Abstract
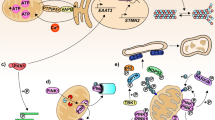
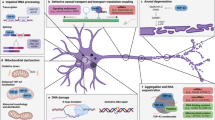
Amyotrophic lateral sclerosis (ALS) represents a rapidly progressing neurodegenerative disease and is characterized by a degeneration of motor neurons. Motor neurons are particularly susceptible to selective and early degeneration because of their extended axon length and their dependency on the cytoskeleton for its stability, signaling, and axonal transport. The motor neuron cytoskeleton comprises actin filaments, neurofilaments like peripherin, and microtubules. The Transactivating Response Region (TAR) DNA Binding Protein (TDP-43) forms characteristic cytoplasmic aggregates in motor neurons of ALS patients, and at least in part, the pathogenesis of ALS seems to be driven by toxic pTDP-43 aggregates in cytoplasm, which lead to a diminished axon formation and reduced axon length. Diminished axon formation and reduced axon length suggest an interaction of TDP-43 with the cytoskeleton of motor neurons. TDP-43 interacts with several cytoskeletal components, e.g., the microtubule-associated protein 1B (MAP1B) or the neurofilament light chain (NFL) through direct binding to its RNA. From a clinical perspective, cytoskeletal biomarkers like phosphorylated neurofilament heavy chain (pNFH) and NFL are already clinically used in ALS patients to predict survival, disease progression, and duration. Thus, in this review, we focus on the interaction of TDP-43 with the different cytoskeleton components such as actin filaments, neurofilaments, and microtubules as well as their associated proteins as one aspect in the complex pathogenesis of ALS.

Similar content being viewed by others
References
Morgan S, Orrell RW (2016) Pathogenesis of amyotrophic lateral sclerosis. Br Med Bull. doi:10.1093/bmb/ldw026
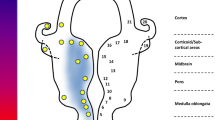
Brettschneider J, Arai K, Del Tredici K, Toledo JB, Robinson JL, Lee EB, Kuwabara S, Shibuya K et al (2014) TDP-43 pathology and neuronal loss in amyotrophic lateral sclerosis spinal cord. Acta Neuropathol 128(3):423–437. doi:10.1007/s00401-014-1299-6
Feiler MS, Strobel B, Freischmidt A, Helferich AM, Kappel J, Brewer BM, Li D, Thal DR et al (2015) TDP-43 is intercellularly transmitted across axon terminals. J Cell Biol 211(4):897–911. doi:10.1083/jcb.201504057
Spiller KJ, Cheung CJ, Restrepo CR, Kwong LK, Stieber AM, Trojanowski JQ, Lee VM (2016) Selective motor neuron resistance and recovery in a new inducible mouse model of TDP-43 proteinopathy. J Neurosci 36(29):7707–7717. doi:10.1523/JNEUROSCI.1457-16.2016
Gershoni-Emek N, Chein M, Gluska S, Perlson E (2015) Amyotrophic lateral sclerosis as a spatiotemporal mislocalization disease: location, location, location. Int Rev Cell Mol Biol 315:23–71. doi:10.1016/bs.ircmb.2014.11.003
Baldwin KR, Godena VK, Hewitt VL, Whitworth AJ (2016) Axonal transport defects are a common phenotype in Drosophila models of ALS. Hum Mol Genet. doi:10.1093/hmg/ddw105
Robberecht W, Philips T (2013) The changing scene of amyotrophic lateral sclerosis. Nat Rev Neurosci 14(4):248–264. doi:10.1038/nrn3430
