Abstract
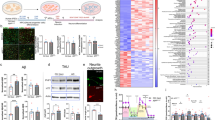
Alzheimer’s disease research has been conducted for many years, yet no effective cure methods have been found. N6-methyladenosine (m6A) RNA methylation, an essential post-transcriptional regulation mechanism, has been discovered to affect essential neurobiological processes, such as brain cell development and aging, which are closely related to neurodegenerative diseases such as Alzheimer’s disease. The relationship between Alzheimer’s disease and the m6A mechanism still needs further investigation. Our work evaluated the alteration profile of m6A regulators and their influences on Alzheimer’s disease in 4 brain regions: the postcentral gyrus, superior frontal gyrus, hippocampus, and entorhinal cortex. We found that the expression levels of the m6A regulators FTO, ELAVL1, and YTHDF2 were altered in Alzheimer’s disease and were related to pathological development and cognitive levels. We also assessed AD-related biological processes influenced by m6A regulators via GSEA and GSVA method. Biological Processes Gene Ontology terms including memory, cognition, and synapse-signaling were found to potentially be affected by m6A regulators in AD. We also found different m6A modification patterns in AD samples among different brain regions, mainly due to differences in m6A readers. Finally, we further evaluated the importance of AD-related regulators based on the WGCNA method, assessed their potential targets based on correlation relationships, and constructed diagnostic models in 3 of all 4 regions using hub regulators, including FTO, YTHDC1, YTHDC2, etc., and their potential targets. This work aims to provide a reference for the follow-up study of m6A and Alzheimer’s disease.








Similar content being viewed by others
Code Availability
Not Applicable
Data Availability
Not Applicable
References
Alzheimer A, Stelzmann RA, Schnitzlein HN, Murtagh FR (1995) An English translation of Alzheimer’s 1907 paper, “Uber eine eigenartige Erkankung der Hirnrinde”. Clin Anat 8 (6):429–431. https://doi.org/10.1002/ca.980080612
Takizawa C, Thompson PL, van Walsem A, Faure C, Maier WC (2015) Epidemiological and economic burden of Alzheimer's disease: a systematic literature review of data across Europe and the United States of America. J Alzheimers Dis 43(4):1271–1284. https://doi.org/10.3233/JAD-141134
McKhann G, Drachman D, Folstein M, Katzman R, Price D, Stadlan EM (1984) Clinical diagnosis of Alzheimer's disease: report of the NINCDS-ADRDA Work Group under the auspices of Department of Health and Human Services Task Force on Alzheimer's Disease. Neurology 34(7):939–944. https://doi.org/10.1212/wnl.34.7.939
McKhann GM, Knopman DS, Chertkow H, Hyman BT, Jack CR Jr, Kawas CH, Klunk WE, Koroshetz WJ et al (2011) The diagnosis of dementia due to Alzheimer's disease: recommendations from the National Institute on Aging-Alzheimer's Association workgroups on diagnostic guidelines for Alzheimer's disease. Alzheimers Dement 7(3):263–269. https://doi.org/10.1016/j.jalz.2011.03.005
Montine TJ, Phelps CH, Beach TG, Bigio EH, Cairns NJ, Dickson DW, Duyckaerts C, Frosch MP et al (2012) National Institute on Aging-Alzheimer's Association guidelines for the neuropathologic assessment of Alzheimer's disease: a practical approach. Acta Neuropathol 123(1):1–11. https://doi.org/10.1007/s00401-011-0910-3
Cummings J, Lee G, Zhong K, Fonseca J, Taghva K (2021) Alzheimer's disease drug development pipeline: 2021. Alzheimers Dement (N Y) 7(1):e12179. https://doi.org/10.1002/trc2.12179
Sevigny J, Chiao P, Bussiere T, Weinreb PH, Williams L, Maier M, Dunstan R, Salloway S et al (2016) The antibody aducanumab reduces Abeta plaques in Alzheimer's disease. Nature 537(7618):50–56. https://doi.org/10.1038/nature19323
