Abstract
Peroxiredoxins (Prxs) are cysteine-based peroxidases that play a central role in kee** the H2O2 at physiological levels. Eukaryotic cells express different Prxs isoforms, which differ in their subcellular locations and substrate specificities. Mitochondrial Prxs are synthesized in the cytosol as precursor proteins containing N-terminal cleavable presequences that act as mitochondrial targeting signals. Due to the fact that presequence controls the import of the vast majority of mitochondrial matrix proteins, the mitochondrial Prxs were initially predicted to be localized exclusively in the matrix. However, recent studies showed that mitochondrial Prxs are also targeted to the intermembrane space by mechanisms that remain poorly understood. While in yeast the IMP complex can translocate Prx1 to the intermembrane space, the maturation of yeast Prx1 and mammalian Prdx3 and Prdx5 in the matrix has been associated with sequential cleavages of the presequence by MPP and Oct1/MIP proteases. In this review, we describe the state of the art of the molecular mechanisms that control the mitochondrial import and maturation of Prxs of yeast and human cells. Once mitochondria are considered the major intracellular source of H2O2, understanding the mitochondrial Prx biogenesis pathways is essential to increase our knowledge about the H2O2-dependent cellular signaling, which is relevant to the pathophysiology of some human diseases.




Similar content being viewed by others
References
Anschau V, Ferrer-Sueta G, Aleixo-Silva RL et al (2020) Reduction of sulfenic acids by ascorbate in proteins, connecting thiol-dependent to alternative redox pathways. Free Radic Biol Med 156:207–216. https://doi.org/10.1016/j.freeradbiomed.2020.06.015
Araiso Y, Tsutsumi A, Qiu J et al (2019) Structure of the mitochondrial import gate reveals distinct preprotein paths. Nature 575:395–401. https://doi.org/10.1038/s41586-019-1680-7
Araki M, Nanri H, Ejima K et al (1999) Antioxidant function of the mitochondrial protein SP-22 in the cardiovascular system. J Biol Chem 274:2271–2278. https://doi.org/10.1074/jbc.274.4.2271
Yamamoto T, Matsui Y, Natori S, Obinata M (1989) Cloning of a housekee**-type gene (MER5) preferentially expressed in murine erythroleukemia cells. Gene 80:337–343. https://doi.org/10.1016/0378-1119(89)90297-7
Backes S, Hess S, Boos F, et al (2018) Tom70 enhances mitochondrial preprotein import efficiency by binding to internal targeting sequences. J Cell Biol 217:1369–1382. https://doi.org/10.1083/jcb.201708044
Balaban RS, Nemoto S, Finkel T (2005) Mitochondria, oxidants, and aging. Cell 120:483–495. https://doi.org/10.1016/j.cell.2005.02.001
Banmeyer I, Marchand C, Clippe A, Knoops B (2005) Human mitochondrial peroxiredoxin 5 protects from mitochondrial DNA damages induced by hydrogen peroxide. FEBS Lett 579:2327–2333. https://doi.org/10.1016/j.febslet.2005.03.027
Beasley EM, Müller S, Schatz G (1993) The signal that sorts yeast cytochrome b2 to the mitochondrial intermembrane space contains three distinct functional regions. EMBO J 12:2303–2311. https://doi.org/10.1002/j.1460-2075.1993.tb05884.x
Boldogh IR, Pon LA (2007) Purification and subfractionation of mitochondria from the yeast Saccharomyces cerevisiae. Methods Cell Biol 80:45–64. https://doi.org/10.1016/S0091-679X(06)80002-6
