Abstract
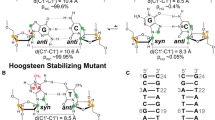
Nucleic acid duplexes featuring a single alpha-anomeric thymidine inserted into each DNA strand via 3′-3′ and 5′-5′ phosphodiester linkages exhibit local conformational dynamics that are not adequately depicted by conventional restrained molecular dynamics (rMD) methods. We have used molecular dynamics with time-averaged NMR restraints (MDtar) to explore its applicability to describing the conformational dynamics of two α-containing duplexes – d(GCGAAT-3′-3′-αT-5′-5′-CGC)2 and d(ATGG-3′-3′-αT-5′-5′-GCTC)•r(gagcaccau). In contrast to rMD, enforcing NOE-based distance restraints over a period of time in MDtar rather than instantaneously results in better agreement with the experimental NOE and J-data. This conclusion is based on the dramatic decreases in average distance and coupling constant violations (Δd av, J rms, and ΔJ av) and improvements in sixth-root R-factors (R x). In both duplexes, the deoxyribose ring puckering behavior predicted independently by pseudorotation analysis is portrayed remarkably well using this approach compared to rMD. This indicates that the local dynamic behavior is encoded within the NOE data, although this is not obvious from the local R x values. In both systems, the backbone torsion angles comprising the 3′-3′ linkage as well as the (high S-) sugars of the α-nucleotide and preceding residue (α−1) are relatively static, while the conformations of the 5′-5′ linkage and the sugar in the neighboring β-nucleotide (α+1) show enhanced flexibility. To reduce the large ensembles generated by MDtar to more manageable clusters we utilized the PDQPRO program. The resulting PDQPRO clusters (in both cases, 13 structures and associated probabilities extracted from a pool of 300 structures) adequately represent the structural and dynamic characteristics predicted by the experimental data.
Similar content being viewed by others
References
Altona, C., Francke, R., de Haan, R., Ippel, J.H., Daalmans, G.J., Westra Hoekzema, A.J.A. and van Wijk, J. (1994) Magn. Reson. Chem., 32, 670–678.
Aramini, J.M. and Germann, M.W. (1999) Biochemistry, 38, 15448–15458.
Aramini, J.M., Kalisch, B.W., Pon, R.T., van de Sande, J.H. and Germann, M.W. (1996) Biochemistry, 35, 9355–9365.
Aramini, J.M., Mujeeb, A. and Germann, M.W. (1998a) Nucleic Acids Res., 26, 5644–5654.
Aramini, J.M., van de Sande, J.H. and Germann, M.W. (1998b) In ACS Symposium Series 682: Molecular Modeling of Nucleic Acids (Eds., Leontis, N.B. and SantaLucia J., Jr.), American Chemical Society, Washington, DC, pp. 92–105.
Bachelin, M., Hessler, G., Kurz, G., Hacia, J.G., Dervan, P.B. and Kessler, H. (1998) Nat. Struct. Biol., 5, 271–276.
Berendsen, H.J.C., Postma, J.P.M., van Gunsteren, W.F., DiNola, A. and Haak, J.R. (1984) J. Chem. Phys., 81, 3684–3690.
Bertrand, J.-R., Imbach, J.-L., Paoletti, C. and Malvy, C. (1989) Biochem. Biophys. Res. Commun., 164, 311–318.
Bhattacharyya, D. and Bansal, M. (1992) J. Biomol. Struct. Dyn., 10, 213–226.
Boiziau, C., Debart, F., Rayner, B., Imbach, J.-L. and Toulmé, J.-J. (1995) FEBS Lett., 361, 41–45.
Bonvin, A.M.J.J. and Brünger, A.T. (1995) J. Mol. Biol., 250, 80–93.
Borgias, B.A. and James, T.L. (1988) J. Magn. Reson., 79, 493–512.
Cheatham III, T.E. and Kollman, P.A. (1997) J. Am. Chem. Soc., 119, 4805–4825.
Darden, T.A., York, D.M. and Pedersen, L.G. (1993) J. Chem. Phys., 98, 10089–10092.
Debart, F., Tosquellas, G., Rayner, B. and Imbach, J.-L. (1994) Bioorg. Med. Chem. Lett., 4, 1041–1046.
Donders, L.A., de Leeuw, F.A.A.M. and Altona, C. (1989) Magn. Reson. Chem., 27, 556–563.
Fedoroff, O.Y., Salazar, M. and Reid, B.R. (1993) J. Mol. Biol., 233, 509–523.
Fennen, J., Torda, A.E. and van Gunsteren, W.F. (1995) J. Biomol. NMR, 6, 163–170.
Ferrin, T.E., Huang, C.C., Jarvis, L.E. and Langridge, R. (1988) J. Mol. Graphics, 6, 13–27.
González, C., Stec, W., Reynolds, M.A. and James, T.L. (1995) Biochemistry, 34, 4969–4982.
Gorin, A.A., Zhurkin, V.B. and Olson, W.K. (1995) J. Mol. Biol., 247, 34–48.
Görler, A., Ulyanov, N.B. and James, T.L. (2000) J. Biomol. NMR, 16, 147–164.
Güntert, P. (1998) Q. Rev. Biophys., 31, 145–237.
Gyi, J.I., Lane, A.N., Conn, G.L. and Brown, T. (1998) Biochemistry, 37, 73–80.
