Abstract
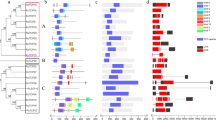
The two-component system (TCS) generally consists of three elements, namely the histidine kinase (HK), response regulator (RR), and histidine phosphotransfer (HP) gene families. This study aimed to assess the expression of TCS genes in P. vulgaris leaf tissue under salt and drought stress and perform a genome-wide analysis of TCS gene family members using bioinformatics methods. This study identified 67 PvTCS genes, including 10 PvHP, 38 PvRR, and 19 PvHK, in the bean genome. PvHK2 had the maximum number of amino acids with 1261, whilst PvHP8 had the lowest number with 87. In addition, their theoretical isoelectric points were between 4.56 (PvHP8) and 9.15 (PvPRR10). The majority of PvTCS genes are unstable. Phylogenetic analysis of TCS genes in A. thaliana, G. max, and bean found that PvTCS genes had close phylogenetic relationships with the genes of other plants. Segmental and tandem duplicate gene pairs were detected among the TCS genes and TCS genes have been subjected to purifying selection pressure in the evolutionary process. Furthermore, the TCS gene family, which has an important role in abiotic stress and hormonal responses in plants, was characterized for the first time in beans, and its expression of TCS genes in bean leaves under salt and drought stress was established using RNAseq and qRT-PCR analyses. The findings of this study will aid future functional and genomic studies by providing essential information about the members of the TCS gene family in beans.











Similar content being viewed by others
Abbreviations
- CHASE:
-
Cyclases/histidine kinases associated sensing extracellular
- HATPase:
-
Histidine kinase-like ATPase
- HisKA:
-
His kinase A
- HK:
-
Histidine kinase
- HP:
-
Histidine phosphotransfer
- RR:
-
Response regulator
- TCS:
-
Two-component system
References
Ahmad B, Azeem F, Ali MA, Nawaz MA, Nadeem H, Abbas A, Batool R, Atif RM, Ijaz U, Nieves-Cordones M, Chung G (2020) Genome-wide identification and expression analysis of two component system genes in Cicer arietinum. Genomics 112:1371–1383. https://doi.org/10.1016/J.YGENO.2019.08.006
Akdogan G, Tufekci ED, Uranbey S, Unver T (2016) miRNA-based drought regulation in wheat. Funct Integr Genomics 16:221–233. https://doi.org/10.1007/s10142-015-0452-1
Asfaw A, Ambachew D, Shah T, Blair MW (2017) Trait associations in diversity panels of the two common bean (Phaseolus vulgaris L.) gene pools grown under well-watered and water-stress conditions. Front Plant Sci 8:733. https://doi.org/10.3389/FPLS.2017.00733/BIBTEX
Aygören AS, Güneş E, Muslu S, Kasapoglu AG, Yigider E, Aydin M, Büyük I, Ilhan E (2022) Genome-wide analysis and characterization of SABATH gene family in phaseolus vulgaris genotypes subject to melatonin under drought and salinity stresses. Plant Mol Biol Rep 41:242–259. https://doi.org/10.1007/S11105-022-01363-5/FIGURES/13
Azarpazhooh E, Boye JI (2012) Composition of processed dry beans and pulses. In: Siddiq M, Uebersax MA (eds) Dry beans and pulses production, processing and nutrition. John Wiley & Sons Ltd, New York, pp 101–128
Bailey TL, Williams N, Misleh C, Li WW (2006) MEME: discovering and analyzing DNA and protein sequence motifs. Nucleic Acids Res 34:W369–W373. https://doi.org/10.1093/nar/gkl198
Bakhshi B, Fard EM (2023) The arrangement of MicroRNAs in the regulation of drought stress response in plants: a systematic review. Plant Mol Biol Rep. https://doi.org/10.1007/s11105-023-01380-y
Braun P, Gingras AC (2012) History of protein–protein interactions: from egg-white to complex networks. Proteomics 12:1478–1498. https://doi.org/10.1002/PMIC.201100563