Brand MD (2020) Riding the tiger — physiological and pathological effects of superoxide and hydrogen peroxide generated in the mitochondrial matrix. Crit Rev Biochem Mol Biol 55:592–661. https://doi.org/10.1080/10409238.2020.1828258
Branda SS, Isaya G (1995) Prediction and identification of new natural substrates of the yeast mitochondrial intermediate peptidase. J Biol Chem 270:27366–27373. https://doi.org/10.1074/jbc.270.45.27366
Burkhart JM, Taskin AA, Zahedi RP, Vögtle F-N (2015) Quantitative profiling for substrates of the mitochondrial presequence processing protease reveals a set of nonsubstrate proteins increased upon proteotoxic stress. J Proteome Res 14:4550–4563. https://doi.org/10.1021/acs.jproteome.5b00327
Callegari S, Cruz-Zaragoza LD, Rehling P (2020) From TOM to the TIM23 complex — handing over of a precursor. Biol Chem 401:709–721. https://doi.org/10.1515/hsz-2020-0101
Calvo SE, Julien O, Clauser KR et al (2017) Comparative analysis of mitochondrial N-termini from mouse, human, and yeast. Mol Cell Proteomics 16:512–523. https://doi.org/10.1074/mcp.M116.063818
Cao Z, Bhella D, Lindsay JG (2007a) Reconstitution of the mitochondrial PrxIII antioxidant defence pathway: general properties and factors affecting PrxIII activity and oligomeric state. J Mol Biol 372:1022–1033. https://doi.org/10.1016/j.jmb.2007.07.018
Cao Z, Lindsay JG, Isaacs NW (2007b) Mitochondrial peroxiredoxins: structure and function. Subcell Biochem 44:295–315. https://doi.org/10.1007/978-1-4020-6051-9_14
Carrie C, Venne AS, Zahedi RP, Soll J (2015) Identification of cleavage sites and substrate proteins for two mitochondrial intermediate peptidases in Arabidopsis thaliana. J Exp Bot 66:2691–2708. https://doi.org/10.1093/jxb/erv064
Caumont-Sarcos A, Moulin C, Poinot L et al (2020) Transmembrane coordination of preprotein recognition and motor coupling by the mitochondrial presequence receptor Tim50. Cell Rep 30:3092-3104.e4. https://doi.org/10.1016/j.celrep.2020.02.031
Chacinska A, Koehler CM, Milenkovic D et al (2009) Importing mitochondrial proteins: machineries and mechanisms. Cell 138:628–644. https://doi.org/10.1016/j.cell.2009.08.005
Chang T-S, Cho C-S, Park S et al (2004) Peroxiredoxin III, a mitochondrion-specific peroxidase, regulates apoptotic signaling by mitochondria. J Biol Chem 279:41975–41984. https://doi.org/10.1074/jbc.M407707200
Collins Y, Chouchani ET, James AM et al (2012) Mitochondrial redox signalling at a glance. J Cell Sci 125:801–806. https://doi.org/10.1242/jcs.098475
Cox AG, Winterbourn CC, Hampton MB (2009) Mitochondrial peroxiredoxin involvement in antioxidant defence and redox signalling. Biochem J 425:313–325. https://doi.org/10.1042/BJ20091541
Culotta VC, Yang M, O’Halloran TV (2006) Activation of superoxide dismutases: putting the metal to the pedal. Biochim Biophys Acta 1763:747–758. https://doi.org/10.1016/j.bbamcr.2006.05.003
D’Autréaux B, Toledano MB (2007) ROS as signalling molecules: mechanisms that generate specificity in ROS homeostasis. Nat Rev Mol Cell Biol 8:813–824. https://doi.org/10.1038/nrm2256
De Armas MI, Esteves R, Viera N et al (2019) Rapid peroxynitrite reduction by human peroxiredoxin 3: implications for the fate of oxidants in mitochondria. Free Radic Biol Med 130:369–378. https://doi.org/10.1016/j.freeradbiomed.2018.10.451
De Simoni S, Goemaere J, Knoops B (2008) Silencing of peroxiredoxin 3 and peroxiredoxin 5 reveals the role of mitochondrial peroxiredoxins in the protection of human neuroblastoma SH-SY5Y cells toward MPP+. Neurosci Lett 433:219–224. https://doi.org/10.1016/j.neulet.2007.12.068
