Key Points
-
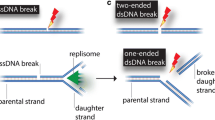
DNA double-strand breaks (DSBs) pose an acute threat to the survival of all cells. There are two major pathways of DSB repair — homologous recombination (HR) and non-homologous end joining (NHEJ).
-
In HR, one or both of the DNA ends are resected by an exonuclease to generate a 3′ single-strand extension that invades the intact sister chromatid. The invading strand serves as a primer for DNA synthesis. HR is generally error-free, which allows the rescue of collapsed DNA-replication forks and provides a defence against exogenous DNA-damaging agents. In NHEJ, there is no requirement for a homologous DNA template. Instead, DSBs are approximated by the DNA-end-binding protein Ku and then at least one of the broken ends is sealed by a specialized DNA ligase. NHEJ can be faithful or mutagenic, depending on whether the ends are sealed directly or remodelled by nucleases or polymerases before sealing. The central role of a dedicated DNA ligase is a distinctive feature of NHEJ.
-
For many years it was thought that bacteria contained only a single DNA ligase and relied solely on HR for DSB repair. However, over the past five years this has been overturned by evidence that many bacterial genera, including Mycobacterium, Pseudomonas, Bacillus and Agrobacterium, contain an NHEJ system that depends on Ku and the dedicated DNA ligase LigD. Biochemical, structural and genetic studies of the bacterial ligases and Ku proteins are beginning to define an NHEJ pathway that has distinctive features and enzymatic components.
-
LigD is a large, multifunctional enzyme that differs from all other DNA ligases in that it has multiple catalytic activities in a single polypeptide. LigD consists of an ATP-dependent ligase domain, a polymerase (POL) domain and a phosphoesterase domain. The minimal LigC ligases have been characterized only in mycobacteria and Agrobacterium tumefaciens, and comprise only nucleotidyltransferase (NTase) and oligonucleotide-binding (OB) domains. The complexity of the bacterial NHEJ apparatus ranges from the simple state found in Pseudomonas aeruginosa (which has one Ku, one LigD and no LigC), to progressively more complex forms, such as that found in Mycobacterium tuberculosis (which has one Ku, one LigD and one LigC), Mycobacterium smegmatis (which has one Ku, one LigD and two LigCs) and A. tumefaciens (which has three Ku paralogues, two LigDs and three LigCs).
-
Bacterial NHEJ has been studied in most detail in mycobacteria, in which the fate of DSBs that have been repaired by mycobacterial NHEJ has illuminated the broad outlines and distinctive features of the bacterial NHEJ pathway.
-
The crystal structures of the POL domains of P. aeruginosa LigD bound to ATP and M. tuberculosis LigD bound to GTP have been solved.
Abstract
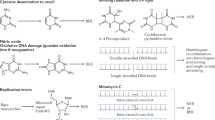
The capacity to rectify DNA double-strand breaks (DSBs) is crucial for the survival of all species. DSBs can be repaired either by homologous recombination (HR) or non-homologous end joining (NHEJ). The long-standing notion that bacteria rely solely on HR for DSB repair has been overturned by evidence that mycobacteria and other genera have an NHEJ system that depends on a dedicated DNA ligase, LigD, and the DNA-end-binding protein Ku. Recent studies have illuminated the role of NHEJ in protecting the bacterial chromosome against DSBs and other clastogenic stresses. There is also emerging evidence of functional crosstalk between bacterial NHEJ proteins and components of other DNA-repair pathways. Although still a young field, bacterial NHEJ promises to teach us a great deal about the nexus of DNA repair and bacterial pathogenesis.




Similar content being viewed by others
References
Cromie, G. A., Connelly, J. C. & Leach, D. R. F. Recombination at double-strand breaks and DNA ends: conserved mechanisms from phage to humans. Mol. Cell 8, 1163–1174 (2001).
Daley, J. M., Palmbos, P. L., Wu, D. & Wilson, T. E. Nonhomologous end joining in yeast. Annu. Rev. Genet. 39, 431–451 (2005).
Lieber, M. R., Yu, L. & Raghavan, S. C. Roles of nonhomologous DNA end joining, V(D)J recombination, and class switch recombination in chromosomal translocations. DNA Repair 5, 1234–1245 (2006).
Lusetti, S. L. & Cox, M. M. The bacterial RecA protein and the recombinational DNA repair of stalled replication forks. Annu. Rev. Biochem. 71, 71–100 (2002).
