Abstract
As an important forest type, deciduous broad-leaved forest is crucial for estimating forest carbon sequestration capacity and evaluating forest carbon balance. This study focuses on the natural deciduous broad-leaved forest of Mazongling Nature Reserve in **zhai County of China. WorldView-2 images were selected as data source. 36 candidate factors including vegetation indices, texture features, and topographic factors were used for modelling. Three machine learning algorithms (i.e., random forest, k-nearest neighbor, and artificial neural network) were used to establish the optimal quantitative retrieval model for natural deciduous broad-leaved biomass. Results showed that the ANN model was the best predictor with R2 = 0.69 and RMSE = 31.53 (Mg·ha−1). Combining the ANN model with the complete spatial coverage of remote sensing data, we developed a distribution map of natural deciduous broad-leaved biomass in the Mazongling forest farm. The estimated average biomass of the study area was 90.34 ± 47.96 Mg·ha−1. In addition, the influence of light saturation on model accuracy is also discussed. This study confirms that remote sensing data in temporal and spatial space can improve the model estimation accuracy.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Forest biomass is a basic measure for evaluating the forest ecosystem, and it is also an essential variable for quantifying the structure and function of the ecosystem (Paulo et al., 2012; Rodrguez-Veig et al. 2019). As an important part of the carbon cycle, effective forest biomass monitoring can help us understand the interactions between the biosphere and the atmosphere (Pang et al., 2017; Rödig et al., 2017; Zhang et al., 2019). Deciduous broad-leaved forest is one of the most widely distributed forest vegetation types in the world, and it plays an important role in regulating climate, as well as maintaining water and soil (Souza & Longhi, 2019). Recently, with increasing and changing climate, deciduous broad-leaved forests are facing unprecedented threats (Laurin et al., 2020; Pope et al., 2020). The effects of climate change on rangelands and broad-leaved forests were studied using free satellite data from the GEE platform in a recent research project (Orusa & Mondino, 2021). The use of remote sensing to estimate deciduous broad-leaved forest biomass plays an important role in the study of forest ecosystems and their contribution to the global carbon cycle.
Traditional biomass calculation methods have the defects of large workload and high costs, such as the clear-cutting method (Liu et al., 2020) and the standard wood method (Jiang et al., 2017). In addition, the regression method is also commonly used (Li et al., 2012; Zaki et al., 2018; Zhang et al., 2020). Therefore, it is challenging to meet the requirements of these methods for estimating forest biomass at large-scale (Han et al., 2019; Koju et al., 2019; Rodig et al., 2017; Wan et al., 2018). Remote sensing technology has the advantages of the wide detection range and short update time, so combining remote sensing data with a small sample set of ground survey data has become a useful approach to estimate forest biomass at large-scale (Gwenzi et al., 2017; Kankare et al., 2013). In temperate and subtropical regions, deciduous forest is the most typical forest type, and the study of deciduous forest biomass change has important implications for climate change (Ghosh & Behera, 2018; Landuyt et al., 2020; Raha et al., 2020). In terms of research data, the biomass estimation of deciduous forest was carried out mainly by optical remote sensing data and lidar data (Joshi & Dhyani, 2019; Kristen et al., 2018; Wang et al., 2020; Nandy et al., 2017; Senger et al., 2020). Environmental variables (e.g., rainfall, humidity and soil) can affect the horizontal distribution of species biomass (Fu et al., 2019). Additionally, some forest parameters, such as stand age, leaf area index and canopy closure, can also improve the accuracy of biomass estimation (Li et al., 2020a, 2016; Mutanga et al., 2012; Nguyen et al., 2018; Yang et al., 2018). Compared with linear regression model, machine learning can improve model accuracy when the biomass is more than 120 Mg·ha−1 (Gao et al., 2018). Most studies on aboveground biomass primarily focus on the coniferous forests, coniferous and broad-leaved mixed forests, and evergreen broad-leaved forests (Dai et al., 2016; Dimitrov & Roumenina, 2013; Hu et al., 2016; Luo et al., 2021; Nie et al., 2017; Shen et al., 2018; Stovall et al., 2017). However, there is limited research on combining optical remote sensing information with machine learning to estimate the biomass of natural deciduous broad-leaved forests.
This study focuses on the development of quantitative models for biomass in the natural deciduous broad-leaved forest of Mazongling Nature Reserve in China. Vegetation indices and texture information were extracted using Worldview-2 remote sensing data. Additionally, terrain factors extracted from DEM (Digital Elevation Model) and ground measured data were obtained. An optimal biomass remote sensing quantitative inversion model was constructed using a machine learning algorithm. This study estimated the biomass of forest and analyzed its distribution. Its results provide a scientific reference for the protection and utilization of forest resources in Mazongling Nature Reserve.
Materials and Methods
Overview of the Study Area
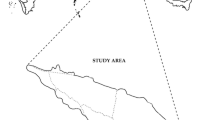
Mazongling Nature Reserve is located in the southwest of **zhai County, Anhui Province, China (115°31′-115°50′E, 31°10′-31°20′N; Fig. 1). It is one part of Anhui Tianma National Nature Reserve, with a total area of 4640.85 ha. The reserve belongs to the north subtropical humid monsoon climate zone, and it protects north subtropical evergreen-deciduous broad-leaved mixed forest as well as rare wild animals and plants. Tree species occurring on Mazongling Nature Reserve include Cunninghamia lanceolata (Lamb.) Hook., Pinus taiwanensis Hmyata, Quercus serrata var. brevipetiolata (A.DC.) Nakai, Castanea seguinii Dode, Cyclobalanopsis glauca (Thunb.) Oerst, and shrubs include Loropetalum chinense (R. Br.) Oliv., Rhododendron simsii Planch., Rhus chinensis Mill. The highest elevation in the reserve is 1671 m, the valley is vertical and horizontal, and the natural vegetation is lush. Its annual average temperature is 13.3 °C, and the average temperature in summer is 20 °C. The annual sunshine hours are 2225.5 h. Rainfall is abundant in the reserve, and the annual rainfall is 1480 mm.