Arai T, Hasegawa M, Akiyama H, Ikeda K, Nonaka T, Mori H, Mann D, Tsuchiya K et al (2006) TDP-43 is a component of ubiquitin-positive tau-negative inclusions in frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Biochem Biophys Res Commun 351(3):602–611. doi:10.1016/j.bbrc.2006.10.093
Neumann M, Sampathu DM, Kwong LK, Truax AC, Micsenyi MC, Chou TT, Bruce J, Schuck T et al (2006) Ubiquitinated TDP-43 in frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Science 314(5796):130–133. doi:10.1126/science.1134108
Mompean M, Buratti E, Guarnaccia C, Brito RM, Chakrabartty A, Baralle FE, Laurents DV (2014) Structural characterization of the minimal segment of TDP-43 competent for aggregation. Arch Biochem Biophys 545:53–62. doi:10.1016/j.abb.2014.01.007
Wang YT, Kuo PH, Chiang CH, Liang JR, Chen YR, Wang S, Shen JC, Yuan HS (2013) The truncated C-terminal RNA recognition motif of TDP-43 protein plays a key role in forming proteinaceous aggregates. J Biol Chem 288(13):9049–9057. doi:10.1074/jbc.M112.438564
Lukavsky PJ, Daujotyte D, Tollervey JR, Ule J, Stuani C, Buratti E, Baralle FE, Damberger FF et al (2013) Molecular basis of UG-rich RNA recognition by the human splicing factor TDP-43. Nat Struct Mol Biol 20(12):1443–1449. doi:10.1038/nsmb.2698
D’Ambrogio A, Buratti E, Stuani C, Guarnaccia C, Romano M, Ayala YM, Baralle FE (2009) Functional map** of the interaction between TDP-43 and hnRNP A2 in vivo. Nucleic Acids Res 37(12):4116–4126. doi:10.1093/nar/gkp342
Buratti E, Baralle FE (2010) The multiple roles of TDP-43 in pre-mRNA processing and gene expression regulation. RNA Biol 7(4):420–429
Yang C, Tan W, Whittle C, Qiu L, Cao L, Akbarian S, Xu Z (2010) The C-terminal TDP-43 fragments have a high aggregation propensity and harm neurons by a dominant-negative mechanism. PLoS One 5(12):e15878. doi:10.1371/journal.pone.0015878
Igaz LM, Kwong LK, Xu Y, Truax AC, Uryu K, Neumann M, Clark CM, Elman LB et al (2008) Enrichment of C-terminal fragments in TAR DNA-binding protein-43 cytoplasmic inclusions in brain but not in spinal cord of frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Am J Pathol 173(1):182–194. doi:10.2353/ajpath.2008.080003
Van Deerlin VM, Leverenz JB, Bekris LM, Bird TD, Yuan W, Elman LB, Clay D, Wood EM et al (2008) TARDBP mutations in amyotrophic lateral sclerosis with TDP-43 neuropathology: a genetic and histopathological analysis. Lancet Neurol 7(5):409–416. doi:10.1016/S1474-4422(08)70071-1
Tripathi VB, Baskaran P, Shaw CE, Guthrie S (2014) Tar DNA-binding protein-43 (TDP-43) regulates axon growth in vitro and in vivo. Neurobiol Dis 65:25–34. doi:10.1016/j.nbd.2014.01.004
Dent EW, Baas PW (2014) Microtubules in neurons as information carriers. J Neurochem 129(2):235–239. doi:10.1111/jnc.12621
Goldman JE, Yen SH (1986) Cytoskeletal protein abnormalities in neurodegenerative diseases. Ann Neurol 19(3):209–223. doi:10.1002/ana.410190302
Langford GM, Kuznetsov SA, Johnson D, Cohen DL, Weiss DG (1994) Movement of axoplasmic organelles on actin filaments assembled on acrosomal processes: evidence for a barbed-end-directed organelle motor. J Cell Sci 107(Pt 8):2291–2298
Senda T, Okabe T, Matsuda M, Fujita H (1994) Quick-freeze, deep-etch visualization of exocytosis in anterior pituitary secretory cells: localization and possible roles of actin and annexin II. Cell Tissue Res 277(1):51–60