Demasi M, Augusto O, Bechara EJH, et al (2021) Oxidative modification of oroteins: from damage to catalysis, signaling, and beyond. Antioxid Redox Signal 35:1016–1080. https://doi.org/10.1089/ars.2020.8176
Edwards R, Gerlich S, Tokatlidis K (2020) The biogenesis of mitochondrial intermembrane space proteins. Biol Chem 401:737–747. https://doi.org/10.1515/hsz-2020-0114
Eldomery MK, Akdemir ZC, Vögtle FN et al (2016) MIPEP recessive variants cause a syndrome of left ventricular non-compaction, hypotonia, and infantile death. Genome Med 8:1–13. https://doi.org/10.1186/s13073-016-0360-6
Esser K, Jan P-S, Pratje E, Michaelis G (2004) The mitochondrial IMP peptidase of yeast: functional analysis of domains and identification of Gut2 as a new natural substrate. Mol Genet Genomics 271:616–626. https://doi.org/10.1007/s00438-004-1011-y
Figueira TR, Barros MH, Camargo AA et al (2013) Mitochondria as a source of reactive oxygen and nitrogen species: from molecular mechanisms to human health. Antioxid Redox Signal 18:2029–2074. https://doi.org/10.1089/ars.2012.4729
Gakh O, Cavadini P, Isaya G (2002) Mitochondrial processing peptidases. Biochim Biophys Acta 1592:63–77. https://doi.org/10.1016/s0167-4889(02)00265-3
Glick BS, Brandt A, Cunningham K et al (1992) Cytochromes c1 and b2 are sorted to the intermembrane space of yeast mitochondria by a stop-transfer mechanism. Cell 69:809–822. https://doi.org/10.1016/0092-8674(92)90292-k
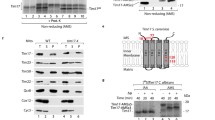
Gomes F, Palma FR, Barros MH et al (2017) Proteolytic cleavage by the inner membrane peptidase (IMP) complex or Oct1 peptidase controls the localization of the yeast peroxiredoxin Prx1 to distinct mitochondrial compartments. J Biol Chem 292:17011–17024. https://doi.org/10.1074/jbc.M117.788588
Gomes F, Turano H, Ramos A, Netto LES (2021) Assessment of submitochondrial protein localization in budding yeast Saccharomyces cerevisiae. J Vis Exp 1–16. https://doi.org/10.3791/62853
Greetham D, Grant CM (2009) Antioxidant activity of the yeast mitochondrial one-Cys peroxiredoxin is dependent on thioredoxin reductase and glutathione in vivo. Mol Cell Biol 29:3229–3240. https://doi.org/10.1128/MCB.01918-08
Hall A, Nelson K, Poole LB, Karplus PA (2011) Structure-based insights into the catalytic power and conformational dexterity of peroxiredoxins. Antioxid Redox Signal 15:795–815. https://doi.org/10.1089/ars.2010.3624
Hamanaka RB, Chandel NS (2010) Mitochondrial reactive oxygen species regulate cellular signaling and dictate biological outcomes. Trends Biochem Sci 35:505–513. https://doi.org/10.1016/j.tibs.2010.04.002
Hansen KG, Herrmann JM (2019) Transport of proteins into mitochondria. Protein J 38:330–342. https://doi.org/10.1007/s10930-019-09819-6
Hendrick JP, Hodges PE, Rosenberg LE (1989) Survey of amino-terminal proteolytic cleavage sites in mitochondrial precursor proteins: leader peptides cleaved by two matrix proteases share a three-amino acid motif. Proc Natl Acad Sci U S A 86:4056–4060. https://doi.org/10.1073/pnas.86.11.4056
Holmström KM, Finkel T (2014) Cellular mechanisms and physiological consequences of redox-dependent signalling. Nat Rev Mol Cell Biol 15:411–421. https://doi.org/10.1038/nrm3801