Research Data
Sample Plot Data
The sampling survey was conducted from July 23 to 31, 2019. To comprehensively investigate the forest resources in the study area, stratified and typical sampling methods were used to establish 35 deciduous broad-leaved forest plots of different ages and site conditions. The sample plots were 20 m × 20 m. All the living trees in the plots with a diameter at breast height greater than 5 cm were measured, and tree heights were measured using a laser range finder. Differential GPS (DGPS) was used to determine the locations of sample plots. The dominant species in the study area were found to be hardwood tree species, so the forest biomass of sample plots were calculated using the general calculation method of hardwood biomass proposed by Li and Lei (2010). Based on the 6th and 7th Chinese National Forest Inventory data, Li and Lei proposed a calculation model for hardwood tree species after comparing three estimation methods (i.e., the Intergovernmental Panel on Climate Change method, the Continuous Biomass Expansion Factor method, and the Empirical (Regression) Model Estimation method). The model has been widely used in China due to its high accuracy and good applicability. Its specific formula is
where W (Mg·ha−1) is the forest biomass, D (cm) is the breast diameter, and H (m) is the tree height. The estimated biomass via Eq. (1), as well as the locations of 35 sample plots (Fig. 1, Table 1), were used to establish a forest biomass model by machine learning.
Remote Sensing Data
Worldview-2 satellite images from June 23, 2019 were used as the remote sensing data. The spatial resolution of panchromatic and multispectral images was 0.46 m and 1.85 m, respectively. Their band information is shown in Table 2. A radiation correction was conducted using ENVI5.3 software to obtain radiance data. The MODTRAN4 + radiative transfer model was used for atmospheric correction of radiance data and to obtain reflectivity data. Gram-Schimdt transform was used to fuse panchromatic images and multispectral data to obtain true color high-resolution images. 1:10,000 topographic maps were used to conduct geometric corrections for the remote sensing data, and their RMSEs were kept within 1 pixel.
Remote Sensing Classification of Forest Types
According to the size of sample plots, the characteristics of forest resources in the study area, and field investigation results, forest resources were categorized into four types: deciduous broad-leaved forest, coniferous forest, coniferous and broad-leaved mixed forest, and non-forest land. After the preprocessing of WorldView-2 data, RF, maximum likelihood method and Mahalanobis distance method were selected in ENVI5.3 to classify forest types. Verification data and the Kappa coefficient were used to test the classification accuracy. After classification, the majority/minority processing was conducted to classify broken patches from the original classification results into the category of background.
Feature Selection
The coordinate of the center point of each sample plot was chosen to be the center pixel. The average pixel value in a window size of 20 × 20 acted as the remote sensing feature. Vegetation index and gray-level co-occurrence matrix (GLCM) texture information were extracted. The window size of the GLCM texture information was defined as 9 × 9 after comparing different sizes, using the default 0° direction and a pixel statistical interval. Terrain factors, such as slope, aspect and elevation, were extracted from Digital Elevation Model data at a resolution of 12.5 m using the ArcGIS10.2 platform. 36 candidate factors were selected. They are NDVI, RVI, EVI, DVI, SAVI, MSAVI, B532_entropy, B3_entropy, B4_entropy, B5_entropy, B532_secondary moment, B3_secondary moment, B4_secondary moment, B5_secondary moment, B532_dissimilarity, B3_dissimilarity, B4_dissimilarity, B5_dissimilarity, B532_mean, B3_mean, B4_mean, B5_mean, B532_homogeneity, B3_homogeneity, B4_homogeneity, B5_homogeneity, B532_correlation, B5_correlation, B532_contrast, B3_contrast, B5_contrast, B532_variance, B3_variance, B4_variance, B5_variance, and Slope. The types and detailed descriptions of the modeling factors are shown in Table 3.
Model Variable Selection
Boruta and Recursive Feature Elimination (RFE) algorithms in R language were used to select variable sets related to the dependent variable. Boruta algorithm is based on the same idea of a random forest classifier. It adds randomness to the system and collects results from an ensemble of randomized samples and to assess the importance of each feature. This iterative process can reduce the misleading impact of random fluctuations and correlations (Amiri et al., 2019). RFE algorithm trains a model on a training set using all predictors. It calculates each variable importance and ranks them in order to seek an optimal variable set model. RFE seeks to improve generalization performance by removing the least important features whose deletion will have the least effect on training errors (Hayet et al., 2020). As the variables used by the Boruta algorithm could be highly correlated, we removed the highly correlated variables using the Pearson correlation coefficient. We set the threshold of the correlation coefficient to 0.9 to ensure that the absolute value of the correlation coefficient of all the prediction variables was below 0.9. This procedure could reduce the excessive abandonment of prediction variables due to the collinearity between prediction variables. Finally, b3_mean, b3_secondary moment, b3_variance, b4_secondary moment, b5_mean, slope, and NDVI were selected as predictors.
Machine Learning Algorithm
We used the k-NN, ANN, and RF machine learning algorithms in the platform of RStudio to construct a forest biomass model.
k-Nearest Neighbour (k-NN) Method
k-NN algorithm is a typical non-parametric algorithm, which estimates biomass based on the observation data of neighboring sampling points (Hoef & Temesgen, 2013). The basic principle of k-NN is that it finds k points, which are the k-nearest neighbors closest to the spatial distance from the prediction variable space of the training set, and it takes the average value of the k-nearest neighbor response variables to predict the value of the object (Mcroberts et al., 2016). Euclidean distance, a linear distance between two observations,\(d_{{(x_{a} ,x_{b} )}}\) is a common distance measure for constructing a forest biomass model based on k-NN. The formula is defined in Eq. (2).
where \(x_{a}\) and \(x_{b}\) are two sample points, and \(P\) is the dimension of each sample.
k-NN method is flexible and transparent, and it has strong generalization ability. However, when there are many features, many feature combinations will be generated, thus reducing the prediction efficiency and model accuracy. Therefore, the super parameter ‘k’, which means the k points closest to the target in the spatial distance, needs to be set when modelling in R language. If k is too small, then the modelling with training data is too sensitive, and the stability of the model is poor. If k is too large, the range of average value becomes too large, and the prediction error is large (Kumar et al., 2021). In practice, k ranges from 3 to 10.