Garner CC, Tucker RP, Matus A (1988) Selective localization of messenger RNA for cytoskeletal protein MAP2 in dendrites. Nature 336(6200):674–677. doi:10.1038/336674a0
Sundell CL, Singer RH (1991) Requirement of microfilaments in sorting of actin messenger RNA. Science 253(5025):1275–1277
Letourneau PC (2009) Actin in axons: stable scaffolds and dynamic filaments. Results Probl Cell Differ 48:65–90. doi:10.1007/400_2009_15
Hoogenraad CC, Akhmanova A (2010) Dendritic spine plasticity: new regulatory roles of dynamic microtubules. Neuroscientist 16(6):650–661. doi:10.1177/1073858410386357
Lepinoux-Chambaud C, Eyer J (2013) Review on intermediate filaments of the nervous system and their pathological alterations. Histochem Cell Biol 140(1):13–22. doi:10.1007/s00418-013-1101-1
Zhao J, Liem RK (2016) Alpha-internexin and peripherin: expression, assembly, functions, and roles in disease. Methods Enzymol 568:477–507. doi:10.1016/bs.mie.2015.09.012
Chakraborti S, Natarajan K, Curiel J, Janke C, Liu J (2016) The emerging role of the tubulin code: From the tubulin molecule to neuronal function and disease. Cytoskeleton (Hoboken) 73(10):521–550. doi:10.1002/cm.21290
Matamoros AJ, Baas PW (2016) Microtubules in health and degenerative disease of the nervous system. Brain Res Bull 126(Pt 3):217–225. doi:10.1016/j.brainresbull.2016.06.016
Dombeck DA, Kasischke KA, Vishwasrao HD, Ingelsson M, Hyman BT, Webb WW (2003) Uniform polarity microtubule assemblies imaged in native brain tissue by second-harmonic generation microscopy. Proc Natl Acad Sci U S A 100(12):7081–7086. doi:10.1073/pnas.0731953100
Leventea E, Hazime K, Zhao C, Malicki J (2016) Analysis of cilia structure and function in zebrafish. Methods Cell Biol 133:179–227. doi:10.1016/bs.mcb.2016.04.016
Hirokawa N, Niwa S, Tanaka Y (2010) Molecular motors in neurons: transport mechanisms and roles in brain function, development, and disease. Neuron 68(4):610–638. doi:10.1016/j.neuron.2010.09.039
Su X, Ohi R, Pellman D (2012) Move in for the kill: motile microtubule regulators. Trends Cell Biol 22(11):567–575. doi:10.1016/j.tcb.2012.08.003
Chen Y, Hancock WO (2015) Kinesin-5 is a microtubule polymerase. Nat Commun 6:8160. doi:10.1038/ncomms9160
Komis G, Illes P, Beck M, Samaj J (2011) Microtubules and mitogen-activated protein kinase signalling. Curr Opin Plant Biol 14(6):650–657. doi:10.1016/j.pbi.2011.07.008
Sun T, Rodriguez M, Kim L (2009) Glycogen synthase kinase 3 in the world of cell migration. Develop Growth Differ 51(9):735–742. doi:10.1111/j.1440-169X.2009.01141.x
Vertessy BG, Orosz F, Kovacs J, Ovadi J (1997) Alternative binding of two sequential glycolytic enzymes to microtubules. Molecular studies in the phosphofructokinase/aldolase/microtubule system. J Biol Chem 272(41):25542–25546
Halpain S, Dehmelt L (2006) The MAP1 family of microtubule-associated proteins. Genome Biol 7(6):224
Riederer BM, Draberova E, Viklicky V, Draber P (1995) Changes of MAP2 phosphorylation during brain development. J Histochem Cytochem 43(12):1269–1284
Roll-Mecak A, McNally FJ (2010) Microtubule-severing enzymes. Curr Opin Cell Biol 22(1):96–103. doi:10.1016/j.ceb.2009.11.001
