Abstract
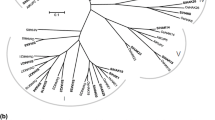
Potassium ions are widely involved in a series of physiological and biochemical processes of plants, which is of great significance to plant growth. High-affinity K+ (HAK) transporter mainly absorbed and transported potassium in plants, but there are few studies on HAK gene in sugarcane. In this study, the coding region of high-affinity K+ transporter genes were cloned from sugarcane and designated as ScHAK9, ScHAK10, ScHAK11 (GenBank accession number: MG564720, MG564721, MG564722). Phylogenetic analysis results confirmed that ScHAK9 and ScHAK11 had the closest relationship with ZmHAK9 and ZmHAK11(Zea mays), ScHAK10 had the closest relationship with SbHAK10 (Sorghum bicolor). In subcellular localization experiments, the fusion protein of ScHAK9 and ScHAK11 with green fluorescent protein was specifically localized in the cell membrane, but ScHAK10 green fluorescent protein was not detected, it was speculated to be expressed in Golgi apparatus. The gene expression level of ScHAK in different tissues of sugarcane at the growth periods was different, and the gene expression level of ScHAK genes were up-regulated by the low-potassium and salt stress. Through the functional characterization experiments of ScHAK genes in K+ uptake-deficient yeasts, it was founded that ScHAK genes possessed K+ transporter activity. The study indicated that ScHAK genes might mediate K+ absorption through the cell membrane and might be participate in maintaining Na+/K+ homeostasis in sugarcane under the adversity stress, and the development of plant organs is regulated by the potassium ions transport of ScHAK genes.





Similar content being viewed by others
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Abbreviations
- HAK :
-
High-affinity K+ transporter genes
- GFP:
-
Green fluorescence protein
- qRT-PCR:
-
Quantitative real-time PCR
- CaMV:
-
Cauliflower mosaic virus
- URA3:
-
Orotidine-5'-phosphate decarboxylase
- GAL1:
-
Recombinant galectin 1
- CYC1:
-
Cytochrome c-1
- AP:
-
Arginine and phosphoric acid
References
Amtmann A, Hammond JP, Armengaud P, White PJ (2005) Nutrient sensing and signalling in plants: potassium and phosphorus. Adv Bot Res 43:209–257. https://doi.org/10.1016/S0065-2296(05)43005-0
Anschutz U, Becker D, Shabala S (2014) Going beyond nutrition: regulation of potassium homoeostasis as a common denominator of plant adaptive responses to environment. J Plant Physiol 171(9):670–687. https://doi.org/10.1016/j.jplph.2014.01.009
Banuelos MA, Garciadeblas B, Cubero B, Navarro AR (2002) Inventory and functional characterization of the HAK potassium transporters of rice. Plant Physiol 130(2):784–795. https://doi.org/10.1104/pp.007781
Boscari A, Clément M, Volkov V, Golldack D, Hybiak J, Miller AJ, Amtmann A, Fricke W (2009) Potassium channels in barley: cloning, functional characterization and expression analyses in relation to leaf growth and development. Plant Cell Environ 32(12):1761–1777. https://doi.org/10.1111/j.1365-3040.2009.02033.x
Chao M, Zhang Z, Song H, Li C, Zhang X, Hu G, Zhang J, Wang Q (2017) Genome-wide identification and expression analysis of Pht1 family genes in cotton (Gossypium hirsutum L.). Cotton Sci 29(1):59–69. https://doi.org/10.11963/issn.1002-7807.201701007