Artificial Neural-Network (ANN) Method
ANN is a multi-layer feed-forward neural network with information forward propagation and error backward propagation (Fig. 2). Firstly, information is processed layer by layer from input layer to hidden layer, and outputs are compared with expected outputs. Reverse propagation is performed when the error between model outputs and expected outputs is greater than a predetermined value. Then, the internal weights and thresholds of the network are adjusted according to the prediction error, and the network is transferred to forward propagation again. This process is repeated until the error reaches the predetermined value, so that the outputs and the predictions are close enough to each other (Dong et al., 2020; Mao et al., 2019).
‘Decay’ and ‘size’ parameters are required when using the ‘nnet’ package of R language to build an ANN model. The parameter of “decay” is used as a penalty for the sum of squares of the weights. The use of “decay” can both help the optimization process and avoid over-fitting (Raji et al., 2020). ‘Decay’ was set as 0.001, 0.01, and 0.1 to reduce the possibility of over-training. ‘Size’ is defined as
where ‘size’ is the number of hidden units, P is the number of nodes in the input layer, O is the number of nodes in the output layer, and m is an integer constant between 0 and 10.
Random Forest (RF) Method
RF is a classifier that contains multiple decision trees, and it uses multiple decision-tree algorithms to carry out repeated predictions for the same inputs (Dong et al., 2020). Multiple random samples can be obtained to establish the corresponding decision trees through several rounds of bootstrap sampling. In this way, a random forest is formed.
The regression procedure of RF is achieved by using the ‘random forest’ data package in R software. Two key parameters are involved in this process: ntree and mtry. ‘Ntree’ is the number of decision trees, which is also the number of times that bootstrap is used to re-sample. ‘Mtry’ is the number of stochastic characteristics, which is also the number of input variables and usually one-third of the number of decision trees. However, ‘mtry’ needs to be tuned to achieve an optimal value (Tavares Júnior et al. 2020).
Model Accuracy Assessment
Model accuracy can be verified using leave-one-out cross-validation. That is to say, for N samples data, each available sample is taken as a test set, and the remaining N-1 samples are used as a training set. This procedure repeats N times, then N classifiers can be obtained, and the average on the results from N times is taken as the final performance index. This method uses almost all the samples to train the model, and the evaluation results are more reliable. There is no randomness and the entire process was repeatable (Wolfrum et al., 2020). The coefficient of determination (R2; Eq. (4)) and root mean square error (RMSE; Eq. (5)) were used to evaluate the models. Generally, greater R2 and lower RMSE indicate a better model fit.
where \(x_{i}\) is the measured value of the i-th sample plot, \(y_{i}\) is the model estimated value of the i-th sample plot, \(N\) represents the number of sample plot, \(\overline{x}\) represents the average value of the measured values, and \(\overline{y}\) is the average of the estimated values.
Results
Forest Type Classification in Mazongling Nature Reserve
The Kappa coefficients for the remote sensing classification of forest types using RF, maximum likelihood, and Mahalanobis distance methods were 0.97, 0.92, and 0.80, respectively. We selected the RF method with the greatest Kappa coefficient to classify the forest types (Fig. 3). Deciduous broad-leaved forest covered 2275.97 ha (49.04%), coniferous forest covered 1163.71 ha (25.08%), coniferous and broad-leaved mixed forest covered 735.38 ha (15.85%), and non-forest covered 465.78 ha (10.04%) of the total study area. Among these four different forest types, deciduous broad-leaved forest was primarily distributed in the Lingtou zone.
Construction of Remote Sensing Quantitative Model of Forest Biomass
The greatest R2 and the smallest RMSE from three models were determined by using leave-one-out cross-validation. The results are shown in Table 4 and Fig. 4.
-
1.
For the RF model, the maximum of RMSE was 36.83 Mg·ha−1 when the mtry was set as 1, and the minimum values was 32.27 Mg·ha−1 when the mtry was set as 7. The model precision was the highest when mtry was 7, R2 and RMSE were 0.68 and 31.85 Mg·ha−1, respectively.
-
2.
For the k-NN model, the maximum of RMSE was 46.11 Mg·ha−1 when k was 9, and the minimum values of RMSE was 40.74 when k was 5. RMSE gradually increased as k increased, and the model was the most accurate when k was 5, R2 and RMSE were 0.48, and 40.74 Mg·ha−1, respectively.
-
3.
For the ANN model, three different values of decay (0.001, 0.01, and 0.1) and hidden layers with sizes of 2 to 12 hide units were compared. The model was found to be the most accurate when decay = 0.1 and size = 2, R2 and RMSE were 0.69, and 31.53 Mg·ha−1, respectively.
Therefore, the most accurate ANN model was selected to construct the remote sensing quantitative estimation model of natural deciduous broad-leaved forest biomass in Mazongling Nature Reserve.
Spatial Distribution of Deciduous Broad-Leaved Forest Biomass in Mazongling Nature Reserve
The verification results of the optimal regression model using leave-one-out cross-validation are shown in Fig. 5. This ANN model had the most accurate prediction (R2 = 0.69, RMSE = 31.53 Mg·ha−1). Therefore, with this optimal ANN model, the above-ground-biomass (AGB) of natural deciduous broad-leaved forest was estimated using WorldView-2 images for Mazongling Nature Reserve (Fig. 6). The estimated biomass from this model was 90.34 ± 47.96 Mg·ha−1. The AGB of natural deciduous broad-leaved forest in Mazongling Nature Reserve was primarily distributed in Lingtou and Heshang** zones, followed by Dacao** and Dongshan zones. The lowest AGB (48 Mg·ha−1) was located in Qian** Village zone.
Discussion
Due to the complex vegetation and numerous tree species in sample plots, we did not use standard wood method. The biomass in the sample plots was calculated using the general calculation method of hardwood biomass proposed by Li and Lei based on the 6th and 7th Chinese National Forest Inventory data (Fu et al., 2022; Huang et al., 2022; Ju et al., 2022). The calculation of biomass of different tree organs is mainly based on the two parameters of tree height and DBH, and the R2 of height curves of other hard broad trees reaches 0.95. The number of sample plots should be increased in the future research that includes all age-class. Model accuracy can be improved by using the biomass model of the same zone, same family or same genus.
