Abstract
This study employs network toxicology, molecular docking, and molecular dynamics simulations to assess the characteristics and potential molecular mechanisms of aspartame-induced hepatocellular carcinoma (HCC). Utilizing ChEMBL, STITCH, and SwissTargetPrediction databases, potential target proteins associated with aspartame are identified. HCC-related targets are determined through bioinformatics and weighted gene co-expression network analysis (WGCNA). Gene enrichment analysis explores the signaling pathways related to aspartame-induced HCC. Further refinement using the STRING database and Cytoscape software highlights 15 key targets. Molecular docking, conducted using Autodock Vina, assesses the relationships between aspartame and each key target. Molecular dynamics simulations evaluate the binding capabilities of aspartame with core targets obtained through molecular docking. The results indicate that aspartame may induce HCC by modulating apoptosis and proliferation of liver cancer cells, affecting inflammatory signaling pathways, and regulating estrogen metabolism, posing to the occurrence and development of liver toxicity and associated inflammation, thereby leading to a risk of hepatocarcinogenesis. This study provides a theoretical foundation for understanding the molecular mechanisms underlying aspartame-induced HCC. Additionally, our network toxicology approach accelerates the elucidation of toxic pathways for uncharacterized food additives.
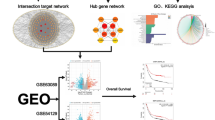
Graphical Abstract










Similar content being viewed by others
Data Availability
Data will be made available on request.
References
Chappell GA, Thompson CM, Wolf JC, Cullen JM, Klaunig JE, Haws LC (2020) Assessment of the mode of action underlying the effects of GenX in mouse liver and implications for assessing human health risks. Toxicol Pathol 48(3):494–508. https://doi.org/10.1177/0192623320905803
Chen H, Wong CC, Liu D, Go MYY, Wu B, Peng S, Yu J (2019) APLN promotes hepatocellular carcinoma through activating PI3K/Akt pathway and is a druggable target. Theranostics 9(18):5246–5260. https://doi.org/10.7150/thno.34713
Cui Q, Wang X, Zhang Y, Shen Y, Qian Y (2023) Macrophage-derived MMP-9 and MMP-2 are closely related to the rupture of the fibrous capsule of hepatocellular carcinoma leading to tumor invasion. Biol Proced Online 25(1):8. https://doi.org/10.1186/s12575-023-00196-0
Daina A, Michielin O, Zoete V (2019) SwissTargetPrediction: updated data and new features for efficient prediction of protein targets of small molecules. Nucleic Acids Res 47(W1):W357–W364. https://doi.org/10.1093/nar/gkz382
Dorokhov YL, Shindyapina AV, Sheshukova EV, Komarova TV (2015) Metabolic methanol: molecular pathways and physiological roles. Physiol Rev 95(2):603–644. https://doi.org/10.1152/physrev.00034.2014
EFSA (2013) Scientific opinion on the re-evaluation of aspartame (E 951) as a food additive. EFSA J 11(12). https://doi.org/10.2903/j.efsa.2013.3496
Fatma Gönül S, Emine D, Birol S (2021) Current approaches to the use of artificial sweetener aspartame. GSC Biol Pharm Sci 14(3):036–041. https://doi.org/10.30574/gscbps.2021.14.3.0044
Gaulton A, Bellis LJ, Bento AP, Chambers J, Davies M, Hersey A, Overington JP (2012) ChEMBL: a large-scale bioactivity database for drug discovery. Nucleic Acids Res 40(Database issue):D1100–D1107. https://doi.org/10.1093/nar/gkr777
Guo Y, **ao Z, Yang L, Gao Y, Zhu Q, Hu L et al (2020) Hypoxia-inducible factors in hepatocellular carcinoma (review). Oncol Rep 43(1):3–15. https://doi.org/10.3892/or.2019.7397
Haschek-Hock WM, Rousseaux CG, Wallig MA, Bolon B (2021) Haschek and Rousseaux’s Handbook of Toxicologic Pathology, Volume 1: principles and practice of toxicologic pathology. Academic press
Hu X, Pan H, Zhou S, Pang Q, Wang Y, Zhu C et al (2022) HS1BP3, transcriptionally regulated by ESR1, promotes hepatocellular carcinoma progression. Biochem Biophys Res Commun 623:111–119. https://doi.org/10.1016/j.bbrc.2022.07.047
Kumar N, Singh A, Sharma DK, Kishore K (2019) Toxicity of food additives. Food Safety Human Health:67–98. https://doi.org/10.1016/b978-0-12-816333-7.00003-5
Li X, Hu Z, Shi H, Wang C, Lei J, Cheng Y (2020) Inhibition of VEGFA increases the sensitivity of ovarian cancer cells to chemotherapy by suppressing VEGFA-mediated autophagy. Onco Targets Ther 13:8161–8171. https://doi.org/10.2147/OTT.S250392
Li YB, Zhang Y, Wang YM, Li YM, Yang FF, Zhang PJ, Liu CX (2019) A strategy for the discovery and validation of toxicity quality marker of Chinese medicine based on network toxicology. Phytomedicine 54:365–370. https://doi.org/10.1016/j.phymed.2018.01.018
More TA, Shaikh Z, Ali A (2021) Artificial sweeteners and their health implications: a review. Biosci Biotechnol Res Asia 18(2):227–237. https://doi.org/10.13005/bbra/2910
O'Brien MH, Pitot HC, Chung SH, Lambert PF, Drinkwater NR, Bilger A (2021) Estrogen receptor-alpha suppresses liver carcinogenesis and establishes sex-specific gene expression. Cancers (Basel) 13(10). https://doi.org/10.3390/cancers13102355
Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, Smyth GK (2015) Limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res 43(7):e47. https://doi.org/10.1093/nar/gkv007
Saeed, A., & Mustafa Khidir, A. (2020). Aspartame sweetener. https://doi.org/10.20959/wjpps20202-15435.
