Abstract
S100a8/a9, largely released by polymorphonuclear neutrophils (PMNs), belongs to the S100 family of calcium-binding proteins and plays a role in a variety of inflammatory diseases. Although S100a8/a9 has been reported to trigger endothelial cell apoptosis, the mechanisms of S100a8/a9-induced endothelial dysfunction during sepsis require in-depth research. We demonstrate that high expression levels of S100a8/a9 suppress Ndufa3 expression in mitochondrial complex I via downregulation of Nrf1 expression. Mitochondrial complex I deficiency contributes to NAD+-dependent Sirt1 suppression, which induces mitochondrial disorders, including excessive fission and blocked mitophagy, and mtDNA released from damaged mitochondria ultimately activates ZBP1-mediated PANoptosis in endothelial cells. Moreover, based on comprehensive scRNA-seq and bulk RNA-seq analyses, S100A8/A9hi neutrophils are closely associated with the circulating endothelial cell count (a useful marker of endothelial damage), and S100A8 is an independent risk factor for poor prognosis in sepsis patients.
Similar content being viewed by others
Introduction
Sepsis is a persistent systemic inflammatory condition caused by an extreme immune response to infection, and it is always accompanied by multiple organ dysfunction [1]. To date, sepsis and septic shock have remained leading causes of death in critically ill patients, and the mortality rate of 30-day septic shock is as high as 34.7% [2]. In the initial stage of infection, the endothelial barrier and antimicrobial substances released by immune cells can cooperatively hinder pathogen dissemination [3, 4]. However, overwhelming amounts of inflammatory mediators damage endothelial barrier integrity, which contributes to microcirculatory disturbance and end-organ injury, especially lung injury [5, 6].
During sepsis, neutrophils are often the first immune cells to be recruited to infected sites [7]. Calprotectin (S100a8/a9), a heterodimeric Ca2+-binding protein mainly released by neutrophils, is thought to induce prolonged inflammation and endothelial injury through binding with its receptors, including toll-like receptor 4 (TLR4) and receptor for advanced glycation end products (RAGE) [8,9,10]. According to previous studies, inflammatory mediators, such as S100a8/a9 and neutrophil extracellular traps (NETs, web-like DNA structures adorned with bactericidal proteins), tend to induce mitochondrial metabolic disturbance and disrupt mitochondrial homeostasis [11, 12].
Mitochondrial dynamics (fission and fusion) and mitophagy are vital for maintaining mitochondrial quantity and quality. Specifically, fusion allows mitochondria to transfer gene products for optimal function [13], while fission induces the isolation of impaired mitochondria for degradation through mitophagy (a selective form of autophagy) [14]. Failure at any stage results in the accumulation of damaged mitochondria in the cytoplasm and ultimately cell death.
Previously, the different cell death patterns were considered independent. However, increasing evidence emphasizes the extensive crosstalk among different cell death patterns, and these pathways may be intertwined. Therefore, a novel form of cell death, PANoptosis, was proposed in 2019 [15]. It is a coordinated cell death pattern that involves apoptosis, pyroptosis and necroptosis [16]. Several studies have shown that Z-DNA binding protein 1 (ZBP1) can sense cytosolic DNA and consequently activate PANoptosis [17].
Here, our study provides evidence that S100A8/A9hi neutrophils are present specifically in lung tissues from septic mice. High expression levels of S100a8/a9 induce mitochondrial disorders in endothelial cells, including excessive fission and blocked mitophagy, mainly through Ndufa3 suppression in mitochondrial complex I. Finally, mtDNA released from damaged mitochondria activates ZBP1-mediated PANoptosis.
Results
Increased numbers of S100A8/A9hi neutrophils are found in the lung tissues of sepsis model mice, and these cells exhibit enhanced interactions with endothelial cells
Since the lung is considered as one of the most susceptible organs to sepsis [18], we selected publicly available scRNA-seq data from the lung tissues of sham and CLP mice for analysis (Fig. 1A). First, to explore the mechanisms underlying the excessive immune response to infection, all immune cells were clustered and identified by their marker genes (Fig. 1B, Supplementary Table 4). According to the relative percentages of each cell type, neutrophils accounted for the largest proportion of immune cells and were considerably more abundant in the CLP group (Fig. 1C). Therefore, neutrophils were extracted and reclustered into five subgroups. We found that a special subgroup, which showed greater expression of S100A8 and S100A9 than the other groups, was present only in the lung tissues of septic mice and represented a large proportion of the total neutrophil population (Fig. 1D–G, Supplementary Fig. 1A−D). Additionally, pseudotime analysis revealed that S100A8/A9hi neutrophils were specifically in the late stage of differentiation, indicating that neutrophils were induced to differentiate into S100A8/A9hi neutrophils during sepsis progression (Fig. 1H). Due to the presence of this subpopulation, neutrophils in the CLP group exhibited increased expressions of S100A8 and S100A9 and exhibited increased immune function and metabolism (Fig. 1I, Supplementary Fig. 1E–K).