The WorldView-2 remote sensing image in this study was acquired in June 2019. Vegetation in the study area is in the growing season and is relatively lush. Because of the problems of different objects having the same spectrum and the same objects having different spectrum in the image (Ashutosh & Roy, 2021), there are omissions and mistakes when carrying out classifications, although its Kappa coefficient is very high. For example, the division of between "coniferous forest" and "coniferous and broad-leaved mixed forest", and that of between "broad-leaved forest" and "coniferous and broad-leaved mixed forest". The WorldView-2 remote sensing images used in this study had a small amount of cloud cover, which slightly impacted the classification of forest types and the inversion of forest biomass. However, it only accounted for 4.8% of the total area in the study area, which met the cloud content requirement (< 10%) for analyzing remote sensing images. Thus, the WorldView-2 images were not de-clouded so as to avoid detailed damaging after the de-clouding process. Regional image replacement can reduce the influence of cloud cover and improve image utilization. However, this study was based on only one year of remote sensing data (i.e., 2019). Therefore, further research on the spatial changes of forest biomass is necessary to improve the accuracy of model estimation.
The predictors selected in this study were able to construct a remote sensing quantitative model of deciduous broad-leaved in Mazongling Nature Reserve. However, the collinearity among predictors was insensitive, and the linear correlation between forest biomass and factors was not high. Additionally, there were positive and negative correlations between biomass and predictors. Therefore, it was not suitable to use a linear model to capture the relationship between biomass and remote sensing factors as well as geographic factors. However, an ANN model with strong nonlinear fitting ability was more suitable to decipher the relationship.
The results showed that the accuracy of the ANN model was the highest with R2 = 0.69. It is lower than that of the multiple linear regression biomass model (Wei, 2019). It is necessary to compare the multiple linear regression models with the machine learning model, so as to fully study the differences between the models and provide a sufficient basis for selecting a more accurate inversion model. Therefore, this study analyzed many references of machine learning algorithms for estimating forest biomass, especially for broad-leaved forest. The results showed that the difference of R2 and RMSE were a bit large. On the one hand, the biomass caused by normal growth is different due to different site conditions (soil, climate, terrain, etc.) of forest type. On the other hand, the difference of modeling candidate factors also plays an important role in model construction. In addition, it was less accurate than Antonio Montagnoli's model using lidar in the Alps (Montagnoli et al., 2015). This could be due to the light saturation in the WorldView-2 remote sensing images. The vegetation density of the deciduous broad-leaved forest in Mazongling Nature Reserve was so high that the electromagnetic radiation information received by remote sensing could no longer reflect changes in biomass. It led to inaccurate estimations for areas with high biomass, causing light saturation of biomass. As a result, the vegetation index and texture factor data fluctuated slightly in some areas, affecting model accuracy and biomass inversion. Therefore, further research is to determine the saturation point of remote sensing and improve the accuracy of remote sensing estimation of forest biomass. This study mainly focuses on the biomass modeling of deciduous broad-leaved forest. Biomass remote sensing inversion model of Pinus forest, Taxodium forest, coniferous and broad-leaved mixed forest, and mixed forest should have been constructed separately, which can help discuss and compare the consistency and difference between the mixed inversion model and the single forest type biomass model (Raj & Jhariya, 2021; Wang et al., Abid, F. (2021). A survey of machine learning algorithms based forest fires prediction and detection systems. Fire Technology, 57(2), 559–590. https://doi.org/10.1007/s10694-020-01056-z Amiri, M., Pourghasemi, H. R., Ghanbarian, G. A., & Afzali, S. F. (2019). Assessment of the importance of gully erosion effective factors using Boruta algorithm and its spatial modeling and map** using three machine learning algorithms. Geoderma, 340(1), 55–69. https://doi.org/10.1016/j.geoderma.2018.12.042 Ashutosh, S., & Roy, P. S. (2021). Three decades of nationwide forest cover map** using Indian remote sensing satellite data: A success story of monitoring forests for conservation in India. Journal of the Indian Society of Remote Sensing, 49(1), 61–70. https://doi.org/10.1007/s12524-020-01279-1 Balbinot, R., Trautenmüller, J. W., Caron, B. O., Junior, S. C., & Breunig, F. M. (2017). Vertical distribution of aboveground biomass in a seasonal deciduous forest. Revista Brasileirade Ciencias Agrarias, 12(3), 361–365. https://doi.org/10.5039/agraria.v12i3a5448 Dai, E. F., Wu, Z., Ge, Q. S., **, W. M., & Wang, X. F. (2016). Predicting the responses of forest distribution and aboveground biomass to climate change under RCP scenarios in southern China. Global Change Biology, 22(11), 3642–3661. https://doi.org/10.1111/gcb.13307 