Abstract
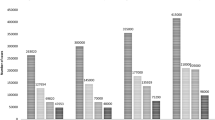
The susceptibility of an individual to oral cancer is mediated by genetic factors and carcinogen-exposure behaviors such as betel quid chewing, tobacco use, and alcohol consumption. This pilot study was aimed to identify the genetic alteration in 100 bp upstream and downstream flanking regions in addition to the exonic regions of 169 cancer-associated genes by using Next Generation sequencing with aim to elucidate the molecular pathogenesis of tobacco- and betel quid-associated oral cancer of Northeast India. To understand the role of chemical compounds present in tobacco and betel quid associated with the progression of oral cancer, single nucleotide polymorphisms (SNPs) and insertion and deletion (Indels) found in this study were analyzed for their association with chemical compounds found in tobacco and betel quid using Comparative Toxogenomic Database. Genes (AR, BRCA1, IL8, and TP53) with novel SNP were found to be associated with arecoline which is the major component of areca nut. Genes (BARD1, BRCA2, CCND2, IGF1R, MSH6, and RASSF1) with novel deletion and genes (APC, BRMS1, CDK2AP1, CDKN2B, GAS1, IGF1R, and RB1) with novel insertion were found to be associated with aflatoxin B1 which is produced by fermented areca nut. Genes (ADH6, APC, AR, BARD1, BRMS1, CDKN1A, E2F1, FGFR4, FLNC, HRAS, IGF1R, IL12B, IL8, NBL1, STAT5B, and TP53) with novel SNP were found to be associated with aflatoxin B1. Genes (ATM, BRCA1, CDKN1A, EGFR, IL8, and TP53) with novel SNP were found to be associated with tobacco specific nitrosamines.

Similar content being viewed by others
Abbreviations
- OSCC:
-
Oral squamous cell carcinoma
- Indels:
-
Short insertion and deletions
- SNP:
-
Single nucleotide polymorphism
- NGS:
-
Next generation sequencing
References
Soya SS, Vinod T, Reddy KS, Gopalakrishnan S, Adithan C. Genetic polymorphisms of glutathione-S-transferase genes (GSTM1, GSTT1 and GSTP1) and upper aerodigestive tract cancer risk among smokers, tobacco chewers and alcoholics in an Indian population. Eur J Cancer. 2007;43:2698–706.
Nair U, Bartsch H, Nair J. Alert for an epidemic of oral cancer due to use of the betel quid substitutes gutkha and pan masala: a review of agents and causative mechanisms. Mutagenesis. 2004;19:251–62.
Bhattacharjee A, Chakraborty A, Purkaystha P. Prevalence of head and neck cancers in the north east—an institutional study. Indian J Otolaryngol Head Neck Surg. 2006;58:15–9.
Mondal P, Datta S, Maiti GP, Baral A, Jha GN, Panda CK, et al. Comprehensive SNP scan of DNA repair and DNA damage response genes reveal multiple susceptibility loci conferring risk to tobacco associated leukoplakia and oral cancer. PLoS One. 2013;8:e56952.
Mondal R, Ghosh SK, Choudhury JH, Seram A, Sinha K, Hussain M, et al. Mitochondrial DNA copy number and risk of oral cancer: a report from Northeast India. PLoS One. 2013;8:e57771.
Chattopadhyay I, Kapur S, Purkayastha J, Phukan R, Kataki A, Mahanta J, et al. Gene expression profile of esophageal cancer in North East India by cDNA microarray analysis. World J Gastroenterol. 2007;13:1438–44.
Phukan RK, Ali MS, Chetia CK, Mahanta J. Betel nut and tobacco chewing; potential risk factors of cancer of oesophagus in Assam. India Br J Cancer. 2001;85:661–7.
Strickland SS, Veena GV, Houghton PJ, Stanford SC, Kurpad AV. Areca nut, energy metabolism and hunger in Asian men. Ann Hum Biol. 2003;30:26–52.
Chiang SL, Jiang SS, Wang YJ, Chiang HC, Chen PH, Tu HP, et al. Characterization of arecoline-induced effects on cytotoxicity in normal human gingival fibroblasts by global gene expression profiling. Toxicol Sci. 2007;100:66–74.
Chien MH, Yang JS, Chu YH, Lin CH, Wei LH, Yang SF, et al. Impacts of CA9 gene polymorphisms and environmental factors on oral-cancer susceptibility and clinicopathologic characteristics in Taiwan. PLoS One. 2012;7:e51051.
Ambatipudi S, Gerstung M, Pandey M, Samant T, Patil A, Kane S, et al. Genome-wide expression and copy number analysis identifies driver genes in gingivobuccal cancers. Genes Chromosom Cancer. 2012;51:161–73.