A The main workflow of the scRNA-seq; B The UMAP plot was based on scRNA-seq data, and it showed five identified immune cell types; C The sankey diagram showed the percentages of five immune cells in sham and CLP groups; D Five clusters of neutrophils were identified in the UMAP plot; E The changes of five neutrophil subclusters percentage were shown on the sankey diagram; F Volcano map showed upregulated genes in S100A8/A9hi neutrophils compared with other subpopulations; G The expression levels of S100A8 and S100A9 in five neutrophil subclusters on dot plot; H The prediction of neutrophil differentiation trajectories using pseudotime analysis; I The expression levels of S100A8 and S100A9 of neutrophils in sham and CLP groups; J, K The number and strength of interactions among neutrophils, endothelial cells and epithelial cells analyzed by CellChat; L The expression levels of ligand-receptor pairs analyzed by CellChat (Since there is no ligand-receptor pair with high expression among epithelial cells in sham group, the dot plot only shows the expressions of ligand-receptor pairs among epithelial cells in CLP group.); M, N The distribution of S100A8/A9hi neutrophils and NET-related gene+ neutrophils on the UMAP plot. Wilcoxon rank sum test was used for the comparison between two groups. *p < 0.05, **p < 0.01 versus sham group.
Previously, we demonstrated that neutrophils could disturb the metabolism of endothelial cells and alveolar epithelial cells to aggravate sepsis-induced acute lung injury (SI-ALI) [11, 19]. Consequently, we next explored the interactions among neutrophils, endothelial cells and epithelial cells by using the “CellChat” R package. The results indicated that the number of interactions among these three cell types increased (Fig. 1J). Notably, from an interaction strength perspective, neutrophils showed enhanced unidirectional effects on endothelial cells, and endothelial cells also affected epithelial cells unidirectionally (Fig. 1K). These findings suggested that endothelial cells might act as a “bridge” in neutrophil-induced epithelial cell damage. We then focused on the ligand‒receptor pairs with upregulated expression in neutrophils and endothelial cells. The results indicated that thrombospondin 1 (Thbs1)-CD47 and Thbs1-CD36 expressions were clearly upregulated (Fig. 1L). Previous studies have revealed that endothelial cells exhibit increased Thbs1 expression after treatment with S100a8/a9 [20]. The above results suggest that neutrophils induce endothelial damage mainly through the release of S100a8/a9.
In addition, our previous study revealed that neutrophil extracellular traps (NETs) released by neutrophils might also damage the endothelial barrier during sepsis [21]. Based on the 137 identified NET formation-related genes, GSEA suggested that neutrophils from the sepsis model mice were able to release more NETs [22] (Supplementary Fig. 1L, Supplementary Table 5). Moreover, the subpopulation with high expression of NET-related genes was mostly composed of S100A8/A9hi neutrophils (Fig. 1M, N, Supplementary Fig. 1M-N). These results indicate that S100A8/A9hi neutrophils, including those in the NET-related gene+ subgroup, play a critical role in endothelial injury during sepsis.
In conclusion, the results of the scRNA-seq data analysis suggest that the presence of S100A8/A9hi neutrophils, which exist especially in the lung tissues of septic model mice, might induce endothelial barrier damage to exacerbate lung injury during sepsis.
High expression levels of S100a8/a9 induce excessive inflammatory responses and acute lung injury, both of which are reversed by an S100a8/a9 inhibitor
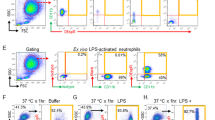
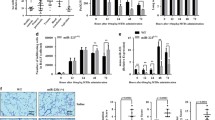
Since S100a8/a9 usually exists as a dimer, we evaluated the number of S100A9+ neutrophils by flow cytometry. The results demonstrated that the percentage of S100A9+ neutrophils in the peripheral blood were significantly increased in septic mice compared to control mice (Fig. 2A, B). According to previous studies, the peak of lung vascular injury occurred at 24 h after CLP [57]. Exclusion criteria included: a history of cardiopulmonary arrest before being admitted to ICU; history of connective tissue diseases such as systemic lupus erythematosus (SLE), vascular embolism and pregnancy.
Cecal ligation and puncture (CLP) mouse model
Eight- to ten-week-old male C57BL/6 mice were used for experiment. Following random grou**, the cecal ligation and puncture (CLP) mice model was established using the procedures described in previous studies [58]. In brief, after intraperitoneal anesthesia with 1% pentobarbital sodium (1 mg/kg), the abdominal cavity was opened. The cecum was carefully separated, ligated using 5-0 suture, and punctured with a 20-gauge needle. Next, we extruded a small quantity of feces from cecum and repositioned it before closing abdominal cavity. Each animal received 0.5 ml/10 g of normal saline for rehydration. The sham group received the same surgery without CLP. When mice required treatments, the following drugs were injected intraperitoneally: Paquinimod (10 mg/kg, MCE, Shanghai, China), β-nicotinamide mononucleotide (NMN, 500 mg/kg, MCE, Shanghai, China), Selisistat (EX-527, 5 mg/kg, MCE, Shanghai, China) [46, 59].
Murine sepsis score (MSS)
Seven clinical variables in MSS, including appearance, level of consciousness, activity, response to stimulus, eyes, respiration rate and respiration quality were used to assess the severity of sepsis. Each indicator has a score of 0-4, with a full score of 28. Higher scores mean more severe injury [60, 61].