Dimitrov, P. K., & Roumenina, E. K. (2013). Combining SPOT 5 imagery with plotwise and standwise forest data to estimate volume and biomass in mountainous coniferous site. Central European Journal of Geosciences, 5(2), 208–222. https://doi.org/10.2478/s13533-012-0124-9 Dong, L. F., Du, H. Q., Han, N., Li, X. J., Zhu, D. E., Mao, F. J., Zhang, M., Zheng, J. L., Liu, H., Huang, Z. H., & He, S. B. (2020). Application of convolutional neural network on Lei Bamboo above-ground-biomass (AGB) estimation using Worldview-2. Remote Sensing, 12(6), 958. https://doi.org/10.3390/rs12060958 Fu, X., Zhang, Y. X., & Wang, X. J. (2022). Prediction of forest biomass carbon pool and carbon sink potential in China before 2060. Scientia Silvae Sinicae, 58(2), 32–41. https://doi.org/10.11707/j.1001-7488.20220204 Fu, Y. Y., He, H. S., Hawbaker, T. J., Henne, P. D., Zhu, Z. L., & Larsen, D. R. (2019). Evaluating k-Nearest Neighbor (kNN) imputation models for species-level aboveground fForest biomass map** in Northeast China. Remote Sensing, 11(17), 2005. https://doi.org/10.3390/rs11172005 Gao, Y. K., Lu, D. S., Li, G. Y., Wang, G. X., Chen, Q., Liu, L. J., & Li, D. Q. (2018). Comparative analysis of modeling algorithms for forest aboveground biomass estimation in a Subtropical Region. Remote Sensing, 10(4), 399–406. https://doi.org/10.3390/rs10040627 Ghosh, S. M., & Behera, M. D. (2018). Aboveground biomass estimation using multi-sensor data synergy and machine learning algorithms in a dense tropical forest. Applied Geography., 96, 29–40. https://doi.org/10.1016/j.apgeog.2018.05.011 Gumma, K. M., Thenkabail, S. P., Teluguntla, G. P., Oliphant, A., **ong, J., Giri, C., Pyla, V., Dixit, S., & Whitbread, M. A. (2020). Agricultural cropland extent and areas of South Asia derived using Landsat satellite 30-m time-series big-data using random forest machine learning algorithms on the Google Earth Engine cloud. Giscience & Remote Sensing, 57(3), 302–322. https://doi.org/10.1080/15481603.2019.1690780 Gwenzi, D., Helmer, E. H., Zhu, X. L., Lefsky, M. A., & Marcano, V. H. (2017). Predictions of tropical forest biomass and biomass growth based on stand height or canopy area are improved by Landsat-scale phenology across Puerto Rico and the U.S. Virgin Islands. Remote Sensing, 9(2), 123. https://doi.org/10.3390/rs9020123 Han, L., Yang, G. J., Dai, H. Y., Xu, B., Yang, H., Feng, H. K., Li, Z. H., & Yang, X. D. (2019). Modeling maize above-ground biomass based on machine learning approaches using UAV remote-sensing data. Plant Methods, 15(1), 1–19. https://doi.org/10.1186/s13007-019-0394-z Hayet, D., Zine, N. G., & Guessoum, S. (2020). Hybrid adapted fast correlation FCBF-support vector machine recursive feature elimination for feature selection. Intelligent Decision Technologies, 14(3), 269–279. https://doi.org/10.3233/IDT-190014 Hoef, J. M. V., & Temesgen, H. (2013). A comparison of the spatial linear model to Nearest Neighbor (k-NN) methods for forestry applications. PLoS ONE, 8(3), e59129. https://doi.org/10.1371/journal.pone.0059129 Hojo, A., Takagi, K., Avtar, R., Tadono, T., & Nakamura, F. (2020). Synthesis of L-Band SAR and forest heights derived from TanDEM-X DEM and 3 digital terrain models for biomass map**. Remote Sensing, 12(3), 349. https://doi.org/10.3390/rs12030349 Hu, Y. Q., Li, W. B., Cui, J. Y., & Su, Z. Y. (2016). Spatial point patterns of dominant species by individualtrees and biomass in a subtropical evergreen broad-leaved forest. Acta Ecologica Sinica, 36(4), 1066–1072. https://doi.org/10.3390/rs12030349 Huang, J. J., Liu, X. T., Zhang, Y. R., & Li, H. K. (2022). Stand biomass growth model of broadleaved forest with parameter classification in Guangdong Province of southern China. Journal of Bei**g Forestry University, 44(5), 19–33. https://doi.org/10.12171/j.1000-1522.20210403 Jian, Y. F., Han, Z. M., Huang, G. T., Wang, X., Li, Y., Zhou, J. J., & Dian, Y. Y. (2021). Estimation of forest biomass using high resolution remote sensing imagery in north subtropical forests. Acta Ecologica Sinica, 41(6), 2161–2169. https://doi.org/10.5846/stxb201910082086 Jiang, Z., Li, D. Y., Chen, B. B., Gao, H. G., Liu, C. H., Zhang, Z. R., Zhou, X., & Li, G. Q. (2017). Clonal growth of Hippophae Rhamniodes ssp. sinensis at the early stage in response to initial planting density and its regulation mechanism of biomass allocation. Scientia Silvae Sinicae, 53(10), 29–39. https://doi.org/10.11707/j.1001-7488.20171004 Joshi, R. K., & Dhyani, S. (2019). Biomass, carbon density and diversity of tree species in tropical dry deciduous forests in central India. Acta Ecologica Sinica, 39(4), 289–299. https://doi.org/10.1016/j.chnaes.2018.09.009 Ju, Y. L., Ji, Y. J., Huang, J. M., & Zhang, W. F. (2022). Inversion of forest aboveground biomass using combination of LiDAR and multispectral data. Journal of Nan**g Forestry University (natural Sciences Edition), 46(1), 58–68. https://doi.org/10.12302/j.issn.1000-2006.202109029 Júnior, I. D. S. T., Torres, C. M. M. E., Leite, H. G., Castro, N. L. M. D., Soares, C. P. B., Castro, R. V. O., & Farias, A. A. (2020). Machine learning: Modeling increment in diameter of individual trees on Atlantic Forest fragments. Ecological Indicators, 117, 106685. https://doi.org/10.1016/j.ecolind.2020.106685 Kaba, J. S., & Abunyewa, A. A. (2021). New aboveground biomass and nitrogen yield in different ages of gliricidia (Gliricidia Sepium Jacq.) trees under