Huang S, Taylor NL, Whelan J, Millar AH (2009) Refining the definition of plant mitochondrial presequences through analysis of sorting signals, n-terminal modifications, and cleavage motifs. Plant Physiol 150:1272–1285. https://doi.org/10.1104/pp.109.137885
Hung V, Zou P, Rhee H-W et al (2014) Proteomic map** of the human mitochondrial intermembrane space in live cells via ratiometric APEX tagging. Mol Cell 55:332–341. https://doi.org/10.1016/j.molcel.2014.06.003
Ieva R, Heißwolf AK, Gebert M et al (2013) Mitochondrial inner membrane protease promotes assembly of presequence translocase by removing a carboxy-terminal targeting sequence. Nat Commun 4:2853. https://doi.org/10.1038/ncomms3853
Ieva R, Schrempp SG, Opaliński L et al (2014) Mgr2 functions as lateral gatekeeper for preprotein sorting in the mitochondrial inner membrane. Mol Cell 56:641–652. https://doi.org/10.1016/j.molcel.2014.10.010
Isaya G, Kalousek F, Rosenberg LE (1992) Amino-terminal octapeptides function as recognition signals for the mitochondrial intermediate peptidase. J Biol Chem 267:7904–7910
Isaya G, Miklos D, Rollins RA (1994) MIP1, a new yeast gene homologous to the rat mitochondrial intermediate peptidase gene, is required for oxidative metabolism in Saccharomyces cerevisiae. Mol Cell Biol 14:5603–5616. https://doi.org/10.1128/mcb.14.8.5603
Knoops B, Clippe A, Bogard C et al (1999) Cloning and characterization of AOEB166, a novel mammalian antioxidant enzyme of the peroxiredoxin family. J Biol Chem 274:30451–30458. https://doi.org/10.1074/jbc.274.43.30451
Knoops B, Goemaere J, Van der Eecken V, Declercq J-P (2011) Peroxiredoxin 5: structure, mechanism, and function of the mammalian atypical 2-Cys peroxiredoxin. Antioxid Redox Signal 15:817–829. https://doi.org/10.1089/ars.2010.3584
Kowaltowski AJ, de Souza-Pinto NC, Castilho RF, Vercesi AE (2009) Mitochondria and reactive oxygen species. Free Radic Biol Med 47:333–343. https://doi.org/10.1016/j.freeradbiomed.2009.05.004
Kropotov A, Usmanova N, Serikov V et al (2007) Mitochondrial targeting of human peroxiredoxin V protein and regulation of PRDX5 gene expression by nuclear transcription factors controlling biogenesis of mitochondria. FEBS J 274:5804–5814. https://doi.org/10.1111/j.1742-4658.2007.06103.x
Lee YJ (2020) Knockout Mouse Models for Peroxiredoxins. Antioxidants 9:182. https://doi.org/10.3390/antiox9020182
Li L, Shoji W, Takano H et al (2007) Increased susceptibility of MER5 (peroxiredoxin III) knockout mice to LPS-induced oxidative stress. Biochem Biophys Res Commun 355:715–721. https://doi.org/10.1016/j.bbrc.2007.02.022
Lu J, Holmgren A (2014) The thioredoxin antioxidant system. Free Radic Biol Med 66:75–87. https://doi.org/10.1016/j.freeradbiomed.2013.07.036
Meier S, Neupert W, Herrmann JM (2005) Proline residues of transmembrane domains determine the sorting of inner membrane proteins in mitochondria. J Cell Biol 170:881–888. https://doi.org/10.1083/jcb.200505126
Mokranjac D (2020) How to get to the other side of the mitochondrial inner membrane — the protein import motor. Biol Chem 401:723–736. https://doi.org/10.1515/hsz-2020-0106
Monteiro G, Horta BB, Pimenta DC et al (2007) Reduction of 1-Cys peroxiredoxins by ascorbate changes the thiol-specific antioxidant paradigm, revealing another function of vitamin C. Proc Natl Acad Sci U S A 104:4886–4891. https://doi.org/10.1073/pnas.0700481104