Flow cytometry
Blood samples from mice were treated with red blood cell lysis buffer [46, 59] (Thermo Fisher). After centrifugation, the supernatant was discarded. The prepared cell suspensions were stained in PBS with the following antibodies: APC anti-mouse/human CD11b (BioLegend, 101211), FITC anti-mouse Ly-6G (BioLegend, 127605), PE anti-S100A9 (Cell Signaling Technology, #93941). Flow cytometry was performed using a BD FACS Aria III flow cytometer according to manufacturer protocol.
Histopathological analysis
Mice were sacrificed for several organs, such as lung, liver, kidney, spleen and intestine, and these tissues were fixed with 4% paraformaldehyde at room temperature for 24 h. Paraffin-embedded tissue sections were stained with hematoxylin and eosin (H&E). According to previous descriptions, the severity of acute lung injury was evaluated through a semiquantitative histology scoring method [62]. Specifically, the score of lung injury was based on these indicators: leukocyte infiltration, alveolar edema, haemorrhage and the thickness of alveolar septa. Two pathologists who blinded to the results graded each indicator from 0 to 3 (0 = normal; 1 = mild; 2 = moderate; 3 = severe), and finally calculated the total lung injury score. Furthermore, paraffin-embedded lung tissue sections were also stained with Masson dye to identify the degree of fibrosis within lung tissues.
TUNEL staining
Paraffin-embedded lung tissue sections were stained with TUNEL to detect cell apoptosis according to manufacturer protocol.
Lung wet-to-dry ratio
We harvested the left lung of mice and obtained its wet weight after drying the surface water. Subsequently, the tissue was dried at 70 °C for 48 h, and the dry weight was acquired. The wet/dry ratio was calculated by dividing the wet weight with dry weight. High wet-to-dry ratio means more severe lung edema.
Semi-quantification of inflammatory mediators
The levels of IL-1β, IL-6 and TNF-α in mouse serum were evaluated by Mouse IL-1β ELISA kit (mIC50300-1, mlbio, Shanghai, China), Mouse IL-6 ELISA kit (IC50325-1, mlbio, Shanghai, China), Mouse TNF-α ELISA kit (mIC50536-1, mlbio, Shanghai, China). Moreover, the levels of S100a8/a9 in mouse serum and human plasma were detected by Mouse S100a8/a9 ELISA kit (ml037985, mlbio, Shanghai, China) and Human S100a8/a9 ELISA kit (ml038517, mlbio, Shanghai, China).
Cell culture and treatments
We obtained Human Umbilical Vein Endothelial Cells (HUVECs) from American Type Culture Collection (ATCC; Manassas, USA) and cultured them in DMEM (Gibco) containing 10% fetal bovine serum (Gibco) and penicillin/streptomycin (Gibco) at 37 °C, 5% CO2 incubator. S100a8&a9 heterodimer protein was purchased from SinoBiological (Bei**g, China). Sirt1 activator SRT1720 was purchased from MCE (5 µM, Shanghai, China) [63]. HUVECs were transfected with Lentivirus-NC or Lentivirus-NRF1 (MOI = 10, Shanghai GeneChem, China) for 72 h to overexpress NRF1, and the sequence was listed in Supplementary Table 3.
Cell viability
A Cell-Counting Kit 8 (Do**do Corp., Kumamoto, Japan) was used to measure relative cell viability based on manufacturer’s instructions.
The Ad-mCherry-GFP-LC3B fluorescence microscopy assay
HUVECs were seeded in 48-well plates (5 × 104 cells/well) one day before transfection. Cells were transfected with adenovirus expressing mCherry-GFP-LC3B fusion protein (MOI = 20, Beyotime, Shanghai, China) for 24 h and then treated with or without S100a8/a9. The images were taken by Olympus microscope. In the absence of autophagy, mCherry-GFP-LC3B under fluorescence microscopy showed dispersed yellow fluorescence. However, in the presence of autophagy, mCherry-GFP-LC3B aggregated on the membrane of autophagosome showing yellow spots. After the fusion between autophagosomes and lysosomes, red dots could be observed, since GFP fluorescence was quenched partially.
Mitochondrial membrane potential assay kit with TMRE
HUVECs were seeded in 48-well plates (2 × 104 cells/well) with or without S100a8/a9 stimulation for 24 h. Then these cells were incubated with TMRE staining working solution (Beyotime, Shanghai, China) at 37 °C, 5% CO2 incubator for 30 min. After that, the supernatant was removed and the cells were washed with medium twice. The samples were then observed under immunofluorescence microscopy. Loss of mitochondria membrane potential was shown by diminished red fluorescence, which occurred in the early stage of cell apoptosis.
Measurement of mitochondrial oxidation
The oxygen consumption rate (OCR) of HUVECs was measured by the Agilent Seahorse XF Cell Mito Stress Test on the Seahorse XFe and XF Extracellular Flux Analyzers. HUVECs (2 × 104 cells/well) were seeded in an XF96 plate and incubated in a medium containing glucose, pyruvate and glutamine. Oligomycin, FCCP and rotenone were used to evaluate OCR. Seahorse Wave software was used to assess all data.