Bryant P, Pozzati G, Elofsson A (2022) Improved prediction of protein–protein interactions using AlphaFold2. Nat Commun 13:1–11. https://doi.org/10.1038/s41467-022-28865-w
Cannon SB, Mitra A, Baumgarten A, Young ND, May G (2004) The roles of segmental and tandem gene duplication in the evolution of large gene families in Arabidopsis thaliana. BMC Plant Biol 4:1–21. https://doi.org/10.1186/1471-2229-4-10/TABLES/2
Chang C, Shockey JA (1999) The ethylene-response pathway: signal perception to gene regulation. Curr Opin Plant Biol 2:352–358. https://doi.org/10.1016/S1369-5266(99)00004-7
Chen C, Chen H, Zhang Y, Thomas HR, Frank MH, He Y, **a R (2020) TBtools: an integrative toolkit developed for interactive analyses of big biological data. Mol Plant 13:1194–1202. https://doi.org/10.1016/j.molp.2020.06.009
Conesa A, Götz S, García-Gómez JM, Terol J, Talon M, Robles M (2005) Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21:3674–3676. https://doi.org/10.1093/bioinformatics/bti610
Dhar YV, Lakhwani D, Pandey A, Singh S, Trivedi PK, Asif MH (2019) Genome-wide identification and interactome analysis of members of two-component system in Banana. BMC Genomics 20:1–15. https://doi.org/10.1186/s12864-019-6050-1
Garg R, Bhattacharjee A, Jain M (2015) Genome-scale transcriptomic insights into molecular aspects of abiotic stress responses in chickpea. Plant Mol Biol Rep 33:388–400. https://doi.org/10.1007/S11105-014-0753-X/FIGURES/6
Gasteiger E, Hoogland C, Gattiker A, Duvaud S, Wilkins MR, Appel RD, Bairoch A (2005) Protein identification and analysis tools on the ExPASy server. In: Walker JM (ed) The proteomics protocols handbook. Springer, Berlin. https://doi.org/10.1385/1-59259-890-0:571
Ghorbanzadeh Z, Hamid R, Jacob F, Mirzaei M, Zeinalabedini M, Abdirad S, Atwell BJ, Haynes PA, Ghaffari MR, Salekdeh GH (2023) MicroRNA profiling of root meristematic zone in contrasting genotypes reveals novel insight into in rice response to water deficiency. J Plant Growth Regul 42:3814–3834. https://doi.org/10.1007/s00344-022-10842-8
Golding C, Lee H (2023) Study and synthesis of melatonin as a strong antioxidant in plants. Biol Mol Chem 1:35–44. https://doi.org/10.22034/BMC.2023.417001.1006
Goodstein DM, Shu S, Howson R, Neupane R, Hayes RD, Fazo J, Mitros T, Dirks W, Hellsten U, Putnam N, Rokhsar DS (2012) Phytozome: a comparative platform for green plant genomics. Nucleic Acids Res 40:D1178–D1186. https://doi.org/10.1093/NAR/GKR944
Götz S, García-Gómez JM, Terol J, Williams TD, Nagaraj SH, Nueda MJ, Robles M, Talon M, Dopazo J, Conesa A (2008) High-throughput functional annotation and data mining with the Blast2GO suite. Nucleic Acids Res 36:3420–3435. https://doi.org/10.1093/NAR/GKN176
Guo AY, Zhu QH, Chen X, Luo JC (2007) GSDS: a gene structure display server. Yi Chuan = Hereditas / Zhongguo Yi Chuan Xue Hui Bian Ji 29:1023–1026. https://doi.org/10.1360/yc-007-1023
Guruprasad K, Reddy BVB, Pandit MW (1990) Correlation between stability of a protein and its dipeptide composition: a novel approach for predicting in vivo stability of a protein from its primary sequence. Protein Eng Des Sel 4:155–161. https://doi.org/10.1093/PROTEIN/4.2.155
Hailu B, Mehari H (2021) Impacts of soil salinity/sodicity on soil-water relations and plant growth in dry land areas: A review. J Nat Sci Res 12:1–10. https://doi.org/10.7176/jnsr/12-3-01
He Y, Liu X, Ye L, Pan C, Chen L, Zou T, Lu G (2016a) Genome-wide identification and expression analysis of two-component system genes in tomato. Int J Mol ScI 17:1204. https://doi.org/10.3390/IJMS17081204