Takata, M. et al. Homologous recombination and nonhomologous end-joining pathways of DNA double-strand break repair have overlap** roles in the maintenance of chromosomal integrity in vertebrate cells. EMBO J. 17, 5497–5508 (1998).
Ferreira, M. G. & Cooper, J. P. Two modes of DNA double-strand break repair are reciprocally regulated through fission yeast cell cycle. Genes Dev. 18, 2249–2254 (2004).
Bertocci, B., De Smet, A., Weill, J. C. & Reynaud, C. A. Nonoverlap** functions of DNA polymerases Mu, Lambda, and terminal deoxynucleotidyltransferase during immunoglobulin V(D)J recombination in vivo. Immunity 25, 31–41 (2006).
Aravind, L. & Koonin, E. V. Prokaryotic homologs of the eukaryotic DNA end-binding protein Ku, novel domains in the Ku protein and prediction of a prokaryotic double-strand break repair system. Genome Res. 11, 1365–1374 (2001).
Doherty, A. J., Jackson, S. P. & Weller, G. R. Identification of bacterial homologues of the Ku DNA repair proteins. FEBS Lett. 500, 186–188 (2001).
d'Adda di Fagagna, F., Weller, G. R., Doherty, A. J. & Jackson, S. P. The Gam protein of bacteriophage Mu is an ortholog of eukaryotic Ku. EMBO Rep. 4, 47–52 (2003).
Cheng, C. & Shuman, S. Characterization of an ATP-dependent DNA ligase encoded by Haemophilus influenzae. Nucleic Acids Res. 25, 1369–1375 (1997).
Magnet, S. & Blanchard, J. S. Mechanistic and kinetic study of the ATP-dependent DNA ligase of Neisseria meningitidis. Biochemistry 43, 710–717 (2004).
Weller, G. R. et al. Identification of a DNA nonhomologous end-joining complex in bacteria. Science 297, 1686–1689 (2002). This paper comprises the first biochemical characterization of M. tuberculosis Ku as a homodimeric DNA-binding protein that stimulates DNA joining by LigD. Ku and LigD were found to be non-essential for the growth of B. subtilis . The deletion of Ku and LigD sensitized stationary phase B. subtilis to ionizing radiation, an effect that was subsequently shown (in Refs 19 & 20 ) to be specific to bacterial spores, which contain a single copy of the chromosome.
Gong, C., Martins, A., Bongiorno, P., Glickman, M. & Shuman, S. Biochemical and genetic analysis of the four DNA ligases of mycobacteria. J. Biol. Chem. 279, 20594–20606 (2004).
Weller, G. R. & Doherty, A. J. A family of DNA repair ligases in bacteria? FEBS Lett. 505, 340–342 (2001).
Della, M. et al. Mycobacterial Ku and ligase proteins constitute a two-component NHEJ repair machine. Science 306, 683–685 (2004). The authors characterize the polymerase activities of M. tuberculosis LigD and indicate a role for POL and ribonucleotides in gap repair before the sealing step of NHEJ in vitro.
Gong, C. et al. Mechanism of nonhomologous end-joining in mycobacteria: a low fidelity repair system driven by Ku, ligase D and ligase C. Nature Struct. Mol. Biol. 12, 304–312 (2005). This paper reports the first study of the NHEJ mechanism in a bacterium, including an assessment of the roles of Ku and ligase B, ligase C and ligase D in the efficiency and fidelity of the repair of blunt and 5′-overhang DNA ends in vivo . An analysis of the molecular outcomes of individual repair events highlighted the high frequency of frame-shift mutations at the repair junctions.
Korycka-Machala, M. et al. Distinct DNA repair pathways involving RecA and nonhomologous end joining in Mycobacterium smegmatis. FEMS Microbiol. Lett. 258, 83–91 (2006).
Wang, S. T. et al. The forespore line of gene expression in Bacillus subtilis. J. Mol. Biol. 358, 16–37 (2006).
Moeller, R. et al. Role of DNA repair by nonhomologous end joining in Bacillus subtilis spore resistance to extreme dryness, mono- and polychromatic UV, and ionizing radiation. J. Bacteriol. 189, 3306–3311 (2007).
Stephanou, N. C. et al. Mycobacterial nonhomologous end joining mediates mutagenic repair of chromosomal double-strand DNA breaks. J. Bacteriol. 189, 5237–5246 (2007).