Transmission electron microscopy (TEM)
HUVECs were fixed in 2.5% glutaraldehyde, and then were post-fixed with 1% osmic acid for 2 h. Next, gradient dehydration was performed by the usage of graded ethanol. The sample was embedded in 812 resin, which followed by thin section staining of 2% uranyl acetate. Finally, the ultrastructural images of mitochondria were acquired by the usage of transmission electron microscope (HT7700, Hitachi).
Cytosolic mtDNA isolation
After lysis, HUVECs were centrifuged at 700 × g for 10 min to remove nuclei. Then, we normalized the supernatant volume based on protein concentration. Cell lysate was further centrifuged at 10,000 × g for 30 min for cytosolic fraction isolation, which included mtDNA and nDNA [17]. mtDNA was assessed by RT-qPCR with gene sequences coding for human NADH dehydrogenase 1 as primers. Nuclear DNA was detected using sequences coding human b-globin as primers [23]. The primers for human NADH dehydrogenase 1 and human b-globin were listed in Supplementary Table 2.
NAD+/NADH measurement
NAD+/NADH Assay Kit with WST-8 (Beyotime, Shanghai, China) was used for measure mitochondrial complex I activity. HUVECs were seeded in a 6-well plate (1 × 106 cells/well) before they were lysed with 200 μl NAD+/NADH extraction solution. Then, 100 μl samples were added in centrifugal tubes and heated at 60 °C for 30 min to decompose NAD+. Supernatant was further mixed with working solution and the absorbance of samples was measured at 450 nm.
Immunofluorescence
HUVECs were seeded in 48-well plates (2 × 104 cells/well) with or without S100a8/a9 stimulation. 4% paraformaldehyde was used for fixation for 10 min. The cells were penetrated by using 0.1% Triton for 5 min and further blocked at room temperature for 30 min. Antibody against Ndufa3 (1:500, sc-365351, Santa Cruz Biotechnology), was used for incubation overnight at 4 °C, and Alexa Fluor® 594-conjugated goat anti-rabbit IgG (1:200, ab150080, Abcam) was used to incubation at room temperature for 1 h next day. Finally, the nuclei were stained with 4,6-diamidino-2-phenylindole (DAPI). In order to visualize the expressions of target proteins in endothelial cells from mice lung tissues, paraffin-embedded tissue sections were deparaffinized, rehydrated for antigen retrieval. The primary antibodies used in this study included anti-S100a8 (1:500, GB11421-100, Servicebio), anti-S100a9 (1:750, GB111149-100, Servicebio), anti-CD31 (1:200, GB12063-100, Servicebio), anti-Ndufa3 (1:500, sc-365351, Santa Cruz Biotechnology), anti-LAMP1 (1:500, sc-20011, Santa Cruz Biotechnology); anti-ZBP1 (1:500, 13285-1-AP, Proteintech). And the secondary antibodies used here included iF488-Tyramide (1:500, G1231-50UL, Servicebio) and iF555-Tyramide (1:500, G1233-50UL, Servicebio).
Quantitative real-time PCR
The total RNA from cells and tissues was extracted by using TRIzol reagent (Thermo Fisher), and the quality and quantity of RNA were measured by NanoDropTM ND-1000. PrimeScript RT reagent kit (RR036A, Takara, Shinga, Japan) was used to reverse-transcribe RNA into cDNA. Then we used TB Green PCR kit (RR820A, Takara) and Bio-Rad system to perform RT-qPCR with two repetitions per well. The primer sequences were listed in Supplementary Table 2.
Western blot
Cells and lung tissues were lysed with RIPA Buffer (Solarbio, Bei**g, China), which contains proteinase inhibitor cocktails. Sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) were used to separate proteins. And then proteins were transferred to polyvinylidene fluoride (PVDF) membranes. The membranes were immersed in blocking buffer and incubated with primary antibodies against S100a8/a9 (1:1000, ab288715, Abcam); phospho-MEK1/2 (1:1000, #9154 S, Cell Signaling Technology); phospho-Erk/2 (1:2000, #4370, Cell Signaling Technology); PGC-1α (1:1000, #AF5395, Affinity); Nrf1 (1:1000, 66832-1-Ig, Proteintech); Total OXPHOS Rodent WB antibody cocktail (6.0 µg/ml, ab110413, Abcam); Sirt1 (1:500, sc-74465, Santa Cruz Biotechnology); phospho-Drp (1:1000, #3455, Cell Signaling Technology); Fis (1:500, sc-376447, Santa Cruz Biotechnology); Mfn1 (1:500, sc-166644, Santa Cruz Biotechnology); Mfn2 (1:500, sc-100560, Santa Cruz Biotechnology); LC3B (1:1000, #2775, Cell Signaling Technology); LAMP1 (1:500, sc-20011, Santa Cruz Biotechnology); ZBP1 (1:500; sc-271483, Santa Cruz Biotechnology); caspase 3 (1:500, sc-56053, Santa Cruz Biotechnology); cleaved caspase 3 (1:500, ab2302, Abcam); GSDMD (1:1000, ab219800, Abcam); N-terminal GSDMD (1:1000, ab215203, Abcam); MLKL (1:5000, 66675-1-Ig, Proteintech); phospho-MLKL (1:1000, ab196436, Abcam); phospho-MLKL (1:1000; ab187091, Abcam); GAPDH (1:1000, GB15004-100, Servicebio); β-Actin (1:1000, GB15003-100, Servicebio).