Ratti A, Buratti E (2016) Physiological functions and pathobiology of TDP-43 and FUS/TLS proteins. J Neurochem 138(Suppl 1):95–111. doi:10.1111/jnc.13625
Colombrita C, Zennaro E, Fallini C, Weber M, Sommacal A, Buratti E, Silani V, Ratti A (2009) TDP-43 is recruited to stress granules in conditions of oxidative insult. J Neurochem 111(4):1051–1061. doi:10.1111/j.1471-4159.2009.06383.x
Nover L, Scharf KD, Neumann D (1989) Cytoplasmic heat shock granules are formed from precursor particles and are associated with a specific set of mRNAs. Mol Cell Biol 9(3):1298–1308
Mahboubi H, Stochaj U (2017) Cytoplasmic stress granules: dynamic modulators of cell signaling and disease. Biochim Biophys Acta 1863(4):884–895. doi:10.1016/j.bbadis.2016.12.022
Takahashi M, Higuchi M, Matsuki H, Yoshita M, Ohsawa T, Oie M, Fujii M (2013) Stress granules inhibit apoptosis by reducing reactive oxygen species production. Mol Cell Biol 33(4):815–829. doi:10.1128/MCB.00763-12
Orru S, Coni P, Floris A, Littera R, Carcassi C, Sogos V, Brancia C (2016) Reduced stress granule formation and cell death in fibroblasts with the A382T mutation of TARDBP gene: evidence for loss of TDP-43 nuclear function. Hum Mol Genet. doi:10.1093/hmg/ddw276
Cohen TJ, Hwang AW, Restrepo CR, Yuan CX, Trojanowski JQ, Lee VM (2015) An acetylation switch controls TDP-43 function and aggregation propensity. Nat Commun 6:5845. doi:10.1038/ncomms6845
Alami NH, Smith RB, Carrasco MA, Williams LA, Winborn CS, Han SS, Kiskinis E, Winborn B et al (2014) Axonal transport of TDP-43 mRNA granules is impaired by ALS-causing mutations. Neuron 81(3):536–543. doi:10.1016/j.neuron.2013.12.018
Ayala YM, Pantano S, D’Ambrogio A, Buratti E, Brindisi A, Marchetti C, Romano M, Baralle FE (2005) Human, Drosophila, and C. elegans TDP43: nucleic acid binding properties and splicing regulatory function. J Mol Biol 348(3):575–588. doi:10.1016/j.jmb.2005.02.038
Romano M, Buratti E, Romano G, Klima R, Del Bel BL, Stuani C, Baralle F, Feiguin F (2014) Evolutionarily conserved heterogeneous nuclear ribonucleoprotein (hnRNP) A/B proteins functionally interact with human and Drosophila TAR DNA-binding protein 43 (TDP-43). J Biol Chem 289(10):7121–7130. doi:10.1074/jbc.M114.548859
Pesiridis GS, Tripathy K, Tanik S, Trojanowski JQ, Lee VM (2011) A “two-hit” hypothesis for inclusion formation by carboxyl-terminal fragments of TDP-43 protein linked to RNA depletion and impaired microtubule-dependent transport. J Biol Chem 286(21):18845–18855. doi:10.1074/jbc.M111.231118
Vanden Broeck L, Naval-Sanchez M, Adachi Y, Diaper D, Dourlen P, Chapuis J, Kleinberger G, Gistelinck M et al (2013) TDP-43 loss-of-function causes neuronal loss due to defective steroid receptor-mediated gene program switching in Drosophila. Cell Rep 3(1):160–172. doi:10.1016/j.celrep.2012.12.014
Pereira A, Doshen J, Tanaka E, Goldstein LS (1992) Genetic analysis of a Drosophila microtubule-associated protein. J Cell Biol 116(2):377–383
Coyne AN, Siddegowda BB, Estes PS, Johannesmeyer J, Kovalik T, Daniel SG, Pearson A, Bowser R et al (2014) Futsch/MAP1B mRNA is a translational target of TDP-43 and is neuroprotective in a Drosophila model of amyotrophic lateral sclerosis. J Neurosci 34(48):15962–15974. doi:10.1523/JNEUROSCI.2526-14.2014