Chen G, Hu Q, Luo LE, Yang T, Zhang S, Hu Y, Yu L, Xu G (2015) Rice potassium transporter OsHAK1 is essential for maintaining potassium-mediated growth and functions in salt tolerance over low and high potassium concentration ranges. Plant Cell Environ 38(12):2747–2765. https://doi.org/10.1111/pce.12585
Feng XM, Wang YJ, Zhang NN, Wu ZL, Zeng QY, Wu JY, Wu XB, Wang L, Zhang JS, Qi YW (2020) Genome-wide systematic characterization of the HAK/KUP/KT gene family and its expression profile during plant growth and in response to low-K+ stress in Saccharum. BMC Plant Biol 20(1):1–17. https://doi.org/10.1186/s12870-019-2227-7
Gierth M, Maser P, Schroeder JI (2005) The potassium transporter AtHAK5 functions in K+ deprivation-induced high-affinity K+ uptake and AKT1 K+ channel contribution to K+ uptake kinetics in Arabidopsis roots. Plant Physiol 137(3):1105–1114. https://doi.org/10.1104/pp.104.057216
He C, Cui K, Duan A, Zeng Y, Zhang J (2012) Genome-wide and molecular evolution analysis of the Poplar KT/HAK/KUP potassium transporter gene family. Ecol & Evol 2(8):1996–2004. https://doi.org/10.1002/ece3.299
Horie T, Sugawara M, Okada T, Taira K, Kaothien-Nakayama P, Katsuhara M, Shinmyo A, Nakayama H (2011) Rice sodium-insensitive potassium transporter, OsHAK5, confers increased salt tolerance in tobacco BY2 cells. J Biosci Bioeng 111(3):346–356. https://doi.org/10.1016/j.jbiosc.2010.10.014
Huang Y, Zeng QY, Ao JH, Chen DW, Zhou WL, Lu YL, Jiang Y, Huang ZR, Li QW (2013) Effects of different potassium levels on nutrient absorption and utilization of N, P K in sugarcane. Guangdong Agric Sci 40(10):50–53. https://doi.org/10.3969/j.issn.1004-874X.2013.10.016
Li J, Long Y, Qi GN, Li J, Xu ZJ, Wu WH, Wang Y (2014) The Os-AKT1channel is critical for K+ uptake in rice roots and is modulated by the rice CBL1-CIPK23 complex. Plant Cell 26(8):3387–3402. https://doi.org/10.1105/tpc.114.123455
Liu JF, Zhang SL, Tang HL, Wu LZ, Dong LJ, Liu LD, Che WL (2015) Overexpression of an Aeluropus littoralis Parl. potassium transporter gene, AlHAK1, in cotton enhances potassium uptake and salt tolerance. Euphytica 203:197–209. https://doi.org/10.1007/s10681-014-1310-2
Lu LM, Yang TZ (2011) Cloning and expression profile analysis of a putative potassium transporter gene NtHAK1 in tobacco. J Nucl Agric Sci 25(03):469–476
Ma TL, Wu WH, Wang Y (2012) Transcriptome analysis of rice root responses to potassium deficiency. BMC Plant Biol 12:161–174. https://doi.org/10.1186/1471-2229-12-161
Maathuis FJM (2006) The role of monovalent cation transporters in plant responses to salinity. J Exp Bot 57(5):1137–1147. https://doi.org/10.1093/jxb/erj001
Maser P, Thomine S, Schroeder JI, Ward JM, Hirschi K, Sze H, Talke IN, Amtmann A, Maathuis FJ, Sanders D, Harper JF, Tchieu J, Gribskov M, Persans MW, Salt DE, Kim SA, Guerinot ML (2001) Phylogenetic relationships within cation transporter families of Arabidopsis. Plant Physiol 126(4):1646–1667. https://doi.org/10.1104/pp.126.4.1646
Nieves-Cordones M, Aleman F, Martinez V, Rubio F (2010) The Arabidopsis thaliana HAK5 K+ transporter is required for plant growth and K+ acquisition from low K+ solutions under saline conditions. Mol Plant 3(2):326–333. https://doi.org/10.1093/mp/ssp102
Rubio F, Santa-Maria GE, Rodriguez-Navarro A (2000) Cloning of Arabidopsis and barley cDNAs encoding HAK potassium transporters in root and shoot cells. Physiol Plantarum 109(1):34–43. https://doi.org/10.1034/j.1399-3054.2000.100106.x
