Abstract
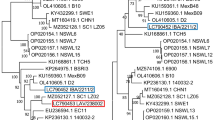
Bovine respiratory syncytial virus (BRSV) is an important viral agent in bovine respiratory disease complex affecting young calves from asymptomatic to fatal. Although BRSV is widely prevalent in Türkiye as in other parts of the world, there are limited molecular studies on BRSV in Türkiye. Therefore, in order to better understand the characteristics of circulating BRSV in Türkiye, a study based on the molecular analysis of both F and G proteins was performed. For this purpose, the presence of BRSV was investigated in 20 calves that died as a result of severe respiratory syndrome in the western region of Türkiye in 2020. Nested PCR was performed for both gene regions, and the products were sequenced. Four samples detected as BRSV positive were identified as genotype III according to both gene regions in molecular analysis. However, they were separated into two distinct clusters due to significant differences in nucleotide (90.09–99.54%) and amino acid (85.42–99.31%) similarities between them. Besides, two positive samples in the same cluster were even more different from previously detected Turkish isolates (90.78–92.17% nt and 87.50–89.58% aa). More over, we detected nine novel aa mutations in the extracellular domain, an immunologically important region in the G protein of the virus, that have not been reported in other world isolates found in Genbank until now. These findings suggest that there may be many different viruses in circulation that have the ability to escape the immune system. We recommend that these findings be taken into account in planning both vaccine and epidemiological studies.


Similar content being viewed by others
Data availability
All data generated or analysed during this study are included in this published article. The datasets generated for this study can be found in NCBI, MW892048, MW892049, MW892050 and MZ476914 for F gene; MW892044, MW892045, MW892046 and MW892047 for G gene of the detected BRSV.
References
Alkan F, Ozkul A, Bilge Dagalp S, Yesilbag K, Oguzoglu TC, Akça Y, Burgu I. Virological and serological studies on the role of PI-3 virus, BRSV, BVDV and BHV-1 on respiratory infections of cattle I The detection of etiological agents by direct immunofluorescence technique. Dtsch Tierarztl Wochenschr. 2000;107(5):193–5.
Alkan F, Özkul A, Karaoglu MT, Bilge S, Akça Y, Burgu I, Yesilbag K, Oğuzoğlu TC. Sığırlarda viral nedenli solunum sistemi enfeksiyonlarinin seroepidemiyolojisi. Ankara Üniv Vet Fak Derg. 1997;44:73–80. https://doi.org/10.1501/Vetfak_0000000658.
Alpay G, Tuncer P, Yeşilbağ K. Bir ada ekosistemindeki sığır, koyun ve keçilerde bazı viral enfeksiyonların serolojik olarak araştırılması. Ankara Üniv Vet Fak Derg. 2014;61:43–8. https://doi.org/10.1501/Vetfak_0000002603.
Avci O, Yavru S, Ekik M. Detection of respiratory viral antigens in cattle lung tissues by direct ELISA. Anim Vet Sci. 2014;2:146–9.
Avci O, Yavru S, Sevik M. Antibody prevalence against respiratory viruses in naturally infected cattle in Central Anatolia. Eurasian J Vet Sci. 2014;30(2):80–4.
Baptista AL, Rezende AL, Fonseca PDA, Massi RP, Nogueira GM, Magalhães LQ, Headley SA, Menezes GL, Alfieri AA, Saut JPE. Bovine respiratory disease complex associated mortality and morbidity rates in feedlot cattle from southeastern Brazil. J Infect Dev Ctries. 2017;11:791–9. https://doi.org/10.3855/jidc.9296.
Bertolotti L, Giammarioli M, Rosati S. Genetic characterization of bovine respiratory syncytial virus strains isolated in Italy: evidence for the circulation of new divergent clades. J Vet Diagn Investig. 2018;30:300–4. https://doi.org/10.1177/1040638717746202.
Beuttemmuller EA, Alfieri AF, Headley SA, Alfieri AA. Brazilian strain of bovine respiratory coronavirus is derived from dual enteric and respiratory tropism. Genet Mol Res. 2017;16(2):gmr16029580. https://doi.org/10.4238/gmr16029580.