Pitcher, R. S. et al. NHEJ protects mycobacteria in stationary phase against the harmful effects of dessication. DNA Repair 6, 1271–1276 (2007).
Zhu, H. & Shuman, S. A primer-dependent polymerase function of Pseudomonas aeruginosa ATP-dependent DNA ligase (LigD). J. Biol. Chem. 280, 418–427 (2005).
Zhu, H. & Shuman, S. Novel 3′-ribonuclease and 3′-phosphatase activities of the bacterial non-homologous end-joining protein, DNA ligase D. J. Biol. Chem. 280, 25973–25981 (2005). This paper reports the identification and characterization of the 3′ ribonucleotide-resection activity of the PE domain of P. aeruginosa LigD, which acts by a novel two-step mechanism.
Zhu, H. & Shuman, S. Characterization of Agrobacterium tumefaciens DNA ligases C and D. Nucleic Acids Res. 35, 3631–3645 (2007).
Akey, D. et al. Crystal structure and nonhomologous end joining function of the ligase domain of Mycobacterium DNA ligase D. J. Biol. Chem. 281, 13412–13423 (2006). This paper reports the 2.4 Å structure of the LigD ligase–AMP intermediate and provides a mutational analysis of the active site. A LigD mutation that specifically ablates the sealing function had only a modest effect on NHEJ efficiency in vivo , which implies the existence of an effective backup NHEJ ligase (probably LigC).
Zhu, H. et al. Atomic structure and nonhomologous end-joining function of the polymerase component of bacterial DNA ligase D. Proc. Natl Acad. Sci. USA 103, 1711–1716 (2006). The determination of the 1.5 Å structure of the POL domain of P. aeruginosa LigD highlighted its structural similarity to archaeal and eukaryal DNA primase. In this study a LigD mutation that abolished the POL activity eliminated non-templated additions during the repair of blunt DNA ends, thereby implicating LigD POL as the direct catalyst of mutagenic NHEJ in vivo.
Pitcher, R. S. et al. Structure and function of a mycobacterial NHEJ DNA repair polymerase. J. Mol. Biol. 366, 391–405 (2007).
Lehman, I. R. DNA ligase: structure, mechanism, and function. Science 186, 790–797 (1974).
Nandakumar, J., Nair, P. A. & Shuman, S. Last stop on the road to repair: structure of E. coli DNA ligase bond to nicked DNA-adenylate. Mol. Cell 26, 257–271 (2007).
Gajiwala, K. & Pinko, C. Structural rearrangement accompanying NAD+ synthesis within a bacterial DNA ligase crystal. Structure 12, 1449–1459 (2004).
Sriskanda, V. & Shuman, S. Conserved residues in domain Ia are required for the reaction of Escherichia coli DNA ligase with NAD+. J. Biol. Chem. 277, 9685–9700 (2002).
Pascal, J. M., O'Brien, P. J., Tomkinson, A. E. & Ellenberger, T. Human DNA ligase I completely encircles and partially unwinds nicked DNA. Nature 432, 473–478 (2004).
Pitcher, R. S., Tonkin, L. M., Green, A. J. & Doherty, A. J. Domain structure of a NHEJ repair ligase from Mycobacterium tuberculosis. J. Mol. Biol. 351, 531–544 (2005).
Yakovleva, L. & Shuman, S. Nucleotide misincorporation, 3′-mismatch extension, and responses to abasic sites and DNA adducts by the polymerase component of bacterial DNA ligase D. J. Biol. Chem. 281, 25026–25040 (2006).
Augustin, M. A., Huber, R. & Kaiser, J. T. Crystal structure of a DNA-dependent RNA polymerase (DNA primase). Nature Struct. Biol. 8, 57–61 (2001).
Ito, N., Nureki, O., Shirouzu, M., Yokoyama, S. & Hanaola, F. Crystal structure of the Pyrococcus horikoshii DNA primase-UTP complex: implications for the mechanism of primer synthesis. Genes Cells 8, 913–923 (2003).
Ramadan, K., Shevelev, I. & Hübscher, U. The DNA-polymerase-X family: controllers of DNA quality? Nature Rev. Mol. Cell. Biol. 5, 1038–1043 (2004).
Zhu, H. & Shuman, S. Essential constituents of the 3′-phosphoesterase domain of bacterial DNA ligase D, a nonhomologous end-joining enzyme. J. Biol. Chem. 280, 33707–33715 (2005).