Santa-Maria GE, Rubio F, Dubcovsky J, Rodriguez-Navarro A (1997) The HAK1 gene of barley is a member of a large gene family and encodes a high-affinity potassium transporter. Plant Cell 9(12):2281–2289. https://doi.org/10.2307/3870585
Sato Y, Nanatani K, Hamamoto S, Shimizu M, Takahashi M, Tabuchi-Kobayashi M, Mizutani A, Schroeder JI, Souma S, Uozumi N (2014) Defining membrane spanning domains and crucial membrane-localized acidic amino acid residues for K+ transport of a KUP/HAK/KT-type Escherichia coli potassium transporter. Biochem J 155(5):315–323. https://doi.org/10.1093/jb/mvu007
Song YF, Zhang L, Dong LH, ** YR, Shi SJ, Bai Y, Liu CK, Feng GL, Feng XG, Wang Q, Liu HB (2013) Research progress on KUP /HAK /KT potassium transporter family in plant. J Agric Sci Technol 15(06):92–98. https://doi.org/10.3969/j.issn.1008⁃0864.2013.0
Song YF, Liu HB, Dong LH, ** YR, Shi SJ, Zhang L, Liu CK, Feng XG, Hu XM, Wang Q (2014) Subcellular localization and expression analysis of Nicotiana sylvestris KUP/HAK/KT family K+ transporter gene NsHAK11. Sci Agric Sin 47(6):1058–1071. https://doi.org/10.3864/j.issn.0578-1752.2014.06.003
Su H, Golldack D, Zhao C, Bohnert HJ (2002) The expression of HAK-type K+ transporters is regulated in response to salinity stress in common ice plant. Plant Physiol 129(4):1482–1493. https://doi.org/10.1104/pp.001149
Takahashi R, Nishio T, Ichizen N, Takano T (2007) High-affinity K+ transporter PhaHAK5 is expressed only in salt-sensitive reed plants and shows Na+ permeability under NaCl stress. Plant Cell Rep 26(9):1673–1679. https://doi.org/10.1007/s00299-007-0364-1
Yang Z, Gao Q, Sun C, Li W, Gu S, Xu C (2009) Molecular evolution and functional divergence of HAK potassium transporter gene family in rice (Oryza sativa L.). J Genet Genom 36(3):161–172. https://doi.org/10.1016/S1673-8527(08)60103-4
Yang TY, Zhang S, Hu YB, Wu FC, Hu QD, Chen G, Cai J, Wu T, Moran N, Yu L, Xu GH (2014) The role of a potassium transporter OsHAK5 in potassium acquisition and transport from roots to shoots in rice at low potassium supply levels. Plant Physiol 166(2):945–959. https://doi.org/10.1104/pp.114.246520
Zhang Z, Zhang J, Chen Y, Li R, Wang H, Wei J (2012) Genome-wide analysis and identification of HAK potassium transporter gene family in maize (Zea mays. L). Mol Biol Rep 39(8):8465–8473. https://doi.org/10.1007/s11033-012-1700-2
Zou N, Li B, Dong G, Kronzucker HJ, Shi W (2012) Ammonium-induced loss of root gravitropism is related to auxin distribution and TRH1 function, and is uncoupled from the inhibition of root elongation in Arabidopsis. J Exp Bot 63(10):3777–3788. https://doi.org/10.1093/jxb/ers068
Funding
This work was supported by Natural Science Foundation of Guangxi Province (2018GXNSFBA281103, 2020GXNSFAA259061).
Author information
Authors and Affiliations
Contributions
HL, CH, and YW designed the research. HL, HZ, HC, SJ, and LX performed the experiments and data analysis. HL wrote the manuscript. ZD, KW and YW contributed with valuable discussions. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Consent for publication
Not applicable.
Ethics approval
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Luo, Hb., Huang, Cm., Wei, Yw. et al. Uncovering expression and functional analysis of newly discovered high-affinity K+ transporter family members from sugarcane. J. Plant Biochem. Biotechnol. 31, 826–836 (2022). https://doi.org/10.1007/s13562-021-00726-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13562-021-00726-5