He Y, Liu X, Zou T, Pan C, Qin L, Chen L, Lu G (2016b) Genome-Wide identification of two-component system genes in cucurbitaceae crops and expression profiling analyses in Cucumber. Front Plant Sci 7:899. https://doi.org/10.3389/FPLS.2016.00899/BIBTEX
Hiz MC, Canher B, Niron H, Turet M (2014) Transcriptome analysis of salt tolerant common bean (Phaseolus vulgaris L.) under saline conditions. PLoS ONE. https://doi.org/10.1371/journal.pone.0092598
Horton P, Park KJ, Obayashi T, Fujita N, Harada H, Adams-Collier CJ, Nakai K (2007) WoLF PSORT: protein localization predictor. Nucleic Acids Res 35:585–587. https://doi.org/10.1093/nar/gkm259
Huang X, Pearce R, Zhang Y (2020) EvoEF2: accurate and fast energy function for computational protein design. Bioinformatics 36:1135–1142. https://doi.org/10.1093/BIOINFORMATICS/BTZ740
Hussain I, Ayub A, Nayab A, Ashraf MA, Hussain S, Siddiqui MH, Sabir MA, Zulfiqar U, Khan TH (2023) Exogenous application of silicon and zinc attenuates drought tolerance in Eruca sativa L. through increasing chlorophyll pigments, osmoprotectants, and modulating defense mechanisms. J Plant Growth Regul. https://doi.org/10.1007/s00344-023-11116-7
Hutchison CE, Kieber JJ (2002) Cytokinin signaling in Arabidopsis. Plant Cell 14:S47–S59. https://doi.org/10.1105/TPC.010444
Hwang I, Chen HC, Sheen J (2002) Two-component signal transduction pathways in Arabidopsis. Plant Physiol 129:500–515. https://doi.org/10.1104/PP.005504
Hwang EW, Shin SJ, Park SC, Jeong MJ, Kwon HB (2011) Identification of miR172 family members and their putative targets responding to drought stress in Solanum tuberosum. Genes Genomics 33:105–110. https://doi.org/10.1007/s13258-010-0135-1
Hwang I, Sheen J, Müller B (2012) Cytokinin signaling networks. Annu Rev Plant Biol 63:353–380. https://doi.org/10.1146/annurev-arplant-042811-105503
İlhan E (2018) Genome-wide analysis of Eucalyptus grandis YABBY transcription factors. Turk Tarim Arast Derg 5:158–166
İncili ÇY, Arslan B, Çelik ENY, Ulu F, Horuz E, Baloglu MC, Çağlıyan E, Burcu G, Bayarslan AU, Altunoglu YC (2023) Comparative bioinformatics analysis and abiotic stress responses of expansin proteins in Cucurbitaceae members: watermelon and melon. Protoplasma 260:509–527. https://doi.org/10.1007/s00709-022-01793-8
Ishida K, Yamashino T, Yokoyama A, Mizuno T (2008) Three type-B response regulators, ARR1, ARR10 and ARR12, play essential but redundant roles in cytokinin signal transduction throughout the life cycle of Arabidopsis thaliana. Plant Cell Physiol 49:47–57. https://doi.org/10.1093/pcp/pcm165
**g X, Zhang H, Huai X, An Q, Qiao Y (2022) Identification and characterization of miRNAs and PHAS loci related to the early development of the embryo and endosperm in Fragaria × ananassa. BMC Genom 23:638. https://doi.org/10.1186/s12864-022-08864-3
Karimi R, Merati M, Shayganfar A (2023) Phytochemical characterization of five commercial Vitis vinifera cultivars in response to salinity. Acta Physiol Plant 45:108. https://doi.org/10.1007/s11738-023-03588-7
Karniol B, Wagner JR, Walker JM, Vierstra RD (2005) Phylogenetic analysis of the phytochrome superfamily reveals distinct microbial subfamilies of photoreceptors. Biochem J 392:103–116. https://doi.org/10.1042/BJ20050826
Kasapoğlu AG, Ilhan E, Kizilkaya D, Hossein-Pour A, Haliloğlu K (2020) Genome-wide analysis of BES1 transcription factor family in sorghum [Sorghum bicolor (L.) Moench] genome. Turk Tarim Arast Derg 7:85–95
Kelley LA, Mezulis S, Yates CM, Wass MN, Sternberg MJE (2015) The Phyre2 web portal for protein modeling, prediction and analysis. Nat Protoc 10(6):845–858. https://doi.org/10.1038/nprot.2015.053
