Abstract
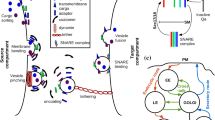
Eukaryotic cells use small membrane-enclosed vesicles to transport molecular cargo between intracellular compartments. Interactions between molecules on vesicles and compartments determine the source and target compartment of each vesicle type. The set of compartment and vesicle types in a cell define the nodes and edges of a transport graph known as the vesicle traffic network. The transmembrane SNARE proteins that regulate vesicle fusion to target compartments travel in cycles through the transport graph, but the paths they follow must be tightly regulated to avoid aberrant vesicle fusion. Here we use graph-theoretic ideas to understand how such molecular constraints place constraints on the structure of the transport graph. We identify edge connectivity (the minimum number of edges that must be removed to disconnect a graph) as a key determinant that separates allowed and disallowed types of transport graphs. As we increase the flexibility of molecular regulation, the required edge connectivity decreases, so more types of vesicle transport graphs are allowed. These results can be used to aid the discovery of new modes of molecular regulation and new vesicle traffic pathways.



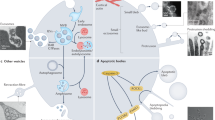
Similar content being viewed by others
References
Baker RW, Jeffrey PD, Zick M, Phillips BP, Wickner WT, and Hughson FM 2015 A direct role for the Sec1/Munc18-family protein Vps33 as a template for SNARE assembly. Science 349 1111–1114
Bang-Jensen J and Gutin GZ 2008 Digraphs: theory, algorithms and applications (Springer)
Bhattacharyya A, Gupta A, Kuppusamy L, Mani S, Shukla A, Srivas M and Thattai M 2019 A formal methods approach to predicting new features of the eukaryotic vesicle traffic system. Acta Inform. 58 1–37
Bonifacino JS and Glick BS 2004 The mechanisms of vesicle budding and fusion. Cell 116 153–166
Cai H, Reinisch K, and Ferro-Novick S 2007 Coats, tethers, Rabs, and SNAREs work together to mediate the intracellular destination of a transport vesicle. Dev. Cell 12 671–682
Cockcroft S and Raghu P 2018 Phospholipid transport protein function at organelle contact sites. Curr. Opin. Cell Biol. 53 52–60
Conibear E and Davis NG 2010 Palmitoylation and depalmitoylation dynamics at a glance. J. Cell Sci. 123 4007–4010
Dubuke ML and Munson M 2016 The secret life of tethers: the role of tethering factors in SNARE complex regulation. Front. Cell Dev. Biol. 4 42
Greaves J and Chamberlain LH 2011 Differential palmitoylation regulates intracellular patterning of SNAP25. J. Cell Sci. 124 1351–1360
Jahn R and Scheller RH 2006 SNAREs-engines for membrane fusion. Nat. Rev. Mol. Cell Biol. 7 631–643
Kamena F and Spang A 2004 Tip20p prohibits back-fusion of COPII vesicles with the endoplasmic reticulum. Science 304 286–289
Karnahl M, Park M, Mayer U, Hiller U, and Jürgens G 2017 ER assembly of SNARE complexes mediating formation of partitioning membrane in Arabidopsis cytokinesis. eLife 6 e25327
Kent HM, Evans PR, Schäfer IB, Gray SR, Sanderson CM, Luzio JP, Peden AA, and Owen DJ 2012 Structural basis of the intracellular sorting of the SNARE VAMP7 by the AP3 adaptor complex. Dev. Cell 22 979–988
Mani S and Thattai M 2016a Stacking the odds for Golgi cisternal maturation. eLife 5 e16231
Mani S and Thattai M 2016b Wine glasses and hourglasses: Non-adaptive complexity of vesicle traffic in microbial eukaryotes. Mol. Biochem. Parasitol. 209 58–63
Pryor PR, Jackson L, Gray SR, Edeling MA, Thompson A, Sanderson CM, Evans PR, Owen DJ, and Luzio JP 2008 Molecular basis for the sorting of the SNARE VAMP7 into endocytic clathrin-coated vesicles by the ArfGAP Hrb. Cell 134 817–827
Ramadas R and Thattai M 2013 New organelles by gene duplication in a biophysical model of eukaryote endomembrane evolution. Biophys. J. 104 2553–2563
Raposo G and Stoorvogel W 2013 Extracellular vesicles: Exosomes, microvesicles, and friends. J. Cell Biol. 200 373–383
Ratamero EM and Royle SJ 2019 Calculating the maximum capacity of intracellular transport vesicles. bioRxiv. https://doi.org/10.1101/555813
Schäfer IB, Hesketh GG, Bright NA, Gray SR, Pryor PR, Evans PR, Luzio JP, and Owen DJ 2012 The binding of Varp to VAMP7 traps VAMP7 in a closed, fusogenically inactive conformation. Nat. Struct. Mol. Biol. 19 1300–1309
Schwartz BL 1961 An analytic method for the “difficult crossing” puzzles. Math. Mag. 34 187
Shukla A, Bhattacharyya A, Kuppusamy L, Srivas M, and Thattai M 2017 Discovering vesicle traffic network constraints by model checking. PloS One 12 e0180692
Stenmark H 2009 Rab GTPases as coordinators of vesicle traffic. Nat. Rev. Mol. Cell Biol. 10 513–525
Südhof TC and Rothman JE 2009 Membrane fusion: grappling with SNARE and SM proteins. Science 323 474–477
Sztul E, Chen P-W, Casanova JE, et al. 2019 ARF GTPases and their GEFs and GAPs: concepts and challenges. Mol. Biol. Cell 30 1249–1271
Van Den Bogaart G, Holt MG, Bunt G, Riedel D, Wouters FS, and Jahn R 2010 One SNARE complex is sufficient for membrane fusion. Nat. Struct. Mol. Biol. 17 358–364
Acknowledgements
We thank Arnab Bhattacharyya for pointing out the importance of edge connectivity to the vesicle traffic problem. We thank Gregory Gutin, Anjali Jaiman, Ramya Purkanti and Ankit Shukla for useful discussions. SM acknowledges funding from the Institute for Basic Science, South Korea, Project Code IBS-R020-D1. MT acknowledges funding from the Simons Foundation (287975) which supports collaborative activities of the Simons Centre for the Study of Living Machines.
Author information
Authors and Affiliations
Corresponding author
Additional information
Corresponding editor: Mohit Kumar Jolly.
Corresponding editor: Mohit Kumar Jolly
This article is part of the Topical Collection: Emergent dynamics of biological networks.
Rights and permissions
About this article
Cite this article
Mani, S., Krishnan, K. & Thattai, M. Graph-theoretic constraints on vesicle traffic networks. J Biosci 47, 11 (2022). https://doi.org/10.1007/s12038-021-00252-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12038-021-00252-5