Monteiro G, Pereira GAG, Netto LES (2002) Regulation of mitochondrial thioredoxin peroxidase I expression by two different pathways: one dependent on cAMP and the other on heme. Free Radic Biol Med 32:278–288. https://doi.org/10.1016/S0891-5849(01)00801-2
Morgenstern M, Stiller SB, Lübbert P et al (2017) Definition of a high-confidence mitochondrial proteome at quantitative scale. Cell Rep 19:2836–2852. https://doi.org/10.1016/j.celrep.2017.06.014
Mossmann D, Meisinger C, Vögtle F-N (2012) Processing of mitochondrial presequences. Biochim Biophys Acta 1819:1098–1106. https://doi.org/10.1016/j.bbagrm.2011.11.007
Moulin C, Caumont-Sarcos A, Ieva R (2019) Mitochondrial presequence import: multiple regulatory knobs fine-tune mitochondrial biogenesis and homeostasis. Biochim Biophys Acta Mol Cell Res 1866:930–944. https://doi.org/10.1016/j.bbamcr.2019.02.012
Murphy MP (2009) How mitochondria produce reactive oxygen species. Biochem J 417:1–13. https://doi.org/10.1042/BJ20081386
Murphy MP (2012) Mitochondrial thiols in antioxidant protection and redox signaling: distinct roles for glutathionylation and other thiol modifications. Antioxid Redox Signal 16:476–495. https://doi.org/10.1089/ars.2011.4289
Netto LES, Antunes F (2016) The roles of peroxiredoxin and thioredoxin in hydrogen peroxide sensing and in signal transduction. Mol Cells 39:65–71. https://doi.org/10.14348/molcells.2016.2349
Nguyên-nhu NT, Berck J, Clippe A et al (2007) Human peroxiredoxin 5 gene organization, initial characterization of its promoter and identification of alternative forms of mRNA. Biochim Biophys Acta - Gene Struct Expr 1769:472–483. https://doi.org/10.1016/j.bbaexp.2007.05.004
Ott M, Amunts A, Brown A (2016) Organization and regulation of mitochondrial protein synthesis. Annu Rev Biochem 85:77–101. https://doi.org/10.1146/annurev-biochem-060815-014334
Pedrajas JR, McDonagh B, Hernández-Torres F et al (2016) Glutathione is the resolving thiol for thioredoxin peroxidase activity of 1-Cys peroxiredoxin without being consumed during the catalytic cycle. Antioxid Redox Signal 24:115–128. https://doi.org/10.1089/ars.2015.6366
Pedrajas JR, Miranda-Vizuete A, Javanmardy N et al (2000) Mitochondria of Saccharomyces cerevisiae contain one-conserved cysteine type peroxiredoxin with thioredoxin peroxidase activity. J Biol Chem 275:16296–16301. https://doi.org/10.1074/jbc.275.21.16296
Pedrajas JR, Padilla CA, McDonagh B, Bárcena JA (2010) Glutaredoxin participates in the reduction of peroxides by the mitochondrial 1-CYS peroxiredoxin in Saccharomyces cerevisiae. Antioxid Redox Signal 13:249–258. https://doi.org/10.1089/ars.2009.2950
Perkins A, Nelson KJ, Parsonage D et al (2015) Peroxiredoxins: guardians against oxidative stress and modulators of peroxide signaling. Trends Biochem Sci 40:435–445. https://doi.org/10.1016/j.tibs.2015.05.001
Pfanner N, Warscheid B, Wiedemann N (2019) Mitochondrial proteins: from biogenesis to functional networks. Nat Rev Mol Cell Biol 20:267–284. https://doi.org/10.1038/s41580-018-0092-0
Poveda-Huertes D, Mulica P, Vögtle FN (2017) The versatility of the mitochondrial presequence processing machinery: cleavage, quality control and turnover. Cell Tissue Res 367:73–81. https://doi.org/10.1007/s00441-016-2492-9
Quirós PM, Langer T, López-Otín C (2015) New roles for mitochondrial proteases in health, ageing and disease. Nat Rev Mol Cell Biol 16:345–359. https://doi.org/10.1038/nrm3984
