Abstract
Purpose
Long non-coding RNAs (lncRNAs) have participated in progression of colorectal cancer. This study aims to study the role of RUNX1/RNCR3/miR-1301-3p/AKT1 axis in colorectal cancer.
Methods
The cancer tissues were from patients with colorectal cancer. The qRT-PCR was used to determine expression of lncRNA RNCR3, miR-1301-3p, and AKT1. Both dual-luciferase reporter assay and ChIP assay were conducted to investigate the binding sites of RUNX1 on RNCR3 promoter. Western blot was performed to analyze expression of AKT1 protein. Both dual-luciferase reporter assay and RIP assay were performed to detect the interacting sites between RNCR3 and miR-1301-3p. The CCK-8 assay, soft agar assay, transwell assay, and annexin-V-FITC/PI staining were applied to analyze the cell growth, invasion, and apoptosis, respectively.
Results
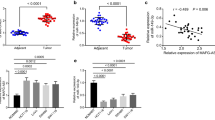
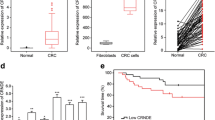
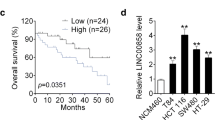
The data demonstrated that RNCR3 was elevated in colorectal cancer, and it was negatively correlated with expression of miR-1301-3p which was decreased in cancers. Then, RNCR3 could interact with and suppress miR-1301-3p expression in HCT116 and SW480. Knockdown of RNCR3 or miR-1301-3p overexpression significantly inhibited cell growth, invasion, and increased apoptosis through suppressing expression of Cyclin A1, PCNA, N-cadherin, Bcl-2, and promoting expression of E-cadherin, Bax in vitro and in vivo. RUNX1 was directly bound to RNCR3 promoter to activate RNCR3 expression. Furthermore, overexpression of RNCR3 blocked tumor inhibitory effects of miR-1301-3p on proliferation, colony formation, invasion, and apoptosis in vitro and in vivo. Additionally, RNCR3 and miR-1301-3p synergistically modulated AKT1 expression.
Conclusion
RUNX1-activated upregulation of RNCR3 promoted colorectal cancer progression by sponging miR-1301-3p to elevate AKT1 levels in vitro and in vivo.






Similar content being viewed by others
Abbreviations
- lncRNAs:
-
Long non-coding RNAs
- CRC:
-
Colorectal cancer
- ncRNAs:
-
Non-coding RNAs
- lncRNAs:
-
Long non-coding RNAs
- miRNAs:
-
MicroRNAs
- NBR2:
-
Neighbor of BRCA1 lncRNA 2
- TTN-AS1:
-
TTN antisense RNA 1
- KLF15:
-
Kruppel-like factor 15
- ROR1-AS1:
-
LncRNA ROR1 antisense RNA 1
- MIR17HG:
-
MiR-17-92a-1 cluster host gene
- GAS5:
-
Growth arrest specific 5
- SLCO4A1-AS1:
-
SLCO4A1 antisense RNA 1
- PARD3:
-
Par-3 family cell polarity regulator
- NEAT1:
-
Nuclear paraspeckle assembly transcript 1
- CDK6:
-
Cyclin-dependent kinase 6
- RNCR3:
-
LncRNA retinal non-coding RNA3
- KLF16:
-
Kruppel-like factor 16
- qRT-PCR:
-
Quantitative real-time polymerase chain reaction
- RUNX1:
-
RUNX family transcription factor 1
- AKT1:
-
AKT serine/threonine kinase 1
- ATCC:
-
American type culture collection
- DMEM:
-
Dulbecco's modified eagle medium
- siRNAs:
-
Small interfering RNAs
- shRNA:
-
Short hairpin RNA
- WT:
-
Wild type
- NC:
-
Negative controls
- FL:
-
Full length of wild type
- UTR:
-
Untranslated regions
- RIP:
-
RNA immunoprecipitation
- anti-Ago2:
-
Anti-Argonaute-2
- CCK-8:
-
Cell Counting Kit-8