different pruning intensities in moist semi-deciduous forest zone of Ghana. Agroforestry Systems, 95(5), 835–842. https://doi.org/10.1007/s10457-019-00414-3 Kankare, V., Räty, M., Yu, X. W., Holopainen, M., Vastaranta, M., Kantola, T., Hyyppä, J., Hyyppä, H., Alho, P., & Viitala, R. (2013). Single tree biomass modelling using airborne laser scanning. ISPRS Journal of Photogrammetry and Remote Sensing, 85, 66–73. https://doi.org/10.1016/j.isprsjprs.2013.08.008 Koju, U. A., Zhang, J. H., Maharjan, S., Zhang, S., Bai, Y., Vijayakumar, D. B. I. P., & Yao, F. M. (2019). A two-scale approach for estimating forest aboveground biomass with optical remote sensing images in a subtropical forest of Nepal. Journal of Forestry Research, 30(6), 2119–2136. https://doi.org/10.1007/s11676-018-0743-1 Kristen, B., Quincey, J., & Margot, K. (2018). Spatial patterns of tree and shrub biomass in a deciduous forest using leaf-off and leaf-on lidar. Canadian Journal of Forest Research, 48(9), 1020–1033. https://doi.org/10.1139/cjfr-2018-0033 Kumar, K. U. P., Gandhi, O., Reddy, M. V., & Srinivasu, S. (2021). Usage of KNN, decision tree and random forest algorithms in machine learning and performance analysis with a comparative measure. Machine Intelligence and Soft Computing, 1280, 473–479. https://doi.org/10.1007/978-981-15-9516-5_39 Lakyda, P., Shvidenko, A., Bilous, A., Myroniuk, V., Matsala, M., Zibtsev, S., Schepaschenko, D., Holiaka, D., Vasylyshyn, R., Lakyda, I., et al. (2019). Impact of disturbances on the carbon cycle of forest ecosystems in Ukrainian Polissya. Forests, 10(4), 337. https://doi.org/10.3390/f10040337 Landuyt, D., Maes, S. L., Depauw, L., Ampoorter, E., Blondeel, H., Perring, M. P., et al. (2020). Drivers of above-ground understorey biomass and nutrient stocks in temperate deciduous forests. Journal of Ecology, 108(3), 982–997. https://doi.org/10.1111/1365-2745.13318 Laurin, G. V., Puletti, N., Grotti, M., Sterenczak, K., Modzelewska, K., Lisiewicz, M., Sadkowski, R., Kuberski, L., Chirici, G., & Papale, D. (2020). Species dominance and above ground biomass in the Białowieża Forest, Poland, described by airborne hyperspectral and lidar data. International Journal of Applied Earth Observation and Geoinformation, 92, 102178. https://doi.org/10.1016/j.jag.2020.102178 Li, C., Li, M. Y., & Li, Y. C. (2020a). Improving estimation of forest aboveground biomass using Landsat 8 imagery by incorporating forest crown density as a dummy variable. Canadian Journal of Forest Research, 50(4), 390–398. https://doi.org/10.1139/cjfr-2019-0216 Li, C. B., **ao, K. Y., Li, N., Song, X. L., Zhang, S., Wang, K., Chu, W. K., & Cao, R. (2020b). A comparative study of support vector machine, random forest and artificial neural network machine learning algorithms in geochemical anomaly information extraction. Acta Geoscientica Sinica, 41(2), 309–319. https://doi.org/10.3975/cagsb.2020.022501 Li, H. K., & Lei, Y. C. (2010). Estimation and evaluation of forest biomass carbon storage in China. China Forestry Publishing House. Li, H. K., Zhao, P. X., Lei, Y. C., & Zeng, W. S. (2012). Comparison on estimation of wood biomass using forest inventory data. Scientia Silvae Sinicae, 48(5), 44–52. https://doi.org/10.11707/j.1001-7488.20120507 Li, Y. C., Li, C., Li, M. Y., & Liu, Z. Z. (2019). Influence of variable selection and forest type on forest aboveground biomass estimation using machine learning algorithms. Forests, 10(12), 1073. https://doi.org/10.3390/f10121073 Li, Y. C., Li, M. Y., Liu, Z. Z., & Li, C. (2020c). Combining Kriging interpolation to improve the accuracy of forest aboveground biomass estimation using remote sensing data. IEEE Access, 8, 128124–128139. https://doi.org/10.1109/ACCESS.2020.3008686 Liao, K. T., Qi, S. H., Wang, C., & Wang, D. (2018). Estimation of forest aboveground biomass and canopy height in Jiangxi Province using GLAS and Landsat TM images. Remote Sensing Technology and Application, 33(4), 713–720. https://doi.org/10.11873/j.issn.1004-0323.2018.4.0713 Liu, F., Wang, C. K., Wang, X. C., Zhang, J. S., Zhang, Z., & Wang, J. J. (2016). Spatial patterns of biomass in the temperate broadleaved deciduous forest within the fetch of the Maoershan flux tower. Acta Ecologica Sinica, 36(20), 6506–6519. https://doi.org/10.5846/stxb201502270392 Liu, L. B., Zhou, Y. C., Cheng, A. Y., Wang, S. J., Cai, X. L., & Ni, J. (2020). Aboveground biomass estimate of a karst forest in central Guizhou Province, Southwestern China based on direct harvest method. Acta Ecologica Sinica, 40(13), 4455–4461. https://doi.org/10.5846/stxb201906141259 López-Serrano, P. M., Domínguez, J. L. C., Corral-Rivas, J. J., Jiménez, E., López-Sánchez, C. A., & Vega-Nieva, D. J. (2020). Modeling of aboveground biomass with Landsat 8 OLI and machine learning in temperate forests. Forests, 11(1), 11. https://doi.org/10.3390/f11010011 López-Serrano, P. M., López-Sánchez, C. A., Álvarez-González, J. G., & García-Gutiérrez, J. (2016). A comparison of machine learning techniques applied to Landsat-5 TM spectral data for biomass estimation. Canadian Journal of Remote Sensing, 42(6), 690–705. https://doi.org/10.1080/07038992.2016.121748 Luo, M., Wang, Y. F., **e, Y. H., Zhou, L., Qiao, J. J., Qiu, S. Y., & Sun, Y. J. (2021). Combination of feature selection and CatBoost for prediction: The first application to the estimation of aboveground biomass. Forests, 12(2), 216. https://doi.org/10.3390/f12020216 