Ostrowitzki S, Lasser RA, Dorflinger E, Scheltens P, Barkhof F, Nikolcheva T, Ashford E, Retout S et al (2017) A phase III randomized trial of gantenerumab in prodromal Alzheimer's disease. Alzheimers Res Ther 9(1):95. https://doi.org/10.1186/s13195-017-0318-y
Egan MF, Kost J, Voss T, Mukai Y, Aisen PS, Cummings JL, Tariot PN, Vellas B et al (2019) Randomized trial of verubecestat for prodromal Alzheimer's disease. N Engl J Med 380(15):1408–1420. https://doi.org/10.1056/NEJMoa1812840
Wang X, Zhao BS, Roundtree IA, Lu Z, Han D, Ma H, Weng X, Chen K et al (2015) N(6)-methyladenosine modulates messenger RNA translation efficiency. Cell 161(6):1388–1399. https://doi.org/10.1016/j.cell.2015.05.014
Wang X, Lu Z, Gomez A, Hon GC, Yue Y, Han D, Fu Y, Parisien M, Dai Q et al (2014) N6-methyladenosine-dependent regulation of messenger RNA stability. Nature 505(7481):117–120. https://doi.org/10.1038/nature12730
Dominissini D, Moshitch-Moshkovitz S, Schwartz S, Salmon-Divon M, Ungar L, Osenberg S, Cesarkas K, Jacob-Hirsch J et al (2012) Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature 485(7397):201–206. https://doi.org/10.1038/nature11112
Zheng G, Dahl JA, Niu Y, Fedorcsak P, Huang CM, Li CJ, Vagbo CB et al (2013) ALKBH5 is a mammalian RNA demethylase that impacts RNA metabolism and mouse fertility. Mol Cell 49(1):18–29. https://doi.org/10.1016/j.molcel.2012.10.015
Widagdo J, Zhao QY, Kempen MJ, Tan MC, Ratnu VS, Wei W, Leighton L, Spadaro PA et al (2016) Experience-dependent accumulation of N6-methyladenosine in the prefrontal cortex is associated with memory processes in mice. J Neurosci 36(25):6771–6777. https://doi.org/10.1523/JNEUROSCI.4053-15.2016
Merkurjev D, Hong WT, Iida K, Oomoto I, Goldie BJ, Yamaguti H, Ohara T, Kawaguchi SY et al (2018) Synaptic N(6)-methyladenosine (m(6)A) epitranscriptome reveals functional partitioning of localized transcripts. Nat Neurosci 21(7):1004–1014. https://doi.org/10.1038/s41593-018-0173-6
Walters BJ, Mercaldo V, Gillon CJ, Yip M, Neve RL, Boyce FM, Frankland PW, Josselyn SA (2017) The role of the RNA demethylase FTO (fat mass and obesity-associated) and mRNA methylation in hippocampal memory formation. Neuropsychopharmacology 42(7):1502–1510. https://doi.org/10.1038/npp.2017.31
Li Q, Li X, Tang H, Jiang B, Dou Y, Gorospe M, Wang W (2017) NSUN2-mediated m5C methylation and METTL3/METTL14-mediated m6A methylation cooperatively enhance p21 translation. J Cell Biochem 118(9):2587–2598. https://doi.org/10.1002/jcb.25957
Lewinska A, Adamczyk-Grochala J, Deregowska A, Wnuk M (2017) Sulforaphane-induced cell cycle arrest and senescence are accompanied by DNA hypomethylation and changes in microRNA profile in breast cancer cells. Theranostics 7(14):3461–3477. https://doi.org/10.7150/thno.20657
Min KW, Zealy RW, Davila S, Fomin M, Cummings JC, Makowsky D, McDowell CH et al (2018) Profiling of m6A RNA modifications identified an age-associated regulation of AGO2 mRNA stability. Aging Cell 17(3):e12753. https://doi.org/10.1111/acel.12753
Han M, Liu Z, Xu Y, Liu X, Wang D, Li F, Wang Y, Bi J (2020) Abnormality of m6A mRNA methylation is involved in Alzheimer's disease. Front Neurosci 14:98. https://doi.org/10.3389/fnins.2020.00098
Huang H, Camats-Perna J, Medeiros R, Anggono V, Widagdo J (2020) Altered expression of the m6A methyltransferase METTL3 in Alzheimer's disease. eNeuro 7(5). https://doi.org/10.1523/ENEURO.0125-20.2020
Li H, Ren Y, Mao K, Hua F, Yang Y, Wei N, Yue C, Li D, Zhang H (2018) FTO is involved in Alzheimer's disease by targeting TSC1-mTOR-Tau signaling. Biochem Biophys Res Commun 498(1):234–239. https://doi.org/10.1016/j.bbrc.2018.02.201
Reitz C, Tosto G, Mayeux R, Luchsinger JA, Group N-LNFS, Alzheimer's Disease Neuroimaging I (2012) Genetic variants in the fat and obesity associated (FTO) gene and risk of Alzheimer's disease. PLoS One 7(12):e50354. https://doi.org/10.1371/journal.pone.0050354