Keskin O, Gursoy A, Ma B, Nussinov R (2008) Principles of protein–protein interactions: What are the preferred ways for proteins to interact? Chem Rev 108:1225–1244. https://doi.org/10.1021/cr040409x
Kiba T, Aoki K, Sakakibara H, Mizuno T (2004) Arabidopsis response regulator, ARR22, ectopic expression of which results in phenotypes similar to the wol cytokinin-receptor mutant. Plant Cell Physiol 45:1063–1077. https://doi.org/10.1093/PCP/PCH128
Kiba T, Naitou T, Koizumi N, Yamashino T, Sakakibara H, Mizuno T (2005) Combinatorial microarray analysis revealing Arabidopsis genes implicated in cytokinin responses through the His→Asp phosphorelay circuitry. Plant Cell Physiol 46:339–355. https://doi.org/10.1093/PCP/PCI033
Kizilkaya D, Kasapoğlu AG, Hosseinpour A, Haliloğlu K, Muslu S, Ilhan E (2020) Genome wide analysis of Sorghum bicolor L. CAMTA transcription factors. Atatürk Üniversitesi Ziraat Fakültesi Dergisi/atatürk Univ. J. Agric. Fac. 51:267–278
Krzywinski M, Schein J, Birol I, Connors J, Gascoyne R, Horsman D, Jones SJ, Marra MA (2009) Circos: an information aesthetic for comparative genomics. Genome Res 19:1639–1645. https://doi.org/10.1101/GR.092759.109
Kumar MN, Verslues PE (2015) Stress physiology functions of the Arabidopsis histidine kinase cytokinin receptors. Physiol Plant 154:369–380. https://doi.org/10.1111/PPL.12290
Lamesch P, Berardini TZ, Li D, Swarbreck D, Wilks C, Sasidharan R, Muller R, Dreher K, Alexander DL, Garcia-Hernandez M, Karthikeyan AS, Lee CH, Nelson WD, Ploetz L, Sing S, Wensel A, Huala E (2012) The Arabidopsis information resource (TAIR): improved gene annotation and new tools. Nucleic Acids Res 40:D1202–D1210. https://doi.org/10.1093/NAR/GKR1090
Le DT, Nishiyama R, Watanabe Y, Mochida K, Yamaguchi-Shinozaki K, Shinozaki K, Tran LSP (2011) Genome-wide expression profiling of soybean two-component system genes in soybean root and shoot tissues under dehydration stress. DNA Res 18:17–29. https://doi.org/10.1093/DNARES/DSQ032
Lescot M, Déhais P, Thijs G, Marchal K, Moreau Y, de Peer YV, Rouze P, Rombauts S (2002) PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res 30:325–327. https://doi.org/10.1093/nar/30.1.325
Letunic I, Bork P (2011) Interactive tree of LIFE v2: online annotation and display of phylogenetic trees made easy. Nucleic Acids Res 39:475–478. https://doi.org/10.1093/nar/gkr201
Li J, Zhang Z, Vang S, Yu J, Wong GKS, Wang J (2009) Correlation between Ka/Ks and Ks is related to substitution model and evolutionary lineage. J Mol Evol 68:414–423. https://doi.org/10.1007/S00239-009-9222-9/TABLES/5
Liddington RC (2004) Structural basis of protein–protein interactions. In: Fu H (ed) Protein-protein interactions: methods and applications. Humana Press, Totowa, pp 3–14
Liu Z, Zhang M, Kong L, Lv Y, Zou M, Lu G, Cao J, Yu X (2014) Genome-wide identification, phylogeny, duplication, and expression analyses of two-component system genes in Chinese cabbage (Brassica rapa ssp. pekinensis). DNA Res 21:379–396. https://doi.org/10.1093/dnares/dsu004
Liu P, Wang S, Wang X, Yang X, Li Q, Wang C, Chen C, Shi Q, Ren Z, Wang L (2020) Genome-wide characterization of two-component system (TCS) genes in melon (Cucumis melo L.). Plant Physiol Biochem 151:197–213. https://doi.org/10.1016/J.PLAPHY.2020.03.017
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25:402–408. https://doi.org/10.1006/METH.2001.1262
Lohrmann J, Harter K (2002) Plant Two-component signaling systems and the role of response regulators. Plant Physiol 128:363–369. https://doi.org/10.1104/PP.010907
Lohrmann J, Buchholz G, Keitel C, Swere U, Kircher S, Baurle I, Kudla J, Schafer E, Harter K (1999) Differential expression and nuclear localization of response regulator-like proteins from Arabidopsis thaliana. Plant Biol 1:495–505. https://doi.org/10.1055/S-2007-978544