Ma, L., Li, W., Shi, N. N., Fu, S. L., Lian, J. Y., & Ye, W. H. (2019). Temporal and spatial patterns of aboveground biomass and its driving forces in a subtropical forest: A case study. Polish Journal of Ecology, 67(2), 95–104. https://doi.org/10.3161/15052249PJE2019.67.2.001 Mao, H. H., Meng, J. H., Ji, F. J., Zhang, Q. K., & Fang, H. T. (2019). Comparison of machine learning regression algorithms for Cotton Leaf Area Index retrieval using Sentinel-2 spectral bands. Applied Sciences, 9(7), 1459. https://doi.org/10.3390/app9071459 Mcroberts, R. E., Domke, G. M., Chen, Q., Næsset, E., & Gobakken, T. (2016). Using genetic algorithms to optimize k-Nearest Neighbors configurations for use with airborne laser scanning data. Remote Sensing of Environment, 184, 387–395. https://doi.org/10.1016/j.rse.2016.07.007 Montagnoli, A., Fusco, S., Terzaghi, M., Kirschbaum, A., Pflugmacher, D., Cohen, W. B., Scippa, G. S., & Chiatante, D. (2015). Estimating forest aboveground biomass by low density lidar data in mixed broad-leaved forests in the Italian Pre-Alps. Forest Ecosystems, 2(1), 10. https://doi.org/10.1186/s40663-015-0035-6 Mutanga, O., Adam, E., & Cho, M. A. (2012). High density biomass estimation for wetland vegetation using worldview-2 imagery and random forest regression algorithm. International Journal of Applied Earth Observation and Geoinformation, 18, 399–406. https://doi.org/10.1016/j.jag.2012.03.012 Nandy, S., Singh, R., Ghosh, S., Watham, T., Kushwaha, S. P. S., Kumar, A. S., & Dadhwal, V. K. (2017). Neural network-based modelling for forest biomass assessment. Carbon Management, 8(4), 305–317. https://doi.org/10.1080/17583004.2017.1357402 Nguyen, T. H., Jones, S., Soto-Berelov, M., Haywood, A., & Hislop, S. (2018). A comparison of imputation approaches for estimating forest biomass using Landsat time-series and inventory data. Remote Sensing, 10(11), 1825. https://doi.org/10.3390/rs10111825 Ni, W. J., Dong, J. C., Sun, G. Q., Zhang, Z. Y., Pang, Y., Tian, X., Li, Z. Y., & Chen, E. X. (2019). Synthesis of leaf-on and leaf-off Unmanned Aerial Vehicle (UAV) stereo imagery for the inventory of aboveground biomass of deciduous forests. Remote Sensing, 11(7), 889. https://doi.org/10.3390/rs11070889 Nie, S., Wang, C., Zeng, H. C., **, X. H., & Li, G. C. (2017). Above-ground biomass estimation using airborne discrete-return and full-waveform LiDAR data in a coniferous forest. Ecological Indicators, 78, 221–228. https://doi.org/10.1016/j.ecolind.2017.02.045 Ningthoujam, R. K., Joshi, P. K., & Roy, P. S. (2018). Retrieval of forest biomass for tropical deciduous mixed forest using ALOS PALSAR mosaic imagery and field plot data. International Journal of Applied Earth Observation & Geoinformation, 69, 206–216. https://doi.org/10.1016/j.jag.2018.03.007 Orusa, T., & Mondino, E. B. (2021). Exploring short-term climate change effects on rangelands and broad-leaved forests by free satellite data in Aosta Valley (Northwest Italy). Climate, 9(3), 47. https://doi.org/10.3390/cli9030047 Orusa, T., Viani, A., Cammareri, D., & Mondino, E. B. (2023). A Google Earth Engine algorithm to map phenological metrics in mountain areas worldwide with Landsat collection and Sentinel-2. Geomatics, 3(1), 221–238. https://doi.org/10.3390/geomatics3010012 Pang, Y., Meng, S. L., & Li, Z. Y. (2017). Temperate forest aboveground biomass estimation using Fourier-Based Textural Ordination (FOTO) indices from high resolution aerial optical image. Scientia Silvae Sinicae, 53(3), 94–104. https://doi.org/10.11707/j.1001-7488.20170311 Paulo, R. A., Adolfo, S., & Mario, J. M. (2012). The dynamics of land-use in Brazilian Amazon. Ecological Economics, 84, 23–36. https://doi.org/10.1016/j.ecolecon.2012.08.014 Peng, D. L., Zhang, H. L., Liu, L. Y., Huang, W. J., Huete, A. R., Zhang, X. Y., Wang, F. M., Yu, L., **e, Q. Y., Wang, C., Luo, S. Z., Li, C. J., & Zhang, B. (2019). Estimating the aboveground biomass for planted forests based on stand age and environmental variables. Remote Sensing, 11(19), 2270. https://doi.org/10.3390/rs11192270 Pope, R. J., Arnold, S. R., Chipperfield, M. P., Reddington, C. L. S., Butt, E. W., Keslake, T. D., Feng, W. H., Latter, B. G., Kerridge, B. J., & Siddans, R. (2020). Substantial Increases in Eastern Amazon and Cerrado Biomass Burning-Sourced Tropospheric Ozone. Geophysical Research Letters, 47(3), e2019GL084143. https://doi.org/10.11707/j.1001-7488.20170311 Raha, D., Dar, J. A., Pandey, P. K., Lone, P. A., Verma, S., Khare, P. K., & Khan, M. L. (2020). Variation in tree biomass and carbon stocks in three tropical dry deciduous forest types of Madhya Pradesh India. Carbon Management, 11(2), 109–120. https://doi.org/10.1080/17583004.2020.1712181 Raj, A., & Jhariya, M. K. (2021). Site quality and vegetation biomass in the tropical Sal mixed deciduous forest of Central India. Landscape and Ecological Engineering, 17(3), 387–399. https://doi.org/10.1007/s11355-021-00450-1 Raji, R. K., Adjeisah, M., Miao, X. H., & Wan, A. L. (2020). A novel respiration pattern biometric prediction system based on artificial neural network. Sensor Review, 40(1), 8–16. https://doi.org/10.1108/SR-10-2019-0235 Rödig, E., Huth, A., Bohn, F., Rebmann, C., & Cuntz, M. (2017). Estimating the carbon fluxes of forests with an individual-based forest model. Forest Ecosystems, 4, 4. https://doi.org/10.1186/s40663-017-0091-1 Rodríguez-Veiga, P., Quegan, S., Carreiras, J., Persson, H. J., Fransson, J. E. S., Hoscilo, A., et