Ambatipudi S, Gerstung M, Gowda R, Pai P, Borges AM, Schäffer AA, et al. Genomic profiling of advanced-stage oral cancers reveals chromosome 11q alterations as markers of poor clinical outcome. PLoS One. 2011;6:e17250.
Sailasree R, Abhilash A, Sathyan KM, Nalinakumari KR, Thomas S, Kannan S. Differential roles of p16INK4A and p14ARF genes in prognosis of oral carcinoma. Cancer Epidemiol Biomarkers Prev. 2008;17:414–20.
India Project Team of the International Cancer Genome Consortium. Mutational landscape of gingivo-buccal oral squamous cell carcinoma reveals new recurrently-mutated genes and molecular subgroups. Nat Commun. 2013;4:2873.
Zhang Q, Zhang J, ** H, Sheng S. Whole transcriptome sequencing identifies tumor-specific mutations in human oral squamous cell carcinoma. BMC Med Genom. 2013;6:28.
West KA, Brognard J, Clark AS, Linnoila IR, Yang X, Swain SM, et al. Rapid Akt activation by nicotine and a tobacco carcinogen modulates the phenotype of normal human airway epithelial cells. J Clin Invest. 2003;111:81–90.
Chaudhry IS, El-Meanawy A, Khiyami A, Tomashefski JF Jr, Machekano RN, Kass L. Short-term exposure to tobacco toxins alters expression of multiple proliferation gene markers in primary human bronchial epithelial cell cultures. J Oncol. 2011; 208563.
Proulx LI, Gaudreault M, Turmel V, Augusto LA, Castonguay A, Bissonnette EY. 4-(Methylnitrosamino)-1-(3-pyridyl)-1-butanone, a component of tobacco smoke, modulates mediator release from human bronchial and alveolar epithelial cells. Clin Exp Immunol. 2005;140:46–53.
Segal-Raz H, Mass G, Baranes-Bachar K, Lerenthal Y, Wang SY, Chung YM, et al. ATM-mediated phosphorylation of polynucleotide kinase/phosphatase is required for effective DNA double-strand break repair. EMBO Rep. 2011;12:713–9.
Woodbine L, Brunton H, Goodarzi AA, Shibata A, Jeggo PA. Endogenously induced DNA double strand breaks arise in heterochromatic DNA regions and require ataxia telangiectasia mutated and Artemis for their repair. Nucleic Acids Res. 2011;39:6986–97.
Dombernowsky SL, Weischer M, Allin KH, Bojesen SE, Tybjaerg-Hansen A, Nordestgaard BG. Risk of cancer by ATM missense mutations in the general population. J Clin Oncol. 2008;26:3057–62.
Lu S, Shen K, Wang Y, Santner SJ, Chen J, Brooks SC, et al. Atm-haploinsufficiency enhances susceptibility to carcinogen-induced mammary tumors. Carcinogenesis. 2006;27:848–55.
Ogmundsdóttir HM, Hilmarsdóttir H, Björnsson J, Holbrook WP. Longitudinal study of TP53 mutations in eight patients with potentially malignant oral mucosal disorders. Oral Pathol Med. 2009;38:716–21.
Sawada Y, Tamada M, Dubin-Thaler BJ, Cherniavskaya O, Sakai R, Tanaka S, et al. Force sensing by mechanical extension of the Src family kinase substrate p130Cas. Cell. 2006;127:1015–26.
Nishikawa Y, Miyazaki T, Nakashiro K, Yamagata H, Isokane M, Goda H, et al. Human FAT1 cadherin controls cell migration and invasion of oral squamous cell carcinoma through the localization of β-catenin. Oncol Rep. 2011;26:587–92.
Matsui S, Utani A, Takahashi K, Mukoyama Y, Miyachi Y, Matsuyoshi N. Human Fat2 is localized at immature adherens junctions in epidermal keratinocytes. J Dermatol Sci. 2007;48:233–6.
Nakaya K, Yamagata HD, Arita N, Nakashiro KI, Nose M, Miki T, et al. Identification of homozygous deletions of tumor suppressor gene FAT in oral cancer using CGH-array. Oncogene. 2007;26:5300–8.
Fu YP, Edvardsen H, Kaushiva A, Arhancet JP, Howe TM, Kohaar I, et al. NOTCH2 in breast cancer: association of SNP rs11249433 with gene expression in ER-positive breast tumors without TP53 mutations. Mol Cancer. 2010;9:113.
Chu D, Zheng J, Wang W, Zhao Q, Li Y, Li J, et al. Notch2 expression is decreased in colorectal cancer and related to tumor differentiation status. Ann Surg Oncol. 2009;16:3259–66.