Loomis WF, Shaulsky G, Wang N (1997) Histidine kinases in signal transduction pathways of eukaryotes. J Cell Sci 110:1141–1145. https://doi.org/10.1242/JCS.110.10.1141
Losa A, Vorster J, Cominelli E, Sparvoli F, Paolo D, Sala T, Ferrari M, Carbonaro M, Marconi S, Camilli E, Reboul E, Waswa B, Ekesa B, Aragão F, Kunert K (2022) Drought and heat affect common bean minerals and human diet—what we know and where to go. Food Energy Secur 11:e351. https://doi.org/10.1002/fes3.351
Lynch M, Conery JS (2000) The evolutionary fate and consequences of duplicate genes. Science (1979) 290:1151–1155. https://doi.org/10.1126/SCIENCE.290.5494.1151
Mähönen AP, Bishopp A, Higuchi M, Nieminen KM, Kınoshita K, Törmakangas K, Ikeda Y, Oka A, Kakimoto T, Helarıutta Y (2006) Cytokinin signaling and its inhibitor AHP6 regulate cell fate during vascular development. Science (1979) 311:94–98. https://doi.org/10.1126/SCIENCE.1118875/SUPPL_FILE/MAHONEN.SOM.PDF
Makino S, Kiba T, Imamura A, Hanaki N, Nakamura A, Suzuki T, Taniguchi M, Ueguchi C, Sugiyama T, Mizuno T (2000) Genes encoding pseudo-response regulators: insight into his-to-asp phosphorelay and circadian rhythm in Arabidopsis thaliana. Plant Cell Physiol 41:791–803. https://doi.org/10.1093/PCP/41.6.791
Marcantonini G, Bartolini D, Zatini L, Costa S, Passerini M, Rende M, Luca G, Basta G, Murdolo G, Calafiore R, Galli F (2022) Natural cryoprotective and cytoprotective agents in cryopreservation: A focus on melatonin. Molecules 27:3254. https://doi.org/10.3390/molecules27103254
Más P (2008) Circadian clock function in Arabidopsis thaliana: time beyond transcription. Trends Cell Biol 18:273–281. https://doi.org/10.1016/J.TCB.2008.03.005
Melotto M, Monteiro-Vitorello CB, Bruschi AG, Camargo LEA (2011) Comparative bioinformatic analysis of genes expressed in common bean (Phaseolus vulgaris L.) seedlings. Genome 48:562–570. https://doi.org/10.1139/G05-010
Miao Y, Gao X, Li B, Wang W, Bai L (2023) Low red to far-red light ratio promotes salt tolerance by improving leaf photosynthetic capacity in cucumber. Front Plant Sci 13:1053780. https://doi.org/10.3389/fpls.2022.1053780
Mir AR, Alam P, Hayat S (2022) Perspective of melatonin-mediated stress resilience and Cu remediation efficiency of Brassica juncea in Cu-contaminated soils. Front Plant Sci 13:910714. https://doi.org/10.3389/fpls.2022.910714
Mistry J, Chuguransky S, Williams L, Qureshi M, Salazar GA, Sonnhammer ELL, Tosatto SCE, Paladin L, Raj S, Richardson LJ, Finn RD, Bateman A (2021) Pfam: The protein families database in 2021. Nucleic Acids Res 49:D412–D419. https://doi.org/10.1093/NAR/GKAA913
Mizuno T (2004) Plant response regulators implicated in signal transduction and circadian rhythm. Curr Opin Plant Biol 7:499–505. https://doi.org/10.1016/J.PBI.2004.07.015
Mochida K, Yoshida T, Sakurai T, Yamaguchi-Shinozaki K, Shinozaki K, Tran LSP (2010) Genome-wide analysis of two-component systems and prediction of stress-responsive two-component system members in soybean. DNA Res 17:303–324. https://doi.org/10.1093/DNARES/DSQ021
Mortazavi A, Williams BA, McCue K, Schaeffer L, Wold B (2008) Map** and quantifying mammalian transcriptomes by RNA-Seq. Nat Methods 5:621–628. https://doi.org/10.1038/nmeth.1226
Muse SV (1996) Estimating synonymous and nonsynonymous substitution rates. Mol Biol Evol 13:105–114. https://doi.org/10.1093/OXFORDJOURNALS.MOLBEV.A025549
Nooren IMA, Thornton JM (2003) Diversity of protein–protein interactions. EMBO J 22:3486–3492. https://doi.org/10.1093/EMBOJ/CDG359