Wang Y, Wang J, Gao L, Stamm S, Andreadis A (2011) An SRp75/hnRNPG complex interacting with hnRNPE2 regulates the 5' splice site of tau exon 10, whose misregulation causes frontotemporal dementia. Gene 485(2):130–138. https://doi.org/10.1016/j.gene.2011.06.020
Shafik AM, Zhang F, Guo Z, Dai Q, Pajdzik K, Li Y, Kang Y, Yao B et al (2021) N6-methyladenosine dynamics in neurodevelopment and aging, and its potential role in Alzheimer's disease. Genome Biol 22(1):17. https://doi.org/10.1186/s13059-020-02249-z
Davis S, Meltzer PS (2007) GEOquery: a bridge between the Gene Expression Omnibus (GEO) and BioConductor. Bioinformatics 23(14):1846–1847. https://doi.org/10.1093/bioinformatics/btm254
Jiang X, Liu B, Nie Z, Duan L, **ong Q, ** Z, Yang C, Chen Y (2021) The role of m6A modification in the biological functions and diseases. Signal Transduct Target Ther 6(1):74. https://doi.org/10.1038/s41392-020-00450-x
Xu K, Cai Y, Zhang M, Zou H, Chang Z, Li D, Bai J, Xu J, Li Y (2021) Pan-cancer characterization of expression and clinical relevance of m(6)A-related tissue-elevated long non-coding RNAs. Mol Cancer 20(1):31. https://doi.org/10.1186/s12943-021-01324-8
Li Y, **ao J, Bai J, Tian Y, Qu Y, Chen X, Wang Q et al (2019) Molecular characterization and clinical relevance of m(6)A regulators across 33 cancer types. Mol Cancer 18(1):137. https://doi.org/10.1186/s12943-019-1066-3
Zhang Y, Xu S, Xu G, Gao Y, Li S, Zhang K, Tian Z, Guo J et al (2020) Dynamic expression of m(6)A regulators during multiple human tissue development and cancers. Front Cell Dev Biol 8:629030. https://doi.org/10.3389/fcell.2020.629030
Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, Smyth GK (2015) limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res 43(7):e47. https://doi.org/10.1093/nar/gkv007
Braak H, Braak E (1991) Neuropathological stageing of Alzheimer-related changes. Acta Neuropathol 82(4):239–259. https://doi.org/10.1007/BF00308809
Folstein MF, Folstein SE, McHugh PR (1975) "Mini-mental state". A practical method for grading the cognitive state of patients for the clinician. J Psychiatr Res 12(3):189–198. https://doi.org/10.1016/0022-3956(75)90026-6
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, Paulovich A et al (2005) Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci U S A 102(43):15545–15550. https://doi.org/10.1073/pnas.0506580102
Wu T, Hu E, Xu S, Chen M, Guo P, Dai Z, Feng T, Zhou L et al (2021) clusterProfiler 4.0: a universal enrichment tool for interpreting omics data. Innovation (Camb) 2(3):100141. https://doi.org/10.1016/j.xinn.2021.100141
Hanzelmann S, Castelo R, Guinney J (2013) GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinformatics 14:7. https://doi.org/10.1186/1471-2105-14-7
Wilkerson MD, Hayes DN (2010) ConsensusClusterPlus: a class discovery tool with confidence assessments and item tracking. Bioinformatics 26(12):1572–1573. https://doi.org/10.1093/bioinformatics/btq170
Langfelder P, Horvath S (2008) WGCNA: an R package for weighted correlation network analysis. BMC Bioinformatics 9:559. https://doi.org/10.1186/1471-2105-9-559
Zhang B, Horvath S (2005) A general framework for weighted gene co-expression network analysis. Stat Appl Genet Mol Biol 4:17. https://doi.org/10.2202/1544-6115.1128
Deng S, Zhang H, Zhu K, Li X, Ye Y, Li R, Liu X, Lin D et al (2021) M6A2Target: a comprehensive database for targets of m6A writers, erasers and readers. Brief Bioinform 22:(3). https://doi.org/10.1093/bib/bbaa055
Liu J, Li K, Cai J, Zhang M, Zhang X, **ong X, Meng H, Xu X et al (2020) Landscape and regulation of m(6)A and m(6)Am methylome across human and mouse tissues. Mol Cell 77(2):426–440 e426. https://doi.org/10.1016/j.molcel.2019.09.032
Tibshirani R (1996) Regression shrinkage and selection via the lasso. doi:https://doi.org/10.1111/j.2517-6161.1996.tb02080.x
Ullrich S, Munch A, Neumann S, Kremmer E, Tatzelt J, Lichtenthaler SF (2010) The novel membrane protein TMEM59 modulates complex glycosylation, cell surface expression, and secretion of the amyloid precursor protein. J Biol Chem 285(27):20664–20674. https://doi.org/10.1074/jbc.M109.055608