Shaher SAA, Mihailescu DF, Amuzescu B (2023) Aspartame safety as a food sweetener and related health hazards. Nutrients 15(16). https://doi.org/10.3390/nu15163627
Shaywitz BA, Sullivan CM, Anderson GM, Gillespie SM, Sullivan B, Shaywitz SE (1994) Aspartame, behavior, and cognitive function in children with attention deficit disorder. Pediatrics 93(1):70–75. https://doi.org/10.1542/peds.93.1.70
Shen H, Yu H, Li QY, Wei YT, Fu J, Dong H, Chen Y (2022) Hepatocyte-derived VEGFA accelerates the progression of non-alcoholic fatty liver disease to hepatocellular carcinoma via activating hepatic stellate cells. Acta Pharmacol Sin 43(11):2917–2928. https://doi.org/10.1038/s41401-022-00907-5
Shi M, Chen MS, Sekar K, Tan CK, Ooi LL, Hui KM (2014) A blood-based three-gene signature for the non-invasive detection of early human hepatocellular carcinoma. Eur J Cancer 50(5):928–936. https://doi.org/10.1016/j.ejca.2013.11.026
Soffritti M, Belpoggi F, Manservigi M, Tibaldi E, Lauriola M, Falcioni L, Bua L (2010) Aspartame administered in feed, beginning prenatally through life span, induces cancers of the liver and lung in male Swiss mice. Am J Ind Med 53(12):1197–1206. https://doi.org/10.1002/ajim.20896
Souche E, Beltran S, Brosens E, Belmont JW, Fossum M, Riess O, Weiss MM (2022) Recommendations for whole genome sequencing in diagnostics for rare diseases. Eur J Hum Genet 30(9):1017–1021. https://doi.org/10.1038/s41431-022-01113-x
Sukocheva OA (2018) Estrogen, estrogen receptors, and hepatocellular carcinoma: are we there yet? World J Gastroenterol 24(1):1–4. https://doi.org/10.3748/wjg.v24.i1.1
Szklarczyk D, Kirsch R, Koutrouli M, Nastou K, Mehryary F, Hachilif R, von Mering C (2023) The STRING database in 2023: protein-protein association networks and functional enrichment analyses for any sequenced genome of interest. Nucleic Acids Res 51(D1):D638–D646. https://doi.org/10.1093/nar/gkac1000
Szklarczyk D, Santos A, von Mering C, Jensen LJ, Bork P, Kuhn M (2016) STITCH 5: augmenting protein-chemical interaction networks with tissue and affinity data. Nucleic Acids Res 44(D1):D380–D384. https://doi.org/10.1093/nar/gkv1277
Tung EK, Mak CK, Fatima S, Lo RC, Zhao H, Zhang C, Ng IO (2011) Clinicopathological and prognostic significance of serum and tissue Dickkopf-1 levels in human hepatocellular carcinoma. Liver Int 31(10):1494–1504. https://doi.org/10.1111/j.1478-3231.2011.02597.x
UniProt C (2023) UniProt: the Universal Protein Knowledgebase in 2023. Nucleic Acids Res 51(D1):D523–D531. https://doi.org/10.1093/nar/gkac1052
Wu L, Zhang C, Long Y, Chen Q, Zhang W, Liu G (2022) Food additives: from functions to analytical methods. Crit Rev Food Sci Nutr 62(30):8497–8517. https://doi.org/10.1080/10408398.2021.1929823
Yin L, Cai Z, Zhu B, Xu C (2018) Identification of key pathways and genes in the dynamic progression of HCC based on WGCNA. Genes (Basel) 9(2). https://doi.org/10.3390/genes9020092
Yue L, Yang K, Jiang F, Dong S, Yang K, Zhu D (2023) Chemical profiling of principle active and toxic constituents in herbs containing aristolochic acids. Chin Herb Med. https://doi.org/10.1016/j.chmed.2023.02.008
Zang E, Jiang L, Cui H, Li X, Yan Y, Liu Q, Li M (2022) Only plant-based food additives: an overview on application, safety, and key challenges in the food industry. Food Rev Int 39(8):5132–5163. https://doi.org/10.1080/87559129.2022.2062764
Zhou Y, He M (2023) GSN synergies with actin-related transfer molecular chain to promote invasion and metastasis of HCC. Clin Transl Oncol 25(2):482–490. https://doi.org/10.1007/s12094-022-02961-1
Funding
This study was supported by the project of Science and Technology Achievement Transfer and Transformation Demonstration Project of Sichuan Provincial (2023ZHCG0071).
Author information
Authors and Affiliations
Contributions
BL proposed and designed the study. BL and LYS performed the analysis. ZQH and YKW contributed to the writing of the manuscript. MF supervised and validated the results. WCF and WTL prepared additional material. TL revised the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, B., Shi, L., Feng, M. et al. An Investigation of the Toxicity and Mechanisms of Food Additives Based on Network Toxicology and GEO Databases: A Case Study of Aspartame. Food Anal. Methods 17, 1057–1072 (2024). https://doi.org/10.1007/s12161-024-02634-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12161-024-02634-5