Zhu, H. & Shuman, S. Substrate specificity and structure-function analysis of the 3′-phosphoesterase component of the bacterial NHEJ protein, DNA Ligase D. J. Biol. Chem. 281, 13873–13881 (2006).
Jacobs, M. A. et al. Comprehensive transposon mutant library of Pseudomonas aeruginosa. Proc. Natl Acad. Sci. USA 100, 14339–14344 (2003).
Darwin, K. H. & Nathan, C. F. Role for nucleotide excision repair in virulence of Mycobacterium tuberculosis. Infect. Immun. 73, 4581–4587 (2005).
Boschoff, H. I., Reed, M. B., Barry, C. E. & Mizrahi, V. DnaE2 polymerase contributes to in vivo survival and the emergence of drug resistance in Mycobacterium tuberculosis. Cell 113, 183–193 (2003).
Cirz, R. T. et al. Inhibition of mutation and combating the evolution of antibiotic resistance. PLoS Biol. 3, 1024–1033 (2005).
Gandhi, N. R. et al. Extensively drug-resistant tuberculosis as a cause of death in patients co-infected with tuberculosis and HIV in a rural area of South Africa. Lancet 368, 1575–1580 (2006).
Wayne, L. G. Synchronized replication of Mycobacterium tuberculosis. Infect. Immun. 17, 528–530 (1977).
Dick, T., Lee, B. H. & Murugasu-Oei, B. Oxygen depletion induced dormancy in Mycobacterium smegmatis. FEMS Microbiol. Lett. 163, 159–164 (1998).
Li, L. et al. Role of the non-homologous DNA end joining pathway in the early steps of retroviral infection. EMBO J. 20, 3272–3281 (2001).
Skalka, A. M. & Katz, R. A. Retroviral DNA integration and the DNA damage response. Cell Death Differ. 12, 871–978 (2005).
Muylaert, I. & Elias, P. Knockdown of DNA ligase IV/XRCC4 by RNA interference inhibits herpes simplex virus type I DNA replication. J. Biol. Chem. 282, 10865–10872 (2007).
Pitcher, R. S. et al. Mycobacteriophage exploit NHEJ to facilitate genome circularization. Mol. Cell 23, 743–748 (2006). This study shows that two DNA viruses that infect M. smegmatis encode homologues of Ku and depend on host-cell LigD for productive replication.
Martinez, J. J., Seveau, S., Veiga, S., Matsuyama, S. & Cossart, P. Ku70, a component of DNA-dependent protein kinase, is a mammalian receptor for Rickettsia conorii. Cell 123, 1013–1023 (2005).
Sinha, K. M., Stephanou, N. C., Gao, F., Glickman, M. S. & Shuman, S. Mycobacterial UvrD1 is a Ku-dependent DNA helicase that plays a role in multiple DNA repair events, including double-strand break repair. J. Biol. Chem. 282, 15114–15125 (2007).
Velankar, S. S., Soultanas, P., Dillingham, M. S., Subramanya, H. S. & Wigley, D. B. Crystal structure of complexes of PcrA DNA helicase with a DNA substrate indicate an inchworm mechanism. Cell 87, 75–84 (1999).
Lee, J. Y. & Yang, W. UvrD helicase unwinds DNA one base pair at a time by a two-part power stroke. Cell 127, 1349–1360 (2006).
Curti, E., Smerdon, S. J. & Davis, E. O. Characterization of the helicase activity and substrate specificity of Mycobacterium tuberculosis UvrD. J. Bacteriol. 189, 1542–1555 (2007).
Acknowledgements
NHEJ research in the laboratories of S.S. and M.S.G. is suported by National Institutes of Health grants A1064693 and GM63611.
Author information
Authors and Affiliations
Corresponding author
Related links
Related links
DATABASES
Entrez Genome Project
Protein Data Bank
Mycobacterium tuberculosis LigD LIG
Mycobacterium tuberculosis LigD POL
FURTHER INFORMATION
Glossary
- Abasic site
-
A common form of DNA damage in which a base is removed from a strand of DNA by the action of DNA-repair enzymes, such as uracil glycosylase, leaving the phosphodiester bond intact.
Rights and permissions
About this article
Cite this article
Shuman, S., Glickman, M. Bacterial DNA repair by non-homologous end joining. Nat Rev Microbiol 5, 852–861 (2007). https://doi.org/10.1038/nrmicro1768
Issue Date:
DOI: https://doi.org/10.1038/nrmicro1768
- Springer Nature Limited