- Annexin-V-FITC/PI staining:
-
Annexin-V/fluorescein isothiocyanate (FITC)/propidium iodide (PI) staining
- RIPA:
-
Radio immunoprecipitation assay
- SDS-PAGE:
-
Sodium dodecyl sulfate (SDS)-polyacrylamide gel electrophoresis
- PVDF:
-
Polyvinylidene difluoride
- PBS:
-
Phosphate-buffered saline
- HRP:
-
Horseradish peroxidase
- GAPDH:
-
Glyceraldehyde-3-phosphate dehydrogenase
- qRT-PCR:
-
Quantitative real-time polymerase chain reaction
- cDNA:
-
Complementary DNA
- ChIP:
-
Chromatin immunoprecipitation
- S.D.:
-
Standard deviation
- ANOVA:
-
One-way analysis of variance
- TCF3:
-
Transcription factor 3
- STAT3:
-
Signal transducer and activator of transcription 3
- LUCAT1:
-
Lung cancer-associated transcript 1
- SP1:
-
Sp1 transcription factor
- RUNX1:
-
Runt-related transcription factor 1
References
Siegel RL, Miller KD, Fedewa SA, Ahnen DJ, Meester RGS, Barzi A, Jemal A. Colorectal cancer statistics, 2017. CA Cancer J Clin. 2017;67(3):177–93.
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68(6):394–424.
Azvolinsky A. Colorectal cancer: to stack or sequence therapy? J Natl Cancer Inst. 2015;107(5):djv138.
Lander ES. Initial impact of the sequencing of the human genome. Nature. 2011;470(7333):187–97.
Kopp F, Mendell JT. Functional classification and experimental dissection of long noncoding RNAs. Cell. 2018;172(3):393–407.
Di W, Weinan X, **n L, Zhiwei Y, **nyue G, **xue T, Mingqi L. Long noncoding RNA SNHG14 facilitates colorectal cancer metastasis through targeting EZH2-regulated EPHA7. Cell Death Dis. 2019;10(7):514.
Tang J, Yan T, Bao Y, Shen C, Yu C, Zhu X, Tian X, Guo F, Liang Q, Liu Q, et al. LncRNA GLCC1 promotes colorectal carcinogenesis and glucose metabolism by stabilizing c-Myc. Nat Commun. 2019;10(1):3499.
Wang Y, Jiang F, **ong Y, Cheng X, Qiu Z, Song R. LncRNA TTN-AS1 sponges miR-376a-3p to promote colorectal cancer progression via upregulating KLF15. Life Sci. 2019;244(1):116936–44.
Marchese FP, Raimondi I, Huarte M. The multidimensional mechanisms of long noncoding RNA function. Genome Biol. 2017;18(1):206.
Prensner JR, Chinnaiyan AM. The emergence of lncRNAs in cancer biology. Cancer Discov. 2011;1(5):391–407.
Liang C, Zhao T, Li H, He F, Zhao X, Zhang Y, Chu X, Hua C, Qu Y, Duan Y, et al. Long Non-coding RNA ITIH4-AS1 accelerates the proliferation and metastasis of colorectal cancer by activating JAK/STAT3 signaling. Mol Therapy Nucleic Acids. 2019;18:183–93.
Bartonicek N, Maag JL, Dinger ME. Long noncoding RNAs in cancer: mechanisms of action and technological advancements. Mol Cancer. 2016;15(1):43.
Kallen AN, Zhou XB, Xu J, Qiao C, Ma J, Yan L, Lu L, Liu C, Yi JS, Zhang H, et al. The imprinted H19 lncRNA antagonizes let-7 microRNAs. Mol Cell. 2013;52(1):101–12.
Wang FZ, Zhang MQ, Zhang L, Zhang MC. Long non-coding RNA ROR1-AS1 enhances colorectal cancer metastasis by targeting miR-375. Eur Rev Med Pharmacol Sci. 2019;23(16):6899–905.
Xu J, Meng Q, Li X, Yang H, Xu J, Gao N, Sun H, Wu S, Familiari G, Relucenti M, et al. Long noncoding RNA MIR17HG promotes colorectal cancer progression via miR-17-5p. Cancer Res. 2019;79(19):4882–955.