Pereira WJ, Melo AT, Coelho AS, Rodrigues FA, Mamidi S, Alencar SA, Lanna AC, Valdisser PA, Brondani C, Nascimento-Júnior IR, Borba TC (2020) Genome-wide analysis of the transcriptional response to drought stress in root and leaf of common bean. Genet Mol Biol 43:e20180259. https://doi.org/10.1590/1678-4685-GMB-2018-0259
Petrov NM, Stoyanova MI, Gaur RK (2019) Post-transcriptional gene silencing as a tool for controlling viruses in plants. Plant Biotechnol Prog Genom Era. https://doi.org/10.1007/978-981-13-8499-8_23
Petry N, Boy E, Wirth JP, Hurrell RF (2015) Review: the potential of the common Bean (Phaseolus vulgaris) as a vehicle for iron biofortification. Nutrients 7:1144–1173. https://doi.org/10.3390/NU7021144
Pils B, Heyl A (2009) Unraveling the evolution of cytokinin signaling. Plant Physiol 151:782–791. https://doi.org/10.1104/PP.109.139188
Quevillon E, Silventoinen V, Pillai S, Harte N, Mulder N, Apweiler R, Lopez R (2005) InterProScan: protein domains identifier. Nucleic Acids Res 33:116–120. https://doi.org/10.1093/nar/gki442
Ramasamy K, Karuppasami KM, Alagarswamy S, Shanmugam KP, Rathinavelu S, Vellingiri G, Muniyappan U, Kanthan T, Kuppusamy A, Rajendran M, Kathirvel A, Kanagarajan S (2023) Role of melatonin in directing plant physiology. Agron 13:2405. https://doi.org/10.3390/agronomy13092405
Raza A, Tabassum J, Fakhar AZ, Sharif R, Chen H, Zhang C, Ju L, Fotopoulos V, Siddique KH, Singh RK, Zhuang W (2022a) Smart reprograming of plants against salinity stress using modern biotechnological tools. Crit Rev Biotechnol. https://doi.org/10.1080/07388551.2022.2093695
Raza A, Charagh S, García-Caparrós P, Rahman MA, Ogwugwa VH, Saeed F, ** W (2022b) Melatonin-mediated temperature stress tolerance in plants. GM Crops Food 13:196–217
Raza A, Salehi H, Rahman MA, Zahid Z, MadadkarHaghjou M, Najafi-Kakavand S, Charagh S, Osman HSİ, Albaqami M, Zhuang Y, Siddique KHM, Zhuang W (2022c) Plant hormones and neurotransmitter interactions mediate antioxidant defenses under induced oxidative stress in plants. Front Plant Sci. https://doi.org/10.3389/fpls.2022.961872
Raza A, Mubarik MS, Sharif R, Habib M, Jabeen W, Zhang C, Chen H, Chen ZH, Siddique KHM, Zhuang W, Varshney RK (2023a) Develo** drought-smart, ready-to-grow future crops. Plant Genome 16:e20279. https://doi.org/10.1002/tpg2.20279
Raza A, Charagh S, Salehi H, Abbas S, Saeed F, Poinern GE, Siddique KHM, Varshney RK (2023b) Nano-enabled stress-smart agriculture: Can nanotechnology deliver drought and salinity-smart crops? J Sustain Agric Environ 2:189–214
Repik A, Rebbapragada A, Johnson MS, Haznedar JÖ, Zhulin IB, Taylor BL (2000) PAS domain residues involved in signal transduction by the Aer redox sensor of Escherichia coli. Mol Microbiol 36:806–816. https://doi.org/10.1046/J.1365-2958.2000.01910.X
Riaño-Pachón DM, Corrêa LGG, Trejos-Espinosa R, Mueller-Roeber B (2008) Green transcription factors: a chlamydomonas overview. Genetics 179:31–39. https://doi.org/10.1534/GENETICS.107.086090
Rodiño AP, Riveiro M, De Ron AM (2020) Implications of the symbiotic nitrogen fixation in common bean under seasonal water stress. Agron 11:70. https://doi.org/10.3390/agronomy11010070
Sánchez-Reinoso AD, Ligarreto-Moreno GA, Restrepo-Díaz H (2020) Evaluation of drought indices to identify tolerant genotypes in common bean bush (Phaseolus vulgaris L.). J Integr Agric 19:99–107. https://doi.org/10.1016/S2095-3119(19)62620-1
Schaller GE (2000) Histidine kinases and the role of two-component systems in plants. Adv Bot Res 32:109–148. https://doi.org/10.1016/S0065-2296(00)32023-7
Schaller GE, Kieber JJ, Shiu S-H (2008) Two-component signaling elements and histidyl-aspartyl phosphorelays. Arabidopsis Book 2008:e0112. https://doi.org/10.1199/tab.0112
