Abstract
Backgound
Hereditary hearing loss is one of the most common genetically heterogeneous defects in human. About 70% of hereditary hearing loss is defined as non-syndromic hearing loss showing loss of hearing ability without any other symptoms. Up to date, the identified genes associated with non-syndromic hearing loss are 128, including 52 genes for DFNA and 76 genes for DFNB. Because of high levels of heterogeneity, it is difficult to identify the causative factors for hearing loss using Sanger sequencing.
Objective
Our aim was to detect causative factors and investigate pathogenic mutations, which co-segregates within the candidate family.
Methods
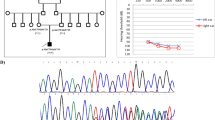
We used Next Generation Sequencing technique to investigate whole-exome sequences of a Korean family with non-syndromic hereditary hearing loss. The family showed autosomal dominant inheritance pattern.
Results
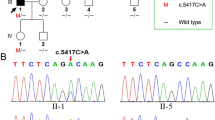
We identified a novel missense variation, c.1978G > A in MYO7A gene, in the family with the autosomal dominant inheritance pattern. c.1978G > A produced Gly660Arg in the motor head domain of Myosin VIIA disrupt the ATP- and actin-binding motif function.
Conclusion
This study is the first to report pathogenic mutations within MYO7A gene in Korean family and our data would facilitate diagnosing the primary cause of hereditary hearing loss in Korean.


Similar content being viewed by others
References
Carrasco-Ramiro F, Peiro-Pastor R, Aguado B (2017) Human genomics projects and precision medicine. Gene Ther 24:551–561
Fettiplace R, Hackney CM (2006) The sensory and motor roles of auditory hair cells. Nat Rev Neurosci 7:19–29
Genomes Project C, Abecasis GR, Auton A, Brooks LD, DePristo MA, Durbin RM, Handsaker RE, Kang HM, Marth GT, McVean GA (2012) An integrated map of genetic variation from 1092 human genomes. Nature 491:56–65
Gibbs D, Diemer T, Khanobdee K, Hu J, Bok D, Williams DS (2010) Function of MYO7A in the human RPE and the validity of shaker1 mice as a model for Usher syndrome 1B. Invest Ophthalmol Vis Sci 51:1130–1135
Gilissen C, Hoischen A, Brunner HG, Veltman JA (2011) Unlocking mendelian disease using exome sequencing. Genome Biol 12:228
Hasson T, Heintzelman MB, Santos-Sacchi J, Corey DP, Mooseker MS (1995) Expression in cochlea and retina of myosin VIIa, the gene product defective in Usher syndrome type 1B. Proc Natl Acad Sci U S A 92:9815–9819
Hasson T, Gillespie PG, Garcia JA, MacDonald RB, Zhao Y, Yee AG, Mooseker MS, Corey DP (1997) Unconventional myosins in inner-ear sensory epithelia. J Cell Biol 137:1287–1307
Inoue A, Ikebe M (2003) Characterization of the motor activity of mammalian myosin VIIA. J Biol Chem 278:5478–5487
Jacobson SG, Aleman TS, Sumaroka A, Cideciyan AV, Roman AJ, Windsor EA, Schwartz SB, Rehm HL, Kimberling WJ (2009) Disease boundaries in the retina of patients with Usher syndrome caused by MYO7A gene mutations. Invest Ophthalmol Vis Sci 50:1886–1894
Lee KY, Choi SY, Bae JW, Kim S, Chung KW, Drayna D, Kim UK, Lee SH (2008) Molecular analysis of the GJB2, GJB6 and SLC26A4 genes in korean deafness patients. Int J Pediatr Otorhinolaryngol 72:1301–1309
Levy G, Levi-Acobas F, Blanchard S, Gerber S, Larget-Piet D, Chenal V, Liu XZ, Newton V, Steel KP, Brown SD et al (1997) Myosin VIIA gene: heterogeneity of the mutations responsible for Usher syndrome type IB. Hum Mol Genet 6:111–116