Godena VK, Romano G, Romano M, Appocher C, Klima R, Buratti E, Baralle FE, Feiguin F (2011) TDP-43 regulates Drosophila neuromuscular junctions growth by modulating Futsch/MAP1B levels and synaptic microtubules organization. PLoS One 6(3):e17808. doi:10.1371/journal.pone.0017808
Li Y, Ray P, Rao EJ, Shi C, Guo W, Chen X, Woodruff EA 3rd, Fushimi K et al (2010) A Drosophila model for TDP-43 proteinopathy. Proc Natl Acad Sci U S A 107(7):3169–3174. doi:10.1073/pnas.0913602107
Hanson KA, Kim SH, Wassarman DA, Tibbetts RS (2010) Ubiquitin modifies TDP-43 toxicity in a Drosophila model of amyotrophic lateral sclerosis (ALS). J Biol Chem 285(15):11068–11072. doi:10.1074/jbc.C109.078527
Hummel T, Krukkert K, Roos J, Davis G, Klambt C (2000) Drosophila Futsch/22C10 is a MAP1B-like protein required for dendritic and axonal development. Neuron 26(2):357–370
Romano M, Feiguin F, Buratti E (2016) TBPH/TDP-43 modulates translation of Drosophila futsch mRNA through an UG-rich sequence within its 5'UTR. Brain Res 1647:50–56. doi:10.1016/j.brainres.2016.02.022
Majumder P, Chu JF, Chatterjee B, Swamy KB, Shen CJ (2016) Co-regulation of mRNA translation by TDP-43 and Fragile X Syndrome protein FMRP. Acta Neuropathol 132 (5):721–738. doi:10.1007/s00401-016-1603-8
Wu CH, Fallini C, Ticozzi N, Keagle PJ, Sapp PC, Piotrowska K, Lowe P, Koppers M et al (2012) Mutations in the profilin 1 gene cause familial amyotrophic lateral sclerosis. Nature 488(7412):499–503. doi:10.1038/nature11280
Tanaka Y, Nonaka T, Suzuki G, Kametani F, Hasegawa M (2016) Gain-of-function profilin 1 mutations linked to familial amyotrophic lateral sclerosis cause seed-dependent intracellular TDP-43 aggregation. Hum Mol Genet 25(7):1420–1433. doi:10.1093/hmg/ddw024
Yuan A, Rao MV, Veeranna N, R.A. (2012) Neurofilaments at a glance. J Cell Sci 125:3257–3263
Bergeron C, Beric-Maskarel K, Muntasser S, Weyer L, Somerville MJ, Percy ME (1994) Neurofilament light and polyadenylated mRNA levels are decreased in amyotrophic lateral sclerosis motor neurons. J Neuropathol Exp Neurol 53(3):221–230
Corbo M, Hays AP (1992) Peripherin and neurofilament protein coexist in spinal spheroids of motor neuron disease. J Neuropathol Exp Neurol 51(5):531–537
Strong MJ, Volkening K, Hammond R, Yang W, Strong W, Leystra-Lantz C, Shoesmith C (2007) TDP43 is a human low molecular weight neurofilament (hNFL) mRNA-binding protein. Mol Cell Neurosci 35(2):320–327. doi:10.1016/j.mcn.2007.03.007
Volkening K, Leystra-Lantz C, Yang W, Jaffee H, Strong MJ (2009) Tar DNA binding protein of 43 kDa (TDP-43), 14-3-3 proteins and copper/zinc superoxide dismutase (SOD1) interact to modulate NFL mRNA stability. Implications for altered RNA processing in amyotrophic lateral sclerosis (ALS). Brain Res 1305:168–182. doi:10.1016/j.brainres.2009.09.105
Polymenidou M, Lagier-Tourenne C, Hutt KR, Huelga SC, Moran J, Liang TY, Ling SC, Sun E et al (2011) Long pre-mRNA depletion and RNA missplicing contribute to neuronal vulnerability from loss of TDP-43. Nat Neurosci 14(4):459–468. doi:10.1038/nn.2779