Rath S, Sharma R, Gupta R et al (2021) MitoCarta3.0: an updated mitochondrial proteome now with sub-organelle localization and pathway annotations. Nucleic Acids Res 49:D1541–D1547. https://doi.org/10.1093/nar/gkaa1011
Rhee HW, Zou P, Udeshi ND et al (2013) Proteomic map** of mitochondria in living cells via spatially restricted enzymatic tagging. Science (80-) 339:1328–1331. https://doi.org/10.1126/science.1230593
Rhee SG, Chae HZ, Kim K (2005) Peroxiredoxins: a historical overview and speculative preview of novel mechanisms and emerging concepts in cell signaling. Free Radic Biol Med 38:1543–1552. https://doi.org/10.1016/j.freeradbiomed.2005.02.026
Rhee SG, Kil IS (2016) Mitochondrial H2O2 signaling is controlled by the concerted action of peroxiredoxin III and sulfiredoxin: linking mitochondrial function to circadian rhythm. Free Radic Biol Med 100:73–80. https://doi.org/10.1016/j.freeradbiomed.2016.10.011
Rhee SG, Kil IS (2017) Multiple functions and regulation of mammalian peroxiredoxins. Annu Rev Biochem 86:749–775. https://doi.org/10.1146/annurev-biochem-060815-014431
Rhee SG, Woo HA, Kil IS, Bae SH (2012) Peroxiredoxin functions as a peroxidase and a regulator and sensor of local peroxides. J Biol Chem 287:4403–4410. https://doi.org/10.1074/jbc.R111.283432
Richter-Dennerlein R, Dennerlein S, Rehling P (2015) Integrating mitochondrial translation into the cellular context. Nat Rev Mol Cell Biol 16:586–592. https://doi.org/10.1038/nrm4051
Riemer J, Schwarzländer M, Conrad M, Herrmann JM (2015) Thiol switches in mitochondria: operation and physiological relevance. Biol Chem 396:465–482. https://doi.org/10.1515/hsz-2014-0293
Schendzielorz AB, Bragoszewski P, Naumenko N et al (2018) Motor recruitment to the TIM23 channel’s lateral gate restricts polypeptide release into the inner membrane. Nat Commun 9:1–10. https://doi.org/10.1038/s41467-018-06492-8
Schendzielorz AB, Schulz C, Lytovchenko O et al (2017) Two distinct membrane potential-dependent steps drive mitochondrial matrix protein translocation. J Cell Biol 216:83–92. https://doi.org/10.1083/jcb.201607066
Schmidt O, Pfanner N, Meisinger C (2010) Mitochondrial protein import: from proteomics to functional mechanisms. Nat Rev Mol Cell Biol 11:655–667. https://doi.org/10.1038/nrm2959
Schulz C, Schendzielorz A, Rehling P (2015) Unlocking the presequence import pathway. Trends Cell Biol 25:265–275. https://doi.org/10.1016/j.tcb.2014.12.001
Sena LA, Chandel NS (2012) Physiological roles of mitochondrial reactive oxygen species. Mol Cell 48:158–167. https://doi.org/10.1016/j.molcel.2012.09.025
Seo MS, Kang SW, Kim K et al (2000) Identification of a new type of mammalian peroxiredoxin that forms an intramolecular disulfide as a reaction intermediate. J Biol Chem 275:20346–20354. https://doi.org/10.1074/jbc.M001943200
Sies H, Jones DP (2020) Reactive oxygen species (ROS) as pleiotropic physiological signalling agents. Nat Rev Mol Cell Biol 21:363–383. https://doi.org/10.1038/s41580-020-0230-3
Sies H, Reichert AS (2019) Selectively addressing mitochondrial glutathione and thioredoxin redox systems. Cell Chem Biol 26:316–318. https://doi.org/10.1016/j.chembiol.2019.02.017
Sim J, Park J, Woo HA, Rhee SG (2021) Maturation of mitochondrially targeted prx v involves a second cleavage by mitochondrial intermediate peptidase that is sensitive to inhibition by h2o2. Antioxidants 10:1–15. https://doi.org/10.3390/antiox10030346