Kern F, Sarg B, Stasyk T, Hess D, Lindner H (2012) The Nogo receptor 2 is a novel substrate of Fbs1. Biochem Biophys Res Commun 417(3):977–981. https://doi.org/10.1016/j.bbrc.2011.12.050
Kurlawala Z, Shah PP, Shah C, Beverly LJ (2017) The STI and UBA domains of UBQLN1 are critical determinants of substrate interaction and proteostasis. J Cell Biochem 118(8):2261–2270. https://doi.org/10.1002/jcb.25880
Younas N, Zafar S, Shafiq M, Noor A, Siegert A, Arora AS, Galkin A, Zafar A et al (2020) SFPQ and Tau: critical factors contributing to rapid progression of Alzheimer's disease. Acta Neuropathol 140(3):317–339. https://doi.org/10.1007/s00401-020-02178-y
Berchtold NC, Coleman PD, Cribbs DH, Rogers J, Gillen DL, Cotman CW (2013) Synaptic genes are extensively downregulated across multiple brain regions in normal human aging and Alzheimer's disease. Neurobiol Aging 34(6):1653–1661. https://doi.org/10.1016/j.neurobiolaging.2012.11.024
Chen J, Zhang J, Gao Y, Li Y, Feng C, Song C, Ning Z, Zhou X et al (2021) LncSEA: a platform for long non-coding RNA related sets and enrichment analysis. Nucleic Acids Res 49(D1):D969–D980. https://doi.org/10.1093/nar/gkaa806
Feng C, Song C, Liu Y, Qian F, Gao Y, Ning Z, Wang Q, Jiang Y et al (2020) KnockTF: a comprehensive human gene expression profile database with knockdown/knockout of transcription factors. Nucleic Acids Res 48(D1):D93–D100. https://doi.org/10.1093/nar/gkz881
Jiang L, Lin W, Zhang C, Ash PEA, Verma M, Kwan J, van Vliet E, Yang Z et al (2021) Interaction of tau with HNRNPA2B1 and N(6)-methyladenosine RNA mediates the progression of tauopathy. Mol Cell 81(20):4209–4227 e4212. https://doi.org/10.1016/j.molcel.2021.07.038
Shi H, Zhang X, Weng YL, Lu Z, Liu Y, Lu Z, Li J, Hao P et al (2018) m(6)A facilitates hippocampus-dependent learning and memory through YTHDF1. Nature 563(7730):249–253. https://doi.org/10.1038/s41586-018-0666-1
Li L, Zang L, Zhang F, Chen J, Shen H, Shu L, Liang F, Feng C et al (2017) Fat mass and obesity-associated (FTO) protein regulates adult neurogenesis. Hum Mol Genet 26(13):2398–2411. https://doi.org/10.1093/hmg/ddx128
Li M, Zhao X, Wang W, Shi H, Pan Q, Lu Z, Perez SP, Suganthan R et al (2018) Ythdf2-mediated m(6)A mRNA clearance modulates neural development in mice. Genome Biol 19(1):69. https://doi.org/10.1186/s13059-018-1436-y
Robertson AG, Kim J, Al-Ahmadie H, Bellmunt J, Guo G, Cherniack AD, Hinoue T, Laird PW et al (2018) Comprehensive molecular characterization of muscle-invasive bladder cancer. Cell 174(4):1033. https://doi.org/10.1016/j.cell.2018.07.036
Bailey P, Chang DK, Nones K, Johns AL, Patch AM, Gingras MC, Miller DK, Christ AN et al (2016) Genomic analyses identify molecular subtypes of pancreatic cancer. Nature 531(7592):47–52. https://doi.org/10.1038/nature16965
Haass C, Kaether C, Thinakaran G, Sisodia S (2012) Trafficking and proteolytic processing of APP. Cold Spring Harb Perspect Med 2(5):a006270. https://doi.org/10.1101/cshperspect.a006270
Serrano-Pozo A, Das S, Hyman BT (2021) APOE and Alzheimer's disease: advances in genetics, pathophysiology, and therapeutic approaches. Lancet Neurol 20(1):68–80. https://doi.org/10.1016/S1474-4422(20)30412-9
Funding
This work was supported in part by grants from the National Natural Science Foundation of China (81701078), the Natural Science Foundation of Heilongjiang Province of China (Outstanding Youth Foundation,YQ2022H003), the University Nursing Program for Young Scholars with Creative Talents in Heilongjiang Province (UNPYSCT-2016190), China Postdoctoral Science Foundation (2016M600261, 2018T110317), Heilongjiang Postdoctoral Financial Assistance (LBH-Z15163), the Innovative Science Research Project of Harbin Medical University (2016JCZX37), and Heilongjiang Touyan Innovation Team Program.