Boulay JL, Miserez AR, Zweifel C, Sivasankaran B, Kana V, Ghaffari A, et al. Loss of NOTCH2 positively predicts survival in subgroups of human glial brain tumors. PLoS One. 2007;2:e576.
Lo Muzio L, Campisi G, Farina A, Rubini C, Pannone G, Serpico R, et al. P-cadherin expression and survival rate in oral squamous cell carcinoma: an immunohistochemical study. BMC Cancer. 2005;5:63.
Au KS, Ward CH, Northrup H. Tuberous sclerosis complex: disease modifiers and treatments. Curr Opin Pediatr. 2008;20:628–33.
Inoki K. Role of TSC-mTOR pathway in diabetic nephropathy. Diabetes Res Clin Pract. 2008;82:S59–62.
Adachi H, Igawa M, Shiina H, Urakami S, Shigeno K, Hino O. Human bladder tumors with 2-hit mutations of tumor suppressor gene TSC1 and decreased expression of p27. J Urol. 2003;170:601–4.
Bauer CM, Ray AM, Halstead-Nussloch BA, Dekker RG, Raymond VM, Gruber SB, et al. Hereditary prostate cancer as a feature of Lynch syndrome. Fam Cancer. 2011;10:37–42.
Lim EJ, Leung C, Pitman A, Stella DL, Brown G, Slattery M, et al. Magnetic resonance colonography for colorectal cancer screening in patients with Lynch syndrome gene mutation. Fam Cancer. 2010;9:555–61.
Friedrich RE, Hagel C, Bartel-Friedrich S. Insulin-like growth factor-1 receptor (IGF-1R) in primary and metastatic undifferentiated carcinoma of the head and neck: a possible target of immunotherapy. Anticancer Res. 2010;30:1641–3.
Wang Y, Azuma Y, Moore D, Osheroff N, Neufeld KL. Interaction between tumor suppressor adenomatous polyposis coli and topoisomerase IIalpha: implication for the G2/M transition. Mol Biol Cell. 2008;19:4076–85.
Qian J, Sarnaik AA, Bonney TM, Keirsey J, Combs KA, Steigerwald K, et al. The APC tumor suppressor inhibits DNA replication by directly binding to DNA via its carboxyl terminus. Gastroenterology. 2008;135:152–62.
Tighe A, Johnson VL, Taylor SS. Truncating APC mutations have dominant effects on proliferation, spindle checkpoint control, survival and chromosome stability. J Cell Sci. 2004;117:6339–53.
Rivero ER, Horta MC, Silva Guerra EN, Ferraz AR, Nunes FD. Loss of heterozygosity of the APC gene in oral squamous cell carcinoma. Pathol Res Pract. 2008;204:793–7.
Chien MH, Liu YF, Hsin CH, Lin CH, Shih CH, Yang SF, et al. Impact of VEGF-C gene polymorphisms and environmental factors on oral cancer susceptibility in Taiwan. PLoS One. 2013;8(4):e60283.
Lin CW, Chuang CY, Tang CH, Chang JL, Lee LM, Lee WJ, et al. Combined effects of icam-1 single-nucleotide polymorphisms and environmental carcinogens on oral cancer susceptibility and clinicopathologic development. PLoS One. 2013;8(9):e72940.
Liu CM, Yeh CJ, Yu CC, Chou MY, Lin CH, Wei LH, et al. Impact of interleukin-8 gene polymorphisms and environmental factors on oral cancer susceptibility in Taiwan. Oral Dis. 2012;18:307–14.
Jamaly S, Khanehkenari MR, Rao R, Patil G, Thakur S, Ramaswamy P, et al. Relationship between p53 overexpression, human papillomavirus infection, and lifestyle in Indian patients with head and neck cancers. Tumour Biol. 2012;33:543–50.
Lee CH, Wong TS, Chan JY, Lu SC, Lin P, Cheng AJ, et al. Epigenetic regulation of the X-linked tumour suppressors BEX1 and LDOC1 in oral squamous cell carcinoma. J Pathol. 2013;230:298–309.
Acknowledgments
The authors would like to thank Genotypic Technology (P) Ltd., Bangalore. The authors would also like to thank the Indian Council of Medical Research for their financial support.
Conflicts of interest
None
Author information
Authors and Affiliations
Corresponding author
Additional information
Dhirendra Singh Yadav and Indranil Chattopadhyay have equal contribution to this study.
Rights and permissions
About this article
Cite this article
Yadav, D.S., Chattopadhyay, I., Verma, A. et al. A pilot study evaluating genetic alterations that drive tobacco- and betel quid-associated oral cancer in Northeast India. Tumor Biol. 35, 9317–9330 (2014). https://doi.org/10.1007/s13277-014-2222-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-014-2222-4