Schmutz J, Cannon SB, Schlueter J, Ma J, Mitros T, Nelson W, Hyten DL, Song Q, Thelen JJ, Cheng J, Xu D (2010) Genome sequence of the palaeopolyploid soybean. Nature 463:178–183. https://doi.org/10.1038/nature08670
Schmutz J, McClean PE, Mamidi S, Wu GA, Cannon SB, Grimwood J, Jenkins J, Shu S, Song Q, Chavarro C, Torres-Torres M (2014) A reference genome for common bean and genome-wide analysis of dual domestications. Nat Genet 46:707–713. https://doi.org/10.1038/ng.3008
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13:2498–2504. https://doi.org/10.1101/GR.1239303
Shida C, Castrucci AML, Lamy-Freund MT (1994) High melatonin solubility in aqueous medium. J Pineal Res 16:198–201. https://doi.org/10.1111/J.1600-079X.1994.TB00102.X
Singh G, Kumar R (2012) Genome-wide Insilico analysis of plant two component signaling system in woody model plant Populus trichocarpa. Res Plant Biol 2:13–23
Singroha G, Sharma P, Sunkur R (2021) Current status of microRNA-mediated regulation of drought stress responses in cereals. Physiol Plant 172:1808–1821. https://doi.org/10.1111/ppl.13451
Stock AM, Robinson VL, Goudreau PN (2000) Two-component signal transduction. Annu Rev Biochem 69:183–215. https://doi.org/10.1146/annurev.biochem.69.1.183
Suleymanov A, Gabbasova I, Komissarov M, Suleymanov R, Garipov T, Tuktarova I, Belan L (2023) Random forest modeling of soil properties in saline semi-arid areas. Agriculture 13:976. https://doi.org/10.3390/agriculture13050976
Suyama M, Torrents D, Bork P (2006) PAL2NAL: robust conversion of protein sequence alignments into the corresponding codon alignments. Nucleic Acids Res 34:W609–W612. https://doi.org/10.1093/NAR/GKL315
Szklarczyk D, Gable AL, Nastou KC, Lyon D, Kirsch R, Pyysalo S, Doncheva NT, Legeay M, Fang T, Bork P, Jensen LJ, von Mering C (2021) The STRING database in 2021: customizable protein–protein networks, and functional characterization of user-uploaded gene/measurement sets. Nucleic Acids Res 49:D605–D612. https://doi.org/10.1093/NAR/GKAA1074
Taniguchi M, Sasaki N, Tsuge T, Aoyama T, Oka A (2007) ARR1 directly activates cytokinin response genes that encode proteins with diverse regulatory functions. Plant Cell Physiol 48:263–277. https://doi.org/10.1093/PCP/PCL063
Terzi E, Bilgintürk M, Gündoğan R, Can Karaça A (2020) Fasulye Proteini İzolatının Çeşitli Gıda Ürünlerinin Kalite Özelliklerine Etkisi. Avrupa Bilim Ve Teknoloji Dergisi. https://doi.org/10.31590/EJOSAT.757599. (In Turkish)
Thomason P, Kay R (2000) Eukaryotic signal transduction via histidine-aspartate phosphorelay. J Cell Sci 113:3141–3150. https://doi.org/10.1242/JCS.113.18.3141
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882. https://doi.org/10.1093/nar/25.24.4876
Thung M, Rao IM (1999) Integrated management of abiotic stresses. In: Singh SP (ed) Common bean improvement in the twenty-first century. Springer, Dordrecht, pp 331–370
Tsai YC, Weir NR, Hill K, Zhang W, Kim HJ, Shiu SH, Schaller GE, Kieber JJ (2012) Characterization of genes involved in cytokinin signaling and metabolism from rice. Plant Physiol 158:1666–1684. https://doi.org/10.1104/PP.111.192765
Urao T, Yakubov B, Yamaguchi-Shinozaki K, Shinozaki K (1998) Stress-responsive expression of genes for two-component response regulator-like proteins in Arabidopsis thaliana. FEBS Lett 427:175–178. https://doi.org/10.1016/S0014-5793(98)00418-9
Wang Y, Tang H, Debarry JD, Tan X, Li J, Wang X, Tae-ho L, ** H, Marler B, Guo H, Kissinger JC, Paterson AH (2012) MCScanX: a toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res 40:e49–e49. https://doi.org/10.1093/NAR/GKR1293