Bulk RNA-seq and scRNA-seq data
Bulk RNA-seq data from pneumonia-induced sepsis patients (GSE65682) and sepsis patients in ICU (GSE185263) were used to reanalyzed. scRNA-seq data of lung tissues from sham and CLP mice (GSE 207651) were selected for further analysis.
ScRNA-seq data pre-processing
We transferred merged matrix into the R statistical environment for further analysis through Seurat package (v. 4.0.4). Cells expressing <200 or >2500 genes, >5% mitochondrial reads were removed. “NormalizeData” function was performed to normalize the gene expression matrix, and 2000 highly variable genes (HVGs) were identified using “FindVariableFeatures” function. Then, the data were integrated among different samples based on identified anchor points using “FindIntegrationAnchors” function. Finally, we used “FindNeighbors” and FindCluster” functions to cluster and identify cells. And cell clusters were visualized by “RunTSNE” and “RunUMAP” functions.
Cluster marker identification and cell annotation
We identified the differentially expressed genes (DEGs) of each cluster using “FindAllMarkers” function, and the clusters were annotated based on classic marker genes [64].
Pseudotime analysis
We constructed differentiation trajectory using “Monocle 2” with DDRTree and the default parameter.
Pathway and functional enrichment analysis
Gene Ontology (GO) analysis was performed by “clusterProfiler” R package. Gene set enrichment analysis (GSEA) was performed by GSEA software. And gene set variation analysis (GSVA) scores were calculated through “gsva” function. We showed gene lists in supplementary tables.
Cell-cell communication analysis
“CellChat” package was used to evaluate the interactions between cells.
Correlation analysis
The correlation of genes in every endothelial subcluster was analyzed by “corrplot” R package.
The analysis of immune cells proportion
22 immune cells proportions were calculated by the CIBERSORT algorithm.
Survival analysis
“Survival” and “Survminer” R packages were used for Survival analysis. Sepsis patients were classified as “S100A8high” and “S100A8low” groups using “surv_cutpoint” function.
Logistic regression analysis
We took an intersection of marker genes of S100A8/A9hi neutrophils and gene lists from peripheral blood leukocytes of sepsis patients, and then conducted univariate logistic regression analysis. 24 genes (p < 0.001) were selected for multivariate logistic regression analysis to seek independent risk factors of high SOFA scores (>median of SOFA scores).
Cox regression analysis
We selected significantly upregulated marker genes in S100A8/A9hi neutrophils according to scRNA-seq data. Then, we took an intersection of marker genes of S100A8/A9hi neutrophils and gene lists from peripheral blood leukocytes of sepsis patients. The top 40 genes ordered by log2FC were chosen for univariate and multivariate cox regression analyses.
Statistical analysis
We carried out all statistical analyses using v.4.0.0 R and GraphPad Prism 8 software. Experimental data were expressed as means ± standard error of the means (SEM). Unpaired t-test and Wilcoxon rank sum test were used for the comparison between two groups, and one-way ANOVA was used for three or more groups. p < 0.05 was considered statistically significant (*/#/▲p < 0.05, **/##/▲▲p < 0.01).
Data availability
All experimental data are available and requested to Professor Changhong Miao.
References
Wang X, Ding Y, Li R, Zhang R, Ge X, Gao R, et al. N6-methyladenosine of Spi2a attenuates inflammation and sepsis-associated myocardial dysfunction in mice. Nat Commun. 2023;14:1185.
Bauer M, Gerlach H, Vogelmann T, Preissing F, Stiefel J, Adam D. Mortality in sepsis and septic shock in Europe, North America and Australia between 2009 and 2019- results from a systematic review and meta-analysis. Crit Care. 2020;24:239.
Lu SL, Omori H, Zhou Y, Lin YS, Liu CC, Wu JJ, et al. VEGF-mediated augmentation of autophagic and lysosomal activity in endothelial cells defends against intracellular streptococcus pyogenes. mBio. 2022;13:e0123322.
Gautier T, Deckert V, Nguyen M, Desrumaux C, Masson D, Lagrost L. New therapeutic horizons for plasma phospholipid transfer protein (PLTP): targeting endotoxemia, infection and sepsis. Pharmacol Ther. 2022;236:108105.
Barichello T, Generoso JS, Singer M, Dal-Pizzol F. Biomarkers for sepsis: more than just fever and leukocytosis-a narrative review. Crit Care. 2022;26:14.
Jiang J, Huang K, Xu S, Garcia JGN, Wang C, Cai H. Targeting NOX4 alleviates sepsis-induced acute lung injury via attenuation of redox-sensitive activation of CaMKII/ERK1/2/MLCK and endothelial cell barrier dysfunction. Redox Biol. 2020;36:101638.
Wang Y, Zhu CL, Li P, Liu Q, Li HR, Yu CM, et al. The role of G protein-coupled receptor in neutrophil dysfunction during sepsis-induced acute respiratory distress syndrome. Front Immunol. 2023;14:1112196.