Burgu I, Toker A, Akca Y, Alkan F. Seroepidemiologic study of bovine respiratory syncytial virus (BRSV) in Turkey. Dtsch Tierarztl Wochenschr. 1990;97(2):88–9.
Çabalar M, Can Şahna K. Doğu ve güneydoğu anadolu bölgesinde süt sığırlarında parainfluenza virus-3, bovine herpes virus-1 ve respiratory syncytial virus enfeksiyonlarının seroepidemiyolojisi. YYÜ Vet Fak Derg. 2000;11:101–5.
Chang Y, Yue H, Tang C. Prevalence and molecular characteristics of bovine respiratory syncytial virus in beef cattle in China. Animals. 2022;12:3511. https://doi.org/10.3390/ani12243511.
Duman R, Yavru S, Kale M, Avcı O. Seroprevalence of viral upper respiratory infections in dairy cattle. Kafkas Univ Vet Fak Derg. 2009;15(4):539–42.
Fulton RW. Bovine respiratory disease research (1983–2009). Anim Health Res Rev. 2009;10:131–9. https://doi.org/10.1017/S146625230999017X.
Furze J, Roberts SR, Wertz G, Taylor G. Antigenically distinct G glycoproteins of BRSV strains share a high degree of genetic homogeneity. Virology. 1997;231:48–58. https://doi.org/10.1006/viro.1997.8490.
Furze J, Wertz G, Lerch Taylor G. Antigenic heterogenicity of the attachment protein of bovine respiratory syncytial virus. J Gen Virol. 1994;75:363–70. https://doi.org/10.1099/0022-1317-75-2-363.
Gershwin LJ, Van Eenennaam AL, Tisioto PC, Kim J, Seabury CM, Taylor JF, Neibergs H L. The host gene expression response to specific BRD pathogens. In: American Association of bovine practitioners conference proceedings; 2015. p. 157–60.
Hall T. BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser. 1999;41:95–8.
Headley SA, Okano W, Balbo LC, Marcasso RA, Oliveira TE, Alfieri AF, NegriFilho LC, Michelazzo MZ, Rodrigues SC, Baptista AL, Saut SJP, Alfieri AA. Molecular survey of infectious agents associated with bovine respiratory disease in a beef cattle feedlot in southern Brazil. J Vet Diagn Invest. 2018;30:249–51. https://doi.org/10.1177/1040638717739945.
International Committee on Taxonomy of Viruses. 2019. https://talk.ictvonline.org/ictv-reports/ictv_online_report/negative-sense-rnaviruses/mononegavirales/w/pneumoviridae/738/genus-orthopneumovirus. Accessed March 2022.
Ince OM, Sevik M, Ozgur EG, Sait A. Risk factors and genetic characterization of bovine respiratory syncytial virus in the inner Aegean Region, Turkey. Trop Anim Health Prod. 2022;54(1):4. https://doi.org/10.1007/s11250-021-03022-5.
Karayel Hacioglu I, Coskun N, Duran Yelken S, Sevinc S, Alkan F. Phylogenetic analysis of bovine respiratory syncytial virus form calves with respiratory disorders. Kafkas Univ Vet Fak Derg. 2019;25(2):251–9. https://doi.org/10.9775/kvfd.2018.20819.
Klem TB, Kjæstad HP, Kummen E, Holen H, Stokstad M. Bovine respiratory syncytial virus outbreak reduced bulls’ weight gain and feed conversion for eight months in a Norwegian beef herd. Acta Vet Scand. 2016;58:8. https://doi.org/10.1186/s13028-016-0190-y.
Klem TB, Rimstad E, Stokstad M. Occurrence and phylogenetic analysis of bovine respiratory syncytial virus in outbreaks of respiratory disease in Norway. BMC Vet Res. 2014;10:1–9. https://doi.org/10.1186/1746-6148-10-15.