Author information
Authors and Affiliations
Contributions
All authors contributed to the article. YongDa Wang and Xu Gao had the idea for the article. The first draft of the manuscript was written by Qing **a and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics Approval
Not applicable
Consent to Participate
Not applicable
Consent for Publication
Not applicable
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information

ESM 1
Supplementary Fig. 1 Threshold picking for WGCNA network construction among four brain regions. (A) R^2 and mean connectivity for different beta of the WCGNA network in the PG region, (B) the SFG region, (C) the HP region, (D) and the EC region. (PNG 255 kb)

ESM 2
Supplementary Fig. 2 Biological processes related to AD. (A) Venn graph of biological processes differences in activities among four brain regions. (B) GSEA results of significant GO: BP terms in all 4 brain regions. (C) Relationships between key biological processes and AD-related regulators in the PG region, (D) the SFG region, (E) the HP region, (F) and the EC region. (PNG 742 kb)

ESM 3
Supplementary Fig. 3 Identification of different modification patterns in AD samples among four brain regions. (A) Heatmaps of consensus clustering results in the PG region, (B) the SFG region, (C) the HP region, (D) and the EC region. (PNG 101 kb)

ESM 4
Supplementary Fig.4 Relationships of key regulators with modules and phenotypes. (A) Gene significance of regulators to phenotypes (GS > 0.2 shown) in the PG region, (B) the SFG region, (C) the HP region, (D) and the EC region. (E) Module membership of key regulators to modules (|MM| > 0.65 shown) in the PG region, (F) the SFG region, (G) the HP region, (H) and the EC region. (PNG 297 kb)

ESM 5
Supplementary Fig. 5 Hub genes potentially regulated by m6A regulators in the SFG region and the HP region. (A) Results of the assessment of regulatory evidence between FTO, (B) RBM15B and target genes in the SFG region and (C) YTHDC1 in the EC region. The bars represent (from right to left) correlation with the corresponding regulator, gene name, evidence of change after perturbation of regulator, evidence of interaction between regulator and gene transcript, and evidence of m6A or m6Am modification on gene transcript. Genes that show evidence of changes after regulator perturbation and interaction with the corresponding regulator are shown in the SFG region. Genes that show evidence of changes after regulator perturbation are shown in the HP region. Colors represent the relation between target genes and AD. (PNG 628 kb)

ESM 6
Supplementary Fig. 6 Hub genes potentially regulated by m6A regulators in the EC and HP regions. (A) Results of the assessment of regulatory evidence between the target gene YTHDC2 in the EC region and (B) YTHDC1 in the HP region. The bars represent (from right to left) correlation with the corresponding regulator, gene name, evidence of change after perturbation of regulator, evidence of interaction between regulator and gene transcript, and evidence of m6A or m6Am modification on gene transcript. Genes that show evidence of changes after regulator perturbation and interaction with the corresponding regulator are shown in the EC region. Genes that show evidence of changes after regulator perturbation are shown in the HP region. Colors represent relations between target genes and AD. (PNG 1592 kb)

ESM 7
Supplementary Fig. 7 Hub genes potentially regulated by m6A regulators in the HP region. (A) Results of the assessment of regulatory evidence between the target genes FTO and (B) YTHDF2 in the HP region. The bars represent (from right to left) correlation with the corresponding regulator, gene name, evidence of change after perturbation of regulator, evidence of interaction between regulator and gene transcript, and evidence of m6A or m6Am modification on gene transcript. Genes that show evidence of changes after regulator perturbation are shown in the HP region. Colors represent the relation between target genes and AD. (PNG 2849 kb)
ESM 8
(XLSX 10 kb)
ESM 9
(XLSX 20 kb)
ESM 10
(XLSX 25 kb)
ESM 11
(XLSX 52 kb)
ESM 12
(XLSX 344 kb)
ESM 13
(XLSX 81 kb)
ESM 14
(XLSX 12 kb)
ESM 15
(XLSX 32 kb)
ESM 16
(XLSX 12 kb)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, Z., **a, Q., Zhao, X. et al. The Landscape of m6A Regulators in Multiple Brain Regions of Alzheimer’s Disease. Mol Neurobiol 60, 5184–5198 (2023). https://doi.org/10.1007/s12035-023-03409-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-023-03409-5