Tousif S, Singh AP, Umbarkar P, Galindo C, Wheeler N, Toro Cora A, et al. Ponatinib drives cardiotoxicity by S100A8/A9-NLRP3-IL-1β mediated inflammation. Circ Res. 2023;132:267–89.
Viemann D, Barczyk K, Vogl T, Fischer U, Sunderkötter C, Schulze-Osthoff K, et al. MRP8/MRP14 impairs endothelial integrity and induces a caspase-dependent and -independent cell death program. Blood. 2007;109:2453–60.
Kovačić M, Mitrović-Ajtić O, Beleslin-Čokić B, Djikić D, Subotički T, Diklić M, et al. TLR4 and RAGE conversely mediate pro-inflammatory S100A8/9-mediated inhibition of proliferation-linked signaling in myeloproliferative neoplasms. Cell Oncol (Dordr). 2018;41:541–53.
Zhang H, Wu D, Wang Y, Guo K, Spencer CB, Ortoga L, et al. METTL3-mediated N6-methyladenosine exacerbates ferroptosis via m6A-IGF2BP2-dependent mitochondrial metabolic reprogramming in sepsis-induced acute lung injury. Clin Transl Med. 2023;13:e1389.
Li Y, Chen B, Yang X, Zhang C, Jiao Y, Li P, et al. S100a8/a9 signaling causes mitochondrial dysfunction and cardiomyocyte death in response to ischemic/reperfusion injury. Circulation [Internet]. 2019 Aug [cited 2024 Jan 8];140. Available from: https://pubmed.ncbi.nlm.nih.gov/31220942/.
Adebayo M, Singh S, Singh AP, Dasgupta S. Mitochondrial fusion and fission: the fine-tune balance for cellular homeostasis. FASEB J. 2021;35:e21620.
Cai C, Wu F, He J, Zhang Y, Shi N, Peng X, et al. Mitochondrial quality control in diabetic cardiomyopathy: from molecular mechanisms to therapeutic strategies. Int J Biol Sci [Internet]. 2022 Aug [cited 2024 Jan 8];18. Available from: https://pubmed.ncbi.nlm.nih.gov/36147470/.
Malireddi RKS, Kesavardhana S, Kanneganti TD. ZBP1 and TAK1: master regulators of NLRP3 inflammasome/pyroptosis, apoptosis, and necroptosis (PAN-optosis). Front Cell Infect Microbiol. 2019;9:406.
Lee S, Karki R, Wang Y, Nguyen LN, Kalathur RC, Kanneganti TD. AIM2 forms a complex with pyrin and ZBP1 to drive PANoptosis and host defence. Nature. 2021;597:415–9.
Bi Y, Xu H, Wang X, Zhu H, Ge J, Ren J, et al. FUNDC1 protects against doxorubicin-induced cardiomyocyte PANoptosis through stabilizing mtDNA via interaction with TUFM. Cell Death Dis. 2022;13:1020.
Park I, Kim M, Choe K, Song E, Seo H, Hwang Y, et al. Neutrophils disturb pulmonary microcirculation in sepsis-induced acute lung injury. Eur Respir J. 2019;53:1800786.
Zhang H, Wang Y, Qu M, Li W, Wu D, Cata JP, et al. Neutrophil, neutrophil extracellular traps and endothelial cell dysfunction in sepsis. Clin Transl Med. 2023;13:e1170.
Viemann D, Strey A, Janning A, Jurk K, Klimmek K, Vogl T, et al. Myeloid-related proteins 8 and 14 induce a specific inflammatory response in human microvascular endothelial cells. Blood. 2005;105:2955–62.
Zhu S, Yu Y, Qu M, Qiu Z, Zhang H, Miao C, et al. Neutrophil extracellular traps contribute to immunothrombosis formation via the STING pathway in sepsis-associated lung injury. Cell Death Discov. 2023;9:315.
Wu J, Zhang F, Zheng X, Zhang J, Cao P, Sun Z, et al. Identification of renal ischemia reperfusion injury subtypes and predictive strategies for delayed graft function and graft survival based on neutrophil extracellular trap-related genes. Front Immunol. 2022;13:1047367.
Huang LS, Hong Z, Wu W, **ong S, Zhong M, Gao X, et al. mtDNA activates cGAS signaling and suppresses the YAP-mediated endothelial cell proliferation program to promote inflammatory injury. Immunity. 2020;52:475–486.e5.
Pan X, Xu S, Zhou Z, Wang F, Mao L, Li H, et al. Fibroblast growth factor-2 alleviates the capillary leakage and inflammation in sepsis. Mol Med. 2020;26:108.
**a S, Menden HL, Korfhagen TR, Kume T, Sampath V. Endothelial immune activation programmes cell-fate decisions and angiogenesis by inducing angiogenesis regulator DLL4 through TLR4-ERK-FOXC2 signalling. J Physiol. 2018;596:1397–417.
Finelli MJ, Murphy KJ, Chen L, Zou H. Differential phosphorylation of Smad1 integrates BMP and neurotrophin pathways through Erk/Dusp in axon development. Cell Rep. 2013;3:1592–606.