Kresic N, Bedekovic T, Brnic D, Simic I, Lojkic I, Turk N. Genetic analysis of bovine respiratory syncytial virus in Croatia. Comp Immunol Microbiol Infect Dis. 2018;58:52–7. https://doi.org/10.1016/j.cimid.2018.09.004.
Kumagai A, Kawauchi K, Andoh K, Hatama S. Sequence and unique phylogeny of G genes of bovine respiratory syncytial viruses circulating in Japan. J Vet Diagn Investig. 2021;33(1):162–6. https://doi.org/10.1177/1040638720975364.
Kumar S, Stecher G, Li M, Knyaz C, Tamura K. MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol. 2018;35(6):1547–9. https://doi.org/10.1093/molbev/msy096.
Langedijk JP, Meloen RH, Taylor G, Furze JM, van Oirschot JT. Antigenic structure of the central conserved region of protein G of bovine respiratory syncytial virus. J Virol. 1997;71:4055–61. https://doi.org/10.1128/jvi.71.5.4055-4061.1997.
Larsen LE, Tjørnehøj K, Viuff B. Extensive sequence divergence among bovine respiratory syncytial viruses isolated during recurrent outbreaks in closed herds. J Clin Microbiol. 2000;38:4222–7. https://doi.org/10.1128/JCM.38.11.4222-4227.2000.
Leme RA, Agnol AMD, Balbo LC, Pereria FL, Possatti F, Alfieri AF, Alfieri AA. Molecular characterization of Brazilian wild-type strains of Bovine respiratory syncytial virus reveals genetic diversity and putative subgroup of the virus. Vet Q. 2020;40:83–96. https://doi.org/10.1080/01652176.2020.1733704.
Lerch RA, Anderson K, Wertz GW. Nucleotide suquence analysis and expression from recombinant vectors demonstrate that the attachment protein G of bovine respiratory syncytial virus is distinct from that of human respiratory syncytial virus. J Virol. 1990;64:5559–69. https://doi.org/10.1128/jvi.64.11.5559-5569.1990.
Neill JD, Ridpath JF, Valayudhan BT. Identification and genome characterization of genotype B and genotype C of bovine parainfluenza type 3 viruses isolated in the United States. BMC Vet Res. 2015;11:112. https://doi.org/10.1186/s12917-015-0431-8.
Prozzi D, Walravens K, Langedijk JPM, Daus F, Kramps JA, Letesson JJ. Antigenic and molecular analyses of the variability of bovine respiratory syncytial virus G glycoprotein. J Gen Virol. 1997;78(2):359–66. https://doi.org/10.1099/0022-1317-78-2-359.
Ronquist F, Huelsenbeck JP. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics. 2003;19(12):1572–4. https://doi.org/10.1093/bioinformatics/btg180.
Sarmiento-Silva RE, Nakamura-Lopez Y, Vaughan G. Epidemiology, molecular epidemiology, and evolution of bovine respiratory syncytial virus. Viruses. 2012;4:3452–67. https://doi.org/10.3390/v4123452.
Schrijver RS, Daus F, Kramps JA, Langedijk JPM, Buijs R, Middel WGJ, Taylor G, Furze J, Huyben MWC, van Oirschot JT. Subgrou** of bovine respiratory syncytial virus strains detected in lung tissue. Vet Microbiol. 1996;53(3–4):253–60. https://doi.org/10.1016/S0378-1135(96)01223-0.
Socha W, Larska M, Rola J. Molecular characterisation of the first polish isolates of bovine respiratory syncytial virus. Bull Vet Inst Pulawy. 2009;53:569–74.
Taylor G, Thomas LH, Furze JM, Cook RS, Wyld SG, Lerch R, Hardy R, Wertz GW. Recombinant vaccinia viruses expressing the F, G or N, but not the M2, protein of bovine respiratory syncytial virus (BRSV) induce resistance to BRSV challenge in the calf and protect aganist the development of pneumonic lesions. J Gen Virol. 1997;44:191–5. https://doi.org/10.1099/0022-1317-78-12-3195.