Sriram SM, Kim BY, Kwon YT (2011) The N-end rule pathway: emerging functions and molecular principles of substrate recognition. Nat Rev Mol Cell Biol 12:735–747. https://doi.org/10.1038/nrm3217
Stein KT, Moon SJ, Nguyen AN, Sikes HD (2020) Kinetic modeling of H2O2 dynamics in the mitochondria of HeLa cells. PLoS Comput Biol 16:1–21. https://doi.org/10.1371/journal.pcbi.1008202
Stojanovski D, Bohnert M, Pfanner N, van der Laan M (2012) Mechanisms of protein sorting in mitochondria. Cold Spring Harb Perspect Biol 4:1–18. https://doi.org/10.1101/cshperspect.a011320
Taylor AB, Smith BS, Kitada S et al (2001) Crystal structures of mitochondrial processing peptidase reveal the mode for specific cleavage of import signal sequences. Structure 9:615–625. https://doi.org/10.1016/s0969-2126(01)00621-9
Teixeira PF, Glaser E (2013) Processing peptidases in mitochondria and chloroplasts. Biochim Biophys Acta 1833:360–370. https://doi.org/10.1016/j.bbamcr.2012.03.012
Toledano MB, Delaunay-Moisan A, Outten CE, Igbaria A (2013) Functions and cellular compartmentation of the thioredoxin and glutathione pathways in yeast. Antioxid Redox Signal 18:1699–1711. https://doi.org/10.1089/ars.2012.5033
Tucker K, Park E (2019) Cryo-EM structure of the mitochondrial protein-import channel TOM complex at near-atomic resolution. Nat Struct Mol Biol 26:1158–1166. https://doi.org/10.1038/s41594-019-0339-2
Vaca Jacome AS, Rabilloud T, Schaeffer-Reiss C et al (2015) N-terminome analysis of the human mitochondrial proteome. Proteomics 15:2519–2524. https://doi.org/10.1002/pmic.201400617
Van der Eecken V, Clippe A, Dekoninck S et al (2013) Abolition of peroxiredoxin-5 mitochondrial targeting during canid evolution. PLoS ONE 8:12–16. https://doi.org/10.1371/journal.pone.0072844
Veling MT, Reidenbach AG, Freiberger EC et al (2017) Multi-omic mitoprotease profiling defines a role for Oct1p in coenzyme Q production. Mol Cell 68:970-977.e11. https://doi.org/10.1016/j.molcel.2017.11.023
Vögtle F-N, Wortelkamp S, Zahedi RP et al (2009) Global analysis of the mitochondrial N-proteome identifies a processing peptidase critical for protein stability. Cell 139:428–439. https://doi.org/10.1016/j.cell.2009.07.045
Vögtle FN, Prinz C, Kellermann J et al (2011) Mitochondrial protein turnover: role of the precursor intermediate peptidase Oct1 in protein stabilization. Mol Biol Cell 22:2135–2143. https://doi.org/10.1091/mbc.E11-02-0169
Watabe S, Hiroi T, Yamamoto Y et al (1997) SP-22 is a thioredoxin-dependent peroxide reductase in mitochondria. Eur J Biochem 249:52–60. https://doi.org/10.1111/j.1432-1033.1997.t01-1-00052.x
Wiedemann N, Pfanner N (2017) Mitochondrial machineries for protein import and assembly. Annu Rev Biochem 86:685–714. https://doi.org/10.1146/annurev-biochem-060815-014352
Funding
Our work was supported by research grants from Fundação de Amparo à Pesquisa do Estado de São Paulo (FAPESP), grants 2013/07937–8 and 2013/08028–1. Fernando Gomes and Helena Turano are also supported by FAPESP, grants 2017/09443–3 and 2017/23839–7, respectively. Angélica Ramos is also supported by Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Gomes, F., Turano, H., Ramos, A. et al. Dissecting the molecular mechanisms of mitochondrial import and maturation of peroxiredoxins from yeast and mammalian cells. Biophys Rev 13, 983–994 (2021). https://doi.org/10.1007/s12551-021-00899-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12551-021-00899-2