wang K, **e F, Chen L, Ji H, et al. Cryoablation triggers type I interferon-dependent antitumor immunity and potentiates immunotherapy efficacy in lung cancer. J Immunother Cancer. 2024;12. https://doi.org/10.1136/jitc-2023-008386.
Gozgit JM, Vasbinder MM, Abo RP, Kunii K, Kuplast-Barr KG, Gui B, et al. PARP7 negatively regulates the type I interferon response in cancer cells and its inhibition triggers antitumor immunity. Cancer Cell. 2021;39:1214–26.e10.
Chin EN, Yu C, Vartabedian VF, Jia Y, Kumar M, Gamo AM, et al. Antitumor activity of a systemic STING-activating non-nucleotide cGAMP mimetic. Science. 2020;369:993–9.
Vacchelli E, Bloy N, Aranda F, Buqué A, Cremer I, Demaria S, et al. Trial watch: immunotherapy plus radiation therapy for oncological indications. Oncoimmunology. 2016;5:e1214790.
Harada H, Kitagawa M, Tanaka N, Yamamoto H, Harada K, Ishihara M, et al. Anti-oncogenic and oncogenic potentials of interferon regulatory factors-1 and -2. Science. 1993;259:971–4.
Matsuyama T, Kimura T, Kitagawa M, Pfeffer K, Kawakami T, Watanabe N, et al. Targeted disruption of IRF-1 or IRF-2 results in abnormal type I IFN gene induction and aberrant lymphocyte development. Cell. 1993;75:83–97.
Yang Z, Sun JK, Lee MM, Chan MK. Restoration of p53 activity via intracellular protein delivery sensitizes triple negative breast cancer to anti-PD-1 immunotherapy. J Immunother Cancer. 2022;10. https://doi.org/10.1136/jitc-2022-005068.
Kumazawa T, Nishimura K, Katagiri N, Hashimoto S, Hayashi Y, Kimura K. Gradual reduction in rRNA transcription triggers p53 acetylation and apoptosis via MYBBP1A. Sci Rep. 2015;5:10854.
Zhuang R, Zhang X, Lu D, Wang J, Zhuo J, Wei X, et al. lncRNA DRHC inhibits proliferation and invasion in hepatocellular carcinoma via c-Myb-regulated MEK/ERK signaling. Mol Carcinog. 2019;58:366–75.
Li XL, Subramanian M, Jones MF, Chaudhary R, Singh DK, Zong X, et al. Long noncoding RNA PURPL suppresses basal p53 levels and promotes tumorigenicity in colorectal cancer. Cell Rep. 2017;20:2408–23.
Wiener D, Schwartz S. The epitranscriptome beyond m(6)A. Nat Rev Genet. 2021;22:119–31.
Arango D, Sturgill D, Alhusaini N, Dillman AA, Sweet TJ, Hanson G, et al. Acetylation of cytidine in mRNA promotes translation efficiency. Cell. 2018;175:1872–86.e24.
Yu XM, Li SJ, Yao ZT, Xu JJ, Zheng CC, Liu ZC, et al. N4-acetylcytidine modification of lncRNA CTC-490G23.2 promotes cancer metastasis through interacting with PTBP1 to increase CD44 alternative splicing. Oncogene. 2023;42:1101–16.
Liu R, Wubulikasimu Z, Cai R, Meng F, Cui Q, Zhou Y, et al. NAT10-mediated N4-acetylcytidine mRNA modification regulates self-renewal in human embryonic stem cells. Nucleic Acids Res. 2023. https://doi.org/10.1093/nar/gkad628.
Liao L, He Y, Li SJ, Yu XM, Liu ZC, Liang YY, et al. Lysine 2-hydroxyisobutyrylation of NAT10 promotes cancer metastasis in an ac4C-dependent manner. Cell Res. 2023;33:355–71.
**e R, Cheng L, Huang M, Huang L, Chen Z, Zhang Q, et al. NAT10 drives cisplatin chemoresistance by enhancing ac4C-associated DNA repair in bladder cancer. Cancer Res. 2023;83:1666–83.
Yang W, Li HY, Wu YF, Mi RJ, Liu WZ, Shen X, et al. ac4C acetylation of RUNX2 catalyzed by NAT10 spurs osteogenesis of BMSCs and prevents ovariectomy-induced bone loss. Mol Ther Nucleic Acids. 2021;26:135–47.
Tsai K, Jaguva Vasudevan AA, Martinez Campos C, Emery A, Swanstrom R, Cullen BR. Acetylation of cytidine residues boosts HIV-1 gene expression by increasing viral RNA stability. Cell Host Microbe. 2020;28:306–12.e6.
Thalalla Gamage S, Sas-Chen A, Schwartz S, Meier JL. Quantitative nucleotide resolution profiling of RNA cytidine acetylation by ac4C-seq. Nat Protoc. 2021;16:2286–307.
Rizvi H, Sanchez-Vega F, La K, Chatila W, Jonsson P, Halpenny D, et al. Molecular determinants of response to anti-programmed cell death (PD)-1 and anti-programmed death-ligand 1 (PD-L1) blockade in patients with non-small-cell lung cancer profiled with targeted next-generation sequencing. J Clin Oncol. 2018;36:633–41.
Funding
This study was supported by grants from the National Natural Science Foundation of China (81770082 and 82273325) and the Natural Science Foundation of Jiangsu province (BE2019719).
Author information
Authors and Affiliations
Contributions
ZMW contributed to study design and acquisition, analysis, interpretation of data, and drafting of the manuscript. JL and HRH contributed to the study design and analysis, interpretation of data, and substantively revised the manuscript. LW contributed to the acquisition, analysis, interpretation of data, and drafting of the manuscript. TFL, ZFW, CLL, YMW, and JXL contributed to the acquisition of data and manuscript review. QPC, XYZ, LWH, and MXY contributed to the analysis and interpretation of data and manuscript review. HBL and YS contributed to the study concept and design and supervised the study.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
This study was approved by the Ethics Committee of **ling Hospital. All animal experiments were performed in accordance with a protocol approved by the Animal Care and Use Committee of **ling Hospital.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, Z., Luo, J., Huang, H. et al. NAT10-mediated upregulation of GAS5 facilitates immune cell infiltration in non-small cell lung cancer via the MYBBP1A-p53/IRF1/type I interferon signaling axis. Cell Death Discov. 10, 240 (2024). https://doi.org/10.1038/s41420-024-01997-2
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41420-024-01997-2
- Springer Nature Limited