Weeda S, Zhang N, Zhao X, Ndip G, Guo Y, Buck GA, Fu C, Ren S (2014) Arabidopsis transcriptome analysis reveals key roles of melatonin in plant defense systems. PLoS ONE 9:e93462. https://doi.org/10.1371/JOURNAL.PONE.0093462
Wohlbach DJ, Quirino BF, Sussman MR (2008) Analysis of the Arabidopsis histidine kinase ATHK1 reveals a connection between vegetative osmotic stress sensing and seed maturation. Plant Cell 20:1101–1117. https://doi.org/10.1105/TPC.107.055871
**ang G, Jia R, Wang F, Wang S, Li Y, Yao Y (2023) Exogenous l-tryptophan treatment increases the concentrations of melatonin and aroma compounds in grape berries and wine. Food Qual Saf. https://doi.org/10.1093/fqsafe/fyad042
Ya-Jiao PA, Di WA, Ling-Hua ZH, Bin-Ying FU, Zhi-Kang LI (2009) Differential expressions of two-component element genes in rice under drought stress. Acta Agron Sin 35:1628–1636. https://doi.org/10.1016/S1875-2780(08)60105-4
Yamaguchi-Shinozaki K, Shinozaki K (2005) Organization of cis-acting regulatory elements in osmotic- and cold-stress-responsive promoters. Trends Plant Sci 10:88–94. https://doi.org/10.1016/J.TPLANTS.2004.12.012
Yang Z (2007) PAML 4: phylogenetic analysis by maximum likelihood. Mol Biol Evol 24:1586–1591. https://doi.org/10.1093/MOLBEV/MSM088
Yang T, Lv R, Li J, Lin H, ** D (2018) Phytochrome A and B negatively regulate salt stress tolerance of Nicotiana tobacum via ABA–jasmonic acid synergistic cross-talk. Plant Cell Physiol 59:2381–2393. https://doi.org/10.1093/PCP/PCY164
Yang S, Chu N, Feng N, Zhou B, Zhou H, Deng Z, Shen X, Zheng D (2023) Global responses of autopolyploid sugarcane Badila (Saccharum officinarum L.) to drought stress based on comparative transcriptome and metabolome profiling. Int J Mol Sci 24:3856. https://doi.org/10.3390/ijms24043856
Yavaş İ, Hussain S (2022) Recent progress on melatonin-induced salinity tolerance in plants: an overview. Turk JAF Sci Tech 10:1447–1454. https://doi.org/10.24925/turjaf.v10i8.1447-1454.5125
Zhang BH, Pan XP, Wang QL, Cobb GP, Anderson TA (2005) Identification and characterization of new plant microRNAs using EST analysis. Cell Res 15:336–360. https://doi.org/10.1038/sj.cr.7290302
Zhao Y, Li X, Ye C, Huang C, Lv X, Li J (2023) The biogenesis, mechanism and function of the tRNA-derived small RNA (tsRNA): a review compared with microRNA. Am J Cancer Res 13:1656
Acknowledgements
The authors gratefully acknowledge financial support from Atatürk University.
Funding
This research was funded by Atatürk University Scientific Research Project Coordination Unit with project no: FCD-2021-9813.
Author information
Authors and Affiliations
Contributions
Conceptualization, A.G.K, E.I., M.A., and E.Y.; methodology, A.G.K, E.I., M.A., B.I., I.B., and E.Y.; validation, M.S.T., G.A., and A.C.; formal analysis, A.G.K, E.I., M.A., B.I., I.B., and E.Y.; investigation, A.G.K, E.I., M.A., and E.Y.; writing-original draft preparation, A.G.K, E.I., M.A., and E.Y.; writing-review and editing, A.G.K, E.I., M.A., B.I., I.B., M.S.T., G.A., A.C., and E.Y.; visualization, A.G.K, E.I., M.A., B.I., I.B., and E.Y.; supervision, A.G.K, E.I., and M.A.; project administration, A.G.K, E.I., M.A., and E.Y. The published version of the manuscript has been reviewed and approved by all authors.
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kasapoglu, A.G., Ilhan, E., Aydin, M. et al. Characterization of Two-Component System gene (TCS) in melatonin-treated common bean under salt and drought stress. Physiol Mol Biol Plants 29, 1733–1754 (2023). https://doi.org/10.1007/s12298-023-01406-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12298-023-01406-5