Sihag S, Cresci S, Li AY, Sucharov CC, Lehman JJ. PGC-1alpha and ERRalpha target gene downregulation is a signature of the failing human heart. J Mol Cell Cardiol. 2009;46:201–12.
Liu L, Li Y, Wang J, Zhang D, Wu H, Li W, et al. Mitophagy receptor FUNDC1 is regulated by PGC-1α/NRF1 to fine tune mitochondrial homeostasis. EMBO Rep. 2021;22:e50629.
Fan H, Ding R, Liu W, Zhang X, Li R, Wei B, et al. Heat shock protein 22 modulates NRF1/TFAM-dependent mitochondrial biogenesis and DRP1-sparked mitochondrial apoptosis through AMPK-PGC1α signaling pathway to alleviate the early brain injury of subarachnoid hemorrhage in rats. Redox Biol. 2021;40:101856.
Klinge CM. Estrogenic control of mitochondrial function. Redox Biol. 2020;31:101435.
Hao L, Zhong W, Dong H, Guo W, Sun X, Zhang W, et al. ATF4 activation promotes hepatic mitochondrial dysfunction by repressing NRF1-TFAM signalling in alcoholic steatohepatitis. Gut. 2021;70:1933–45.
McElroy GS, Reczek CR, Reyfman PA, Mithal DS, Horbinski CM, Chandel NS. NAD+ regeneration rescues lifespan, but not ataxia, in a mouse model of brain mitochondrial complex I dysfunction. Cell Metab. 2020;32:301–08.e6.
Qu M, Chen Z, Qiu Z, Nan K, Wang Y, Shi Y, et al. Neutrophil extracellular traps-triggered impaired autophagic flux via METTL3 underlies sepsis-associated acute lung injury. Cell Death Discov. 2022;8:375.
Sharma A, Ahmad S, Ahmad T, Ali S, Syed MA. Mitochondrial dynamics and mitophagy in lung disorders. Life Sci. 2021;284:119876.
Zhao YG, Codogno P, Zhang H. Machinery, regulation and pathophysiological implications of autophagosome maturation. Nat Rev Mol Cell Biol. 2021;22:733–50.
Huang G, Xu X, Ju C, Zhong N, He J, Tang XX. Identification and validation of autophagy-related gene expression for predicting prognosis in patients with idiopathic pulmonary fibrosis. Front Immunol. 2022;13:997138.
Yang YY, Gao ZX, Mao ZH, Liu DW, Liu ZS, Wu P. Identification of ULK1 as a novel mitophagy-related gene in diabetic nephropathy. Front Endocrinol (Lausanne). 2022;13:1079465.
Ren H, Kang N, Yin S, Xu C, Qu T, Dai D. Characteristic of molecular subtypes based on PANoptosis-related genes and experimental verification of hepatocellular carcinoma. Aging (Albany NY). 2023;15:4159–81.
Moussa MD, Santonocito C, Fagnoul D, Donadello K, Pradier O, Gaussem P, et al. Evaluation of endothelial damage in sepsis-related ARDS using circulating endothelial cells. Intensive Care Med. 2015;41:231–8.
Schlichting DE, Waxman AB, O’Brien LA, Wang T, Naum CC, Rubeiz GJ, et al. Circulating endothelial and endothelial progenitor cells in patients with severe sepsis. Microvasc Res. 2011;81:216–21.
Atreya MR, Cvijanovich NZ, Fitzgerald JC, Weiss SL, Bigham MT, Jain PN, et al. Prognostic and predictive value of endothelial dysfunction biomarkers in sepsis-associated acute kidney injury: risk-stratified analysis from a prospective observational cohort of pediatric septic shock. Crit Care. 2023;27:260.
Marinković G, Koenis DS, de Camp L, Jablonowski R, Graber N, de Waard V, et al. S100A9 links inflammation and repair in myocardial infarction. Circ Res. 2020;127:664–76.
Li C, Li S, Jia C, Yang L, Song Z, Wang Y. Low concentration of S100A8/9 promotes angiogenesis-related activity of vascular endothelial cells: bridges among inflammation, angiogenesis, and tumorigenesis? Med Inflamm. 2012;2012:248574.
Vercellino I, Sazanov LA. The assembly, regulation and function of the mitochondrial respiratory chain. Nat Rev Mol Cell Biol. 2022;23:141–61.
Grivennikova VG, Gladyshev GV, Vinogradov AD. Deactivation of mitochondrial NADH:ubiquinone oxidoreductase (respiratory complex I): Extrinsically affecting factors. Biochim Biophys Acta Bioenerg. 2020;1861:148207.
Li HR, Liu Q, Zhu CL, Sun XY, Sun CY, Yu CM, et al. β-Nicotinamide mononucleotide activates NAD+/SIRT1 pathway and attenuates inflammatory and oxidative responses in the hippocampus regions of septic mice. Redox Biol. 2023;63:102745.
Winnik S, Auwerx J, Sinclair DA, Matter CM. Protective effects of sirtuins in cardiovascular diseases: from bench to bedside. Eur Heart J. 2015;36:3404–12.
Kadono K, Kageyama S, Nakamura K, Hirao H, Ito T, Kojima H, et al. Myeloid Ikaros-SIRT1 signaling axis regulates hepatic inflammation and pyroptosis in ischemia-stressed mouse and human liver. J Hepatol. 2022;76:896–909.