al. (2019). Forest biomass retrieval approaches from earth observation in different biomes. International Journal of Applied Earth Observation and Geoinformation, 77, 53–68. https://doi.org/10.1016/j.jag.2018.12.008 Senger, D. F., Hortua, D. A. S., Engel, S., Schnuawa, M., Moosdorf, N., & Gillis, L. G. (2020). Impacts of wetland dieback on carbon dynamics: A comparison between intact and degraded mangroves. Science of the Total Environment, 753, 141817. https://doi.org/10.1016/j.scitotenv.2020.141817 Shen, W. J., Li, M. S., Huang, C. Q., Tao, X., & Wei, A. S. (2018). Annual forest aboveground biomass changes mapped using ICESat/GLAS measurements, historical inventory data, and time-series optical and radar imagery for Guangdong province, China. Agricultural and Forest Meteorology, 259, 23–38. https://doi.org/10.1016/j.agrformet.2018.04.005 Souza, A. F., & Longhi, S. J. (2019). Disturbance history mediates climate change effects on subtropical forest biomass and dynamics. Ecology and Evolution, 9(12), 7184–7199. https://doi.org/10.1002/ece3.5289 Stovall, A. E. L., Vorster, A. G., Anderson, R. S., Evangelista, P. H., & Shugart, H. H. (2017). Non-destructive aboveground biomass estimation of coniferous trees using terrestrial LiDAR. Remote Sensing of Environment, 200, 31–42. https://doi.org/10.1016/j.rse.2017.08.013 Tavares Júnior, I. D. S., Moreira, M. E. T. C., Leite, H. G., Lemos, M. D. C. N, Boechat Soares, C. P., Oliveira Castro, R. V., & Farias, A. A. (2020). Machine learning: Modeling increment in diameter of individual trees on Atlantic Forest fragments. Ecological indicators, 117, 106685. https://doi.org/10.1016/j.ecolind.2020.106685 Wan, R. R., Wang, P., Wang, X. L., Yao, X., & Dai, X. (2018). Modeling wetland aboveground biomass in the Poyang Lake National Nature Reserve using machine learning algorithms and Landsat-8 imagery. Journal of Applied Remote Sensing, 12(4), 046029. https://doi.org/10.1117/1.JRS.12.046029 Wang, X. H., **ng, Y. Q., Huang, J. P., You, H. T., & Chang, X. Q. (2020). Combined spaceborne LiDAR and MODIS multispectral data to estimate regional biomass. Remote Sensing Information., 35(5), 140–147. https://doi.org/10.3969/j.issn.1000-3177.2020.05.018 Wei, X. M. (2019). Estimation of forest aboveground biomass based on multi-source data. Geomatics and Information Science of Wuhan University, 44(9), 1385–1390. https://doi.org/10.13203/j.whugis20190149 Wolfrum, E. J., Payne, C., Schwartz, A., Jacobs, J., & Kressin, R. W. (2020). A performance comparison of low-cost near-infrared (nir) spectrometers to a conventional laboratory spectrometer for rapid biomass compositional analysis. BioEnergy Research, 13(4), 1121–1129. https://doi.org/10.1007/s12155-020-10135-6 Wu, C. F., Shen, H. H., Wang, K., Shen, A. H., Deng, J. S., & Gan, M. Y. (2016). Landsat imagery-based above ground biomass estimation and change investigation related to human activities. Sustainability, 8, 159. https://doi.org/10.3390/su8020159 Yang, S. X., Feng, Q. S., Liang, T. G., Liu, B. K., Zhang, W. J., & **e, H. J. (2018). Modeling grassland above-ground biomass based on artificial neural network and remote sensing in the Three-River Headwaters Region. Remote Sensing of Environment, 204, 448–455. https://doi.org/10.1016/j.rse.2017.10.011 Zaki, N. A. M., Latif, Z. A., & Suratman, M. N. (2018). Modelling above-ground live trees biomass and carbon stock estimation of tropical lowland Dipterocarp forest: Integration of field-based and remotely sensed estimates. International Journal of Remote Sensing, 39(8), 2312–2340. https://doi.org/10.1080/01431161.2017.1421793 Zhang, Q. Y., Wang, H. B., Peng, D. L., Chen, J., & Liu, W. J. (2019). Estimation of forest vegetation carbon storage in Hunan Province, China based on k-NN method and domestic high-resolution data. Chinese Journal of Applied Ecology, 30, 3385–3394. https://doi.org/10.13287/j.1001-9332.201910.016 Zhang, Y. Z., Ma, J., Liang, S. L., Li, X. S., & Li, M. Y. (2020). An evaluation of eight machine learning regression algorithms for forest aboveground biomass estimation from multiple satellite data products. Remote Sensing, 12(24), 4015. https://doi.org/10.3390/rs12244015References
Acknowledgements
The authors are thankful to Taijun Guang and Hongchao Li for surveying and data processing in the study.
Funding
This research was funded by Anhui Provincial Natural Science Foundation (Grant NO.1808085QC74), Anhui Dabie Mountains Forest Ecosystem Research Station (Grant NO.2020132041), and Graduate Innovation Fund of Anhui Agricultural University (Grant NO.2020ysj-18).
Author information
Authors and Affiliations
Contributions
Conceptualization, Q.H., X.T. and D.Y.; Formal analysis, X.T., Q.H. and D.Y.; Funding acquisition, X.T. and Q.H.; Methodology, X.T. and Q.H.; Investigation, X.T., Q.O., M.X., P.F. and D.Y.; Software, D.Y., X.T. and Q.O.; Supervision, Q.H. and X.T.; Writing—original draft preparation, X.T. and H.L.; Writing—review and editing, Q.H. and X.T. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
This article is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
About this article
Cite this article
Tang, X., Yu, D., Lv, H. et al. Construction of Remote Sensing Quantitative Model for Biomass of Deciduous Broad-Leaved Forest in Mazongling Nature Reserve Based on Machine Learning. J Indian Soc Remote Sens (2024). https://doi.org/10.1007/s12524-024-01901-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12524-024-01901-6