Keepers, J.W. and James, T.L. (1984) J. Magn. Reson., 57, 404–426.
Kemmink, J. and Scheek, R.M. (1995) J. Biomol. NMR, 6, 33–40.
Koga, M., Wilk, A., Moore, M.F., Scremin, C.L., Zhou, L. and Beaucage, S.L. (1995) J. Org. Chem., 60, 1520–1530.
Lavignon, M., Tounekti, N., Rayner, B., Imbach, J.-L., Keith, G., Paoletti, J. and Malvy, C. (1992) Antisense Res. Dev., 2, 315–324.
Liu, H., Spielmann, H.P., Ulyanov, N.B., Wemmer, D.E. and James, T.L. (1995) J. Biomol. NMR, 6, 390–402.
Mujeeb, A., Kerwin, S.M., Kenyon, G.L. and James, T.L. (1993) Biochemistry, 32, 13419–13431.
Nanzer, A.P., Torda, A.E., Bisang, C., Weber, C., Robinson, J.A. and van Gunsteren, W.F. (1997) J. Mol. Biol., 267, 1012–1025.
Pearlman, D.A. and Kollman, P.A. (1991) J. Biol. Mol., 220, 457–479.
Pearlman, D.A., Case, D.A., Caldwell, J.W., Ross, W.S., Cheatham III, T.E., Ferguson, D.M., Seibel, G.L., Singh, U.C., Weiner, P.K. and Kollman, P.A. (1995) Amber 4.1, University of California, San Francisco, CA.
Pikkemaat, J.A. and Altona, C. (1996) Magn. Reson. Chem., 34, S33-S39.
Ravishanker, G., Swaminathan, S., Beveridge, D.L., Lavery, R. and Sklenar, H. (1989) J. Biomol. Struct. Dyn., 6, 669–699.
Ryckaert, J.P., Ciccotti, G. and Berendsen, H.J.C. (1977) J. Comput. Phys., 23, 327–341.
Schmitz, U. and James, T.L. (1995) Methods Enzymol., 261, 3–44.
Schmitz, U., Ulyanov, N.B., Kumar, A. and James, T.L. (1993) J. Biol. Mol., 234, 373–389.
Schmitz, U., Donati, A., James, T.L., Ulyanov, N.B. and Yao, L. (1998) Biopolymers, 46, 329–342.
Scott, W.R.P., Mark, A.E. and van Gunsteren, W.F. (1998) J. Biomol. NMR, 12, 501–508.
Simmerling, C., Miller, J.L. and Kollman, P.A. (1998) J. Am. Chem. Soc., 120, 7149–7155.
Slamon, D.J., Clark, G.M., Wong, S.G., Levin, W.J., Ullrich, A. and McGuire, W.L. (1987) Science, 235, 177–182.
Slamon, D.J., Godolphin, W., Jones, L.A., Holt, J.A., Wong, S.G., Keith, D.E., Levin, W.J., Stuart, S.G., Udove, J., Ullrich, A. and Press, M.F. (1989) Science, 244, 707–712.
Szyperski, T., Götte, M., Billeter, M., Perola, E., Cellai, L., Heumann, H. and Wüthrich, K. (1999) J. Biomol. NMR, 13, 343–355.
Tan, T.M.C., Kalisch, B.W., van de Sande, J.H., Ting, R.C.Y. and Tan, Y.H. (1998) Antisense Nucleic Acid Drug Dev., 8, 95–101.
Thomas, P.D., Basus, V.J. and James, T.L. (1991) Proc. Natl. Acad. Sci. USA, 88, 1237–1241.
Torda, A.E., Scheek, R.M. and van Gunsteren, W.F. (1990) J. Biol. Mol., 214, 223–235.
Ulyanov, N.B. and Zhurkin, V.B. (1984) J. Biomol. Struct. Dyn., 2, 361–385.
Ulyanov, N.B. and James, T.L. (1995) Methods Enzymol., 261, 90–120.
Ulyanov, N.B., Schmitz, U., Kumar, A. and James, T.L. (1995) Biophys. J., 68, 13–24.
van Wijk, J., Huckriede, B.D., Ippel, J.H. and Altona, C. (1992) Methods Enzymol., 211, 286–306.
van Wijk, J., Haasnoot, K., de Leeuw, F., Huckriede, D. and Altona, C. (1995) PSEUROT 6.2. A Program for the Conformational Analysis of Five Membered Rings. University of Leiden, The Netherlands.
Vichier-Guerre, S., Pompon, A., Lefebvre, I. and Imbach, J.-L. (1994) Antisense Res. Dev., 4, 9–18.
Yao, L., James, T.L., Kealey, J.T., Santi, D.V. and Schmitz, U. (1997) J. Biomol. NMR, 9, 229–244. 302
Author information
Authors and Affiliations
Electronic Supplementary Material
Rights and permissions
About this article
Cite this article
Aramini, J.M., Mujeeb, A., Ulyanov, N.B. et al. Conformational dynamics in mixed α/β-oligonucleotides containing polarity reversals: A molecular dynamics study using time-averaged restraints*. J Biomol NMR 18, 287–302 (2000). https://doi.org/10.1023/A:1026798010342
Issue Date:
DOI: https://doi.org/10.1023/A:1026798010342