Xu C, Wang L, Fozouni P, Evjen G, Chandra V, Jiang J, et al. SIRT1 is downregulated by autophagy in senescence and ageing. Nat Cell Biol. 2020;22:1170–9.
Zhang Y, Wang Y, Xu J, Tian F, Hu S, Chen Y, et al. Melatonin attenuates myocardial ischemia-reperfusion injury via improving mitochondrial fusion/mitophagy and activating the AMPK-OPA1 signaling pathways. J Pineal Res. 2019;66:e12542.
Chen W, Zhao H, Li Y. Mitochondrial dynamics in health and disease: mechanisms and potential targets. Signal Transduct Target Ther. 2023;8:333.
Hu Q, Zhang H, Gutiérrez Cortés N, Wu D, Wang P, Zhang J, et al. Increased Drp1 acetylation by lipid overload induces cardiomyocyte death and heart dysfunction. Circ Res. 2020;126:456–70.
Jiao H, Wachsmuth L, Kumari S, Schwarzer R, Lin J, Eren RO, et al. Z-nucleic-acid sensing triggers ZBP1-dependent necroptosis and inflammation. Nature. 2020;580:391–5.
Koehler H, Cotsmire S, Zhang T, Balachandran S, Upton JW, Langland J, et al. Vaccinia virus E3 prevents sensing of Z-RNA to block ZBP1-dependent necroptosis. Cell Host Microbe. 2021;29:1266–76.e5.
Lei Y, VanPortfliet JJ, Chen YF, Bryant JD, Li Y, Fails D, et al. Cooperative sensing of mitochondrial DNA by ZBP1 and cGAS promotes cardiotoxicity. Cell. 2023;186:3013–32.e22.
Sundaram B, Pandian N, Mall R, Wang Y, Sarkar R, Kim HJ, et al. NLRP12-PANoptosome activates PANoptosis and pathology in response to heme and PAMPs. Cell. 2023;186:2783–2801.e20.
Shankar-Hari M, Phillips GS, Levy ML, Seymour CW, Liu VX, Deutschman CS, et al. Develo** a new definition and assessing new clinical criteria for septic shock: for the third international consensus definitions for sepsis and septic shock (Sepsis-3). JAMA. 2016;315:775–87.
Zhang H, Liu J, Zhou Y, Qu M, Wang Y, Guo K, et al. Neutrophil extracellular traps mediate m6A modification and regulates sepsis-associated acute lung injury by activating ferroptosis in alveolar epithelial cells. Int J Biol Sci. 2022;18:3337–57.
Wu F, Zhang YT, Teng F, Li HH, Guo SB. S100a8/a9 contributes to sepsis-induced cardiomyopathy by activating ERK1/2-Drp1-mediated mitochondrial fission and respiratory dysfunction. Int Immunopharmacol. 2023;115:109716.
Deng F, Hu JJ, Lin ZB, Sun QS, Min Y, Zhao BC, et al. Gut microbe-derived milnacipran enhances tolerance to gut ischemia/reperfusion injury. Cell Rep Med. 2023;4:100979.
Shrum B, Anantha RV, Xu SX, Donnelly M, Haeryfar SMM, McCormick JK, et al. A robust scoring system to evaluate sepsis severity in an animal model. BMC Res Notes. 2014;7:233.
Li H, Li Y, Song C, Hu Y, Dai M, Liu B, et al. Neutrophil extracellular traps augmented alveolar macrophage pyroptosis via AIM2 inflammasome activation in LPS-induced ALI/ARDS. J Inflamm Res. 2021;14:4839–58.
Wu Q, Hu Y, Jiang M, Wang F, Gong G. Effect of autophagy regulated by Sirt1/FoxO1 pathway on the release of factors promoting thrombosis from vascular endothelial cells. Int J Mol Sci. 2019;20:4132.
Wang F, Chen M, Ma J, Wang C, Wang J, **a H, et al. Integrating bulk and single-cell sequencing reveals the phenotype-associated cell subpopulations in sepsis-induced acute lung injury. Front Immunol. 2022;13:981784.
Acknowledgements
This research was supported by Shanghai Municipal 2021 “Science and Technology Innovation Action Plan” (No. 21JC1401400), Natural Science Foundation of Shanghai (No. 21ZR1413400), National Natural Science Foundation of China (No. 82102253) and Shanghai Sailing Program (No. 21YF1406800), Shanghai Pujiang Talent Program (23PJD013).
Author information
Authors and Affiliations
Contributions
HZ and CHM contributed to the conception and funding of the study. YHZW completed most of the experiments, collated and analyzed data, and wrote the manuscript. YXS and YWS conducted several experiments. XHL participated in data collection and correction. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by Stephen Tait
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, Y., Shi, Y., Shao, Y. et al. S100A8/A9hi neutrophils induce mitochondrial dysfunction and PANoptosis in endothelial cells via mitochondrial complex I deficiency during sepsis. Cell Death Dis 15, 462 (2024). https://doi.org/10.1038/s41419-024-06849-6
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41419-024-06849-6
- Springer Nature Limited