Abstract
Background:
The Fat mass and obesity-associated gene (FTO) was the first gene reliably associated with body mass index in genome-wide association studies on a population level. At present, the genetic variations within the FTO gene are still the common variants that have the largest influence on body mass index.
Methods:
In the current study, we amplified the entire FTO gene, in total 412 Kbp, in over 200 long-range PCR fragments from each individual, from 524 severely obese and 527 lean Swedish children, and sequenced the products as two DNA pools using massive parallel sequencing (SOLiD).
Results:
The sequencing achieved very high coverage (median 18 000 reads) and we detected and estimated allele frequencies for 705 single nucleotide polymorphisms (SNPs) (19 novel) and 40 indels (24 novel) using a sophisticated statistical approach to remove false-positive SNPs. We identified 19 obesity-associated SNPs within intron one of the FTO gene, and validated our findings with genoty**. Ten of the validated obesity-associated SNPs have a stronger obesity association (P<0.007) than the commonly studied rs9939609 SNP (P<0.012).
Conclusions:
This study provides a comprehensive obesity-associated variation map of FTO, identifies novel lead SNPs and evaluates putative causative variants. We conclude that intron one is the only region within the FTO gene associated with obesity, and finally, we establish next generation sequencing of pooled DNA as a powerful method to investigate genetic association with complex diseases and traits.
Similar content being viewed by others
Introduction
SOLiD sequencing
Libraries were prepared from each pool and emulsion PCR was carried out according to the instructions from Applied Biosystems (Foster City, CA, USA). Sequencing was performed with Applied Biosystems' SOLiD 3 platform using a 50-bp read length on standard slides according to the protocol of the manufacturer. The reads were mapped to the FTO reference sequence (hg18, chr16:52 285 069-52 716 675) using the corona lite algorithm (Applied Biosystems) with default settings.
Variant calling and filtering
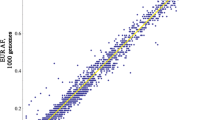
The methodology used for identification and filtration of false-positives SNPs, insertions and deletion (indels) are covered in greater detail in Supplementary Information 1. Briefly, SNPs are reported by the SOLiD system as valid adjacent miss-matches (that is, not sequencing errors) compared with the reference sequence. However, to limit the number of false-positive SNPs, we implemented a strategy that we previously used on data from SOLiD sequencing of pooled DNA.26 This method focuses on filtering excessively amplified reads containing valid adjacent miss-matches, probably derived from errors in previous PCR amplification steps and not true polymorphic sites. A quality score of unique valid adjacent mismatches (UVAM) was calculated for each candidate polymorphic site and a UVAM threshold was determined by comparing the score distribution of the detected candidate sites to sites that were found in the dbSNP database and the 1000 genomes project (Figures 2a–d). Sites that had a lower quality score than the threshold was considered false positives and excluded. The filtration step was validated by genoty** 48 candidate SNPs before filtration with a 48-plex GoldenGate assay from Illumina (San Diego, CA, USA).28
To remove false-positive SNPs, the distribution of unique valid adjacent miss-matches (UVAM) from the SOLiD sequencing were compared for known SNPs from the dbSNP database (a) and results from the 1000 genomes project (b) with the candidate SNPs from the sequencing of obese (c) and lean group (d). The clear difference in distribution between previously known and novel candidate SNPs indicates that most of the candidates are false positives. Based on this, a binomial hypergeometric distribution was used to model the probability of non-SNPs to have a certain UVAM score in the obese (e) and lean group (f). This allowed us to set a threshold for each group (marked in red) that corresponds to a false discovery rate of 1%. The color reproduction of this figure is available at the International Journal of Obesity online.
Indels with an estimated minor allele frequency >1% were detected using two methods for gapped read map** and included for further analysis.
Data analysis
Association tests based on allele frequencies were performed using Fisher's exact test together with approximation of 95% confidence interval off odds ratios for each SNP and indel using the software R (http://www.r-project.org, R Foundation for statistical computing, Vienna, Austria). To adjust for multiple testing, the significance level was adjusted for the 44 haplotype blocks that we identified across the sequenced region in a previous study of a Swedish population, which contains SNPs in linkage disequilibrium.29 Consequently, P-values below 1.14 × 10−3 were considered significant.
Results from the individual genoty** (GoldenGate and TaqMan assay) were analyzed with PLINK (http://pngu.mgh.harvard.edu/purcell/plink/). The validated SNPs were checked for deviation from Hardy–Weinberg equilibrium using a Pearson's χ2-test (1 d.f) before being tested for association with obesity by using logarithmic regression to calculate 95% confidence intervals of odds ratios, controlled for sex and assuming an additive model. For details on the information with ENCODE data see Supplementary Information 1.
Genoty**
For details on genoty** of candidate SNPs and validation in Latvian and Greek populations see Supplementary Information 1.