Liu Y, Atkinson RA, Fernandez-Martos CM, Kirkcaldie MT, Cui H, Vickers JC, King AE (2015) Changes in TDP-43 expression in development, aging, and in the neurofilament light protein knockout mouse. Neurobiol Aging 36(2):1151–1159. doi:10.1016/j.neurobiolaging.2014.10.001
Swarup V, Phaneuf D, Bareil C, Robertson J, Rouleau GA, Kriz J, Julien JP (2011) Pathological hallmarks of amyotrophic lateral sclerosis/frontotemporal lobar degeneration in transgenic mice produced with TDP-43 genomic fragments. Brain 134(Pt 9):2610–2626. doi:10.1093/brain/awr159
Escurat M, Djabali K, Gumpel M, Gros F, Portier MM (1990) Differential expression of two neuronal intermediate-filament proteins, peripherin and the low-molecular-mass neurofilament protein (NF-L), during the development of the rat. J Neurosci 10(3):764–784
Corrado L, Carlomagno Y, Falasco L, Mellone S, Godi M, Cova E, Cereda C, Testa L et al (2011) A novel peripherin gene (PRPH) mutation identified in one sporadic amyotrophic lateral sclerosis patient. Neurobiol Aging 32(3):552. doi:10.1016/j.neurobiolaging.2010.02.011 e551-556
Gros-Louis F, Lariviere R, Gowing G, Laurent S, Camu W, Bouchard JP, Meininger V, Rouleau GA et al (2004) A frameshift deletion in peripherin gene associated with amyotrophic lateral sclerosis. J Biol Chem 279(44):45951–45956. doi:10.1074/jbc.M408139200
Leung CL, He CZ, Kaufmann P, Chin SS, Naini A, Liem RK, Mitsumoto H, Hays AP (2004) A pathogenic peripherin gene mutation in a patient with amyotrophic lateral sclerosis. Brain Pathol 14(3):290–296
Comley L, Allodi I, Nichterwitz S, Nizzardo M, Simone C, Corti S, Hedlund E (2015) Motor neurons with differential vulnerability to degeneration show distinct protein signatures in health and ALS. Neuroscience 291:216–229. doi:10.1016/j.neuroscience.2015.02.013
He CZ, Hays AP (2004) Expression of peripherin in ubiquinated inclusions of amyotrophic lateral sclerosis. J Neurol Sci 217(1):47–54
Mizuno Y, Fujita Y, Takatama M, Okamoto K (2011) Peripherin partially localizes in Bunina bodies in amyotrophic lateral sclerosis. J Neurol Sci 302(1–2):14–18. doi:10.1016/j.jns.2010.12.023
**ao S, Tjostheim S, Sanelli T, McLean JR, Horne P, Fan Y, Ravits J, Strong MJ et al (2008) An aggregate-inducing peripherin isoform generated through intron retention is upregulated in amyotrophic lateral sclerosis and associated with disease pathology. J Neurosci 28(8):1833–1840. doi:10.1523/JNEUROSCI.3222-07.2008
Muresan V, Ladescu Muresan Z (2016) Shared molecular mechanisms in Alzheimer’s disease and amyotrophic lateral sclerosis: neurofilament-dependent transport of sAPP, FUS, TDP-43 and SOD1, with endoplasmic reticulum-like tubules. Neurodegener Dis 16(1–2):55–61. doi:10.1159/000439256
Wolozin B (2012) Regulated protein aggregation: stress granules and neurodegeneration. Mol Neurodegener 7:56. doi:10.1186/1750-1326-7-56
McDonald KK, Aulas A, Destroismaisons L, Pickles S, Beleac E, Camu W, Rouleau GA, Vande Velde C (2011) TAR DNA-binding protein 43 (TDP-43) regulates stress granule dynamics via differential regulation of G3BP and TIA-1. Hum Mol Genet 20(7):1400–1410. doi:10.1093/hmg/ddr021