Taylor G, Wyld S, Valarcher JF, Guzman E, Thom M, Widdison S, Buchholz UJ. Recombinant bovine respiratory syncytial virus with deletion of the SH gene induces increased apoptosis and pro-inflammatory cytokines in vitro, and is attenuated and induces protective immunity in calves. J Gen Virol. 2014;95:1244–54. https://doi.org/10.1099/vir.0.064931-0.
Thonur L, Maley M, Gilray J, Crook T, Laming E, Turnbull D, Nath M, Willoughby K. One-step multiplex real time RT-PCR for the detection of bovine respiratory syncytial virus, bovine herpesvirus 1 and bovine parainfluenza virus 3. BMC Vet Res. 2012;8:37. https://doi.org/10.1186/1746-6148-8-37.
Timurkan MO, Aydin H, Sait A. Identification and molecular characterisation of bovine parainfluenza virus-3 and bovine respiratory syncytial virus-first report form Turkey. J Vet Res. 2019;63:167–73. https://doi.org/10.2478/jvetres-2019-0022.
Toker EB, Yeşilbağ K. Molecular characterization and comprarison of diagnostic methods for bovine respiratory viruses (BPIV-3, BRSV, BVDV, and BoHV-1) in field samples in Northwestern Turkey. Trop Anim Health Prod. 2021;53:79. https://doi.org/10.1007/s11250-020-02489-y.
Urban-Chmiel R, Wernicki A, Puchalski A, Dec M, Stęgierska D, Grooms DL, Barbu NI. Detection of bovine respiratory syncytial virus infections in young dairy and beef cattle in Poland. Vet Q. 2014;35:33–6. https://doi.org/10.1080/01652176.2014.984366.
Valarcher JF, Schelcher F, Bourhy H. Evolution of bovine respiratory syncytial virus. J Virol. 2000;74:10714–28. https://doi.org/10.1128/JVI.74.22.1071410728.2000.
Valarcher JF, Taylor G. Bovine respiratory syncytial virus infection. Vet Res. 2007;38(2):153–80. https://doi.org/10.1051/vetres:2006053.
Valentova V. The antigenic and genetic variability of bovine respiratory syncytial virus with emphasis on the G protein. Vet Med. 2003;48:254–66. https://doi.org/10.17221/5778-VETMED.
Valentova V, Antonis AFG, Kovarcik K. Restriction enzyme analysis of RT-PCR amplicons as a rapid method for detection of genetic diversity among bovine respiratory syncytial virus isolates. Vet Microbiol. 2005;108:1–12. https://doi.org/10.1016/j.vetmic.2005.02.008.
Yavru S, Şimşek A, Yapkıç O, Kale M. Serological evaluation of viral infections in bovine respiratory tract. Acta Vet-Beograd. 2005;55:219–26. https://doi.org/10.2298/AVB0503219Y.
Yazici Z, Ozan E, Tamer C, Muftuoglu B, Barry G, Kurucay HN, Elhag AE, Cagirgan AA, Gumusova S, Albayrak H. Circulation of indigenous bovine respiratory syncytial virus strains in Turkish Cattle: the first isolation and molecular characterization. Animals. 2020;10:1700. https://doi.org/10.3390/ani10091700.
Yeşilbağ K, Güngör B. Seroprevalence of bovine respiratory viruses in North-Western Turkey. Trop Anim Health Prod. 2008;40(1):55–60. https://doi.org/10.1007/s11250-007-9053-x.
Acknowledgements
This study was supported and officially permissied by Ministry of Agriculture and Forestry of The Republic of Türkiye.
Funding
This study was supported and funded by Agriculture and Foresty of the Republic of Türkiye and İzmir/Bornova Veterinary Control Institute.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflicts of interest
The authors declare that they have no conflict of interest.
Consent for publication
The manuscript was approved and permissioned as officialy by Ministry of Agriculture and Forestry.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kaplan, M., Özan, E., Pekmez, K. et al. Molecular characterization of G and F protein genes of bovine respiratory syncytial virus detected from dead calves caused by severe respiratory syndrome: emergence of novel mutations and their importance. VirusDis. 34, 539–549 (2023). https://doi.org/10.1007/s13337-023-00846-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13337-023-00846-7