Liu L, Wang HJ, Meng T, Lei C, Yang XH, Wang QS, ** B, Zhu JF. lncRNA GAS5 inhibits cell migration and invasion and promotes autophagy by targeting miR-222-3p via the GAS5/PTEN-signaling pathway in CRC. Mol Therapy Nucleic Acids. 2019;17:644–56.
Wang Z, ** J. LncRNA SLCO4A1-AS1 promotes colorectal cancer cell proliferation by enhancing autophagy via miR-508-3p/PARD3 axis. Aging (Albany NY). 2019;11(14):4876–89.
He Z, Dang J, Song A, Cui X, Ma Z, Zhang Z. NEAT1 promotes colon cancer progression through sponging miR-495-3p and activating CDK6 in vitro and in vivo. J Cell Physiol. 2019;234(11):19582–91.
Zhang L, Cao Y, Wei M, Jiang X, Jia D. Long noncoding RNA-RNCR3 overexpression deleteriously affects the growth of glioblastoma cells through miR-185-5p/Kruppel-like factor 16 axis. J Cell Biochem. 2018;119(11):9081–9.
Tian C, Deng Y, ** Y, Shi S, Bi H. Long non-coding RNA RNCR3 promotes prostate cancer progression through targeting miR-185-5p. Am J Transl Res. 2018;10(5):1562–70.
Wang L, Zhao Y, Xu M, Zhou F, Yan J. Serum miR-1301-3p, miR-335-5p, miR-28-5p, and their target B7–H3 may serve as novel biomarkers for colorectal cancer. J BUON. 2019;24(3):1120–7.
Fernando TR, Contreras JR, Zampini M, Rodriguez-Malave NI, Alberti MO, Anguiano J, Tran TM, Palanichamy JK, Gajeton J, Ung NM, et al. The lncRNA CASC15 regulates SOX4 expression in RUNX1-rearranged acute leukemia. Mol Cancer. 2017;16(1):126.
Chen X, Zeng K, Xu M, Hu X, Liu X, Xu T, He B, Pan Y, Sun H, Wang S. SP1-induced lncRNA-ZFAS1 contributes to colorectal cancer progression via the miR-150-5p/VEGFA axis. Cell Death Dis. 2018;9(10):982.
Lu Y, Hu Z, Mangala LS, Stine ZE, Hu X, Jiang D, **ang Y, Zhang Y, Pradeep S, Rodriguez-Aguayo C, et al. MYC targeted long noncoding RNA DANCR promotes cancer in part by reducing p21 levels. Cancer Res. 2018;78(1):64–74.
Liu HT, Liu S, Liu L, Ma RR, Gao P. EGR1-mediated transcription of lncRNA-HNF1A-AS1 promotes cell-cycle progression in gastric cancer. Cancer Res. 2018;78(20):5877–90.
Wang X, Guo S, Zhao R, Liu Y, Yang G. STAT3-activated long non-coding RNA lung cancer associated transcript 1 drives cell proliferation, migration, and invasion in hepatoblastoma through regulation of the miR-301b/STAT3 Axis. Hum Gene Ther. 2019;30(6):702–13.
Jiang L, Wang R, Fang L, Ge X, Chen L, Zhou M, Zhou Y, **ong W, Hu Y, Tang X, et al. HCP5 is a SMAD3-responsive long non-coding RNA that promotes lung adenocarcinoma metastasis via miR-203/SNAI axis. Theranostics. 2019;9(9):2460–74.
Zhang L, Liu SK, Song L, Yao HR. SP1-induced up-regulation of lncRNA LUCAT1 promotes proliferation, migration and invasion of cervical cancer by sponging miR-181a. Artif Cells Nanomed Biotechnol. 2019;47(1):556–64.
Zhao W, Li L. SP1-induced upregulation of long non-coding RNA HCP5 promotes the development of osteosarcoma. Pathol Res Pract. 2019;215(3):439–45.
Zheng L, Hu N, Zhou X. TCF3-activated LINC00152 exerts oncogenic role in osteosarcoma through regulating miR-1182/CDK14 axis. Pathol Res Pract. 2019;215(2):373–80.