Vanderweyde T, Apicco DJ, Youmans-Kidder K, Ash PE, Cook C, Lummertz da Rocha E, Jansen-West K, Frame AA et al (2016) Interaction of tau with the RNA-binding protein TIA1 regulates tau pathophysiology and toxicity. Cell Rep 15(7):1455–1466. doi:10.1016/j.celrep.2016.04.045
Zetterberg H, Skillback T, Mattsson N, Trojanowski JQ, Portelius E, Shaw LM, Weiner MW, Blennow K et al (2016) Association of cerebrospinal fluid neurofilament light concentration with Alzheimer disease progression. JAMA Neurol 73(1):60–67. doi:10.1001/jamaneurol.2015.3037
Pijnenburg YA, Verwey NA, van der Flier WM, Scheltens P, Teunissen CE (2015) Discriminative and prognostic potential of cerebrospinal fluid phosphoTau/tau ratio and neurofilaments for frontotemporal dementia subtypes. Alzheimers Dement (Amst) 1(4):505–512. doi:10.1016/j.dadm.2015.11.001
Steinacker P, Feneberg E, Weishaupt J, Brettschneider J, Tumani H, Andersen PM, von Arnim CA, Bohm S et al (2016) Neurofilaments in the diagnosis of motoneuron diseases: a prospective study on 455 patients. J Neurol Neurosurg Psychiatry 87(1):12–20. doi:10.1136/jnnp-2015-311387
Brettschneider J, Petzold A, Sussmuth SD, Ludolph AC, Tumani H (2006) Axonal damage markers in cerebrospinal fluid are increased in ALS. Neurology 66(6):852–856. doi:10.1212/01.wnl.0000203120.85850.54
Weydt P, Oeckl P, Huss A, Muller K, Volk AE, Kuhle J, Knehr A, Andersen PM et al (2016) Neurofilament levels as biomarkers in asymptomatic and symptomatic familial amyotrophic lateral sclerosis. Ann Neurol 79(1):152–158. doi:10.1002/ana.24552
Reijn TS, Abdo WF, Schelhaas HJ, Verbeek MM (2009) CSF neurofilament protein analysis in the differential diagnosis of ALS. J Neurol 256(4):615–619. doi:10.1007/s00415-009-0131-z
Lu CH, Macdonald-Wallis C, Gray E, Pearce N, Petzold A, Norgren N, Giovannoni G, Fratta P et al (2015) Neurofilament light chain: a prognostic biomarker in amyotrophic lateral sclerosis. Neurology 84(22):2247–2257. doi:10.1212/WNL.0000000000001642
Oeckl P, Jardel C, Salachas F, Lamari F, Andersen PM, Bowser R, de Carvalho M, Costa J et al (2016) Multicenter validation of CSF neurofilaments as diagnostic biomarkers for ALS. Amyotroph Lateral Scler Frontotemporal Degener 17(5–6):404–413
Li S, Ren Y, Zhu W, Yang F, Zhang X, Huang X (2016) Phosphorylated neurofilament heavy chain levels in paired plasma and CSF of amyotrophic lateral sclerosis. J Neurol Sci 367:269–274. doi:10.1016/j.jns.2016.05.062
Tortelli R, Copetti M, Ruggieri M, Cortese R, Capozzo R, Leo A, D’Errico E, Mastrapasqua M et al (2015) Cerebrospinal fluid neurofilament light chain levels: marker of progression to generalized amyotrophic lateral sclerosis. Eur J Neurol 22(1):215–218. doi:10.1111/ene.12421
Tortelli R, Copetti M, Panza F, Cortese R, Capozzo R, D’Errico E, Fontana A, Simone IL et al (2016) Time to generalisation as a predictor of prognosis in amyotrophic lateral sclerosis. J Neurol Neurosurg Psychiatry 87(6):678–679. doi:10.1136/jnnp-2014-308478
Ganesalingam J, An J, Shaw CE, Shaw G, Lacomis D, Bowser R (2011) Combination of neurofilament heavy chain and complement C3 as CSF biomarkers for ALS. J Neurochem 117(3):528–537. doi:10.1111/j.1471-4159.2011.07224.x
Ganesalingam J, An J, Bowser R, Andersen PM, Shaw CE (2013) pNfH is