Li L, Yuan H, Wang H, Guan J, Lan L, Wang D, Zong L, Liu Q, Han B, Huang D et al (2018) Identification of a MYO7A mutation in a large chinese DFNA11 family and genotype-phenotype review for DFNA11. Acta Otolaryngol 138:463–470
Lin X, Tang W, Ahmad S, Lu J, Colby CC, Zhu J, Yu Q (2012) Applications of targeted gene capture and next-generation sequencing technologies in studies of human deafness and other genetic disabilities. Hear Res 288:67–76
Liu XZ (2002) The clinical presentation of DFNB2. Adv Otorhinolaryngol 61:120–123
Luijendijk MW, Van Wijk E, Bischoff AM, Krieger E, Huygen PL, Pennings RJ, Brunner HG, Cremers CW, Cremers FP, Kremer H (2004) Identification and molecular modelling of a mutation in the motor head domain of myosin VIIA in a family with autosomal dominant hearing impairment (DFNA11). Hum Genet 115:149–156
Ma Y, **ao Y, Zhang F, Han Y, Li J, Xu L, Bai X, Wang H (2016) Novel compound heterozygous mutations in MYO7A gene associated with autosomal recessive sensorineural hearing loss in a chinese family. Int J Pediatr Otorhinolaryngol 83:179–185
Morton NE (1991) Genetic epidemiology of hearing impairment. Ann N Y Acad Sci 630:16–31
Morton CC, Nance WE (2006) Newborn hearing screening–a silent revolution. N Engl J Med 354:2151–2164
Rabbani B, Tekin M, Mahdieh N (2014) The promise of whole-exome sequencing in medical genetics. J Hum Genet 59:5–15
Raviv D, Dror AA, Avraham KB (2010) Hearing loss: a common disorder caused by many rare alleles. Ann N Y Acad Sci 1214:168–179
Sang Q, Yan X, Wang H, Feng R, Fei X, Ma D, **ng Q, Li Q, Zhao X, ** L et al (2013) Identification and functional study of a new missense mutation in the motor head domain of myosin VIIA in a family with autosomal dominant hearing impairment (DFNA11). PLoS ONE 8:e55178
Schwarz JM, Cooper DN, Schuelke M, Seelow D (2014) Mutation taster2: mutation prediction for the deep-sequencing age. Nat Methods 11:361–362
Sikkema-Raddatz B, Johansson LF, de Boer EN, Almomani R, Boven LG, van den Berg MP, van Spaendonck-Zwarts KY, van Tintelen JP, Sijmons RH, Jongbloed JD et al (2013) Targeted next-generation sequencing can replace Sanger sequencing in clinical diagnostics. Hum Mutat 34:1035–1042
Sirmaci A, Edwards YJ, Akay H, Tekin M (2012) Challenges in whole exome sequencing: an example from hereditary deafness. PLoS ONE 7:e32000
Smith RJ, Bale JF Jr, White KR (2005) Sensorineural hearing loss in children. Lancet 365:879–890
Tamagawa Y, Kitamura K, Ishida T, Ishikawa K, Tanaka H, Tsuji S, Nishizawa M (1996) A gene for a dominant form of non-syndromic sensorineural deafness (DFNA11) maps within the region containing the DFNB2 recessive deafness gene. Hum Mol Genet 5:849–852
Zina ZB, Masmoudi S, Ayadi H, Chaker F, Ghorbel AM, Drira M, Petit C (2001) From DFNB2 to Usher syndrome: variable expressivity of the same disease. Am J Med Genet 101:181–183
Acknowledgements
We appreciate the families for participating in the genetic study. This research was supported by Kyungpook National University Development Project Research Fund, 2018.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kim, YR., Kim, HM., Lee, B. et al. Identification of novel missense mutation related with non-syndromic sensorineural deafness, DFNA11 in korean family by NGS. Genes Genom 45, 225–230 (2023). https://doi.org/10.1007/s13258-022-01357-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13258-022-01357-3