Shen X, Zhong J, Yu P, Zhao Q, Huang T. YY1-regulated LINC00152 promotes triple negative breast cancer progression by affecting on stability of PTEN protein. Biochem Biophys Res Commun. 2019;509(2):448–54.
** J, Feng J, Zeng S, Huang P. Long noncoding RNA UFC1 is activated by E2F1 and exerts oncogenic properties by functioning as a ceRNA of FOXP3. Cancer Med. 2018;7(7):3301–10.
Cao Y, Lin M, Bu Y, Ling H, He Y, Huang C, Shen Y, Song B, Cao D. p53-inducible long non-coding RNA PICART1 mediates cancer cell proliferation and migration. Int J Oncol. 2017;50(5):1671–82.
Li Q, Lai Q, He C, Fang Y, Yan Q, Zhang Y, Wang X, Gu C, Wang Y, Ye L, et al. RUNX1 promotes tumour metastasis by activating the Wnt/beta-catenin signalling pathway and EMT in colorectal cancer. J Exp Clin Cancer Res. 2019;38(1):334.
Li J, Zhao LM, Zhang C, Li M, Gao B, Hu XH, Cao J, Wang GY. The lncRNA FEZF1-AS1 promotes the progression of colorectal cancer through regulating OTX1 and targeting miR-30a-5p. Oncol Res. 2020;28(1):51–63.
Wen J, Wang H, Dong T, Gan P, Fang H, Wu S, Li J, Zhang Y, Du R, Zhu Q. STAT3-induced upregulation of lncRNA ABHD11-AS1 promotes tumour progression in papillary thyroid carcinoma by regulating miR-1301-3p/STAT3 axis and PI3K/AKT signalling pathway. Cell Prolif. 2019;52(2):e12569.
Peng X, Yan B, Shen Y. MiR-1301-3p inhibits human breast cancer cell proliferation by regulating cell cycle progression and apoptosis through directly targeting ICT1. Breast Cancer. 2018;25(6):742–52.
Zhi T, Jiang K, Zhang C, Xu X, Wu W, Nie E, Yu T, Zhou X, Bao Z, ** X, et al. MicroRNA-1301 inhibits proliferation of human glioma cells by directly targeting N-Ras. Am J Cancer Res. 2017;7(4):982–98.
Behan FM, Iorio F, Picco G, Goncalves E, Beaver CM, Migliardi G, Santos R, Rao Y, Sassi F, Pinnelli M, et al. Prioritization of cancer therapeutic targets using CRISPR-Cas9 screens. Nature. 2019;568(7753):511–6.
Wang LL, Zhang L, Cui XF. Downregulation of long noncoding RNA LINC01419 inhibits cell migration, invasion, and tumor growth and promotes autophagy via inactivation of the PI3K/Akt1/mTOR pathway in gastric cancer. Ther Adv Med Oncol. 2019;11:1758835919874651.
Acknowledgements
We thank our team for technical cooperation.
Funding
This study received no financial supporting from government subjects.
Author information
Authors and Affiliations
Contributions
GX designed the project. H-nW, D-wY, JY, LM, KL, YZ, C-xD, and KZ performed the animal experiments, and they cultured the cell lines and conducted the cell experiments. GX, H-nW, and D-wY analyzed the experimental data. GX, H-nW, and D-w wrote the manuscript, and GX offered discussion and suggestions. GXX provided the financial support.
Corresponding author
Ethics declarations
Conflict of interest
All authors declare no competing financial interests.
Ethical approval
All applicable international, national, and/or institutional guidelines for the care and use of animals were followed. All procedures performed in our studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards.
Informed consent
Informed consent was obtained from all included participants.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Xu, G., Wang, H., Yuan, D. et al. RUNX1-activated upregulation of lncRNA RNCR3 promotes cell proliferation, invasion, and suppresses apoptosis in colorectal cancer via miR-1301-3p/AKT1 axis in vitro and in vivo. Clin Transl Oncol 22, 1762–1777 (2020). https://doi.org/10.1007/s12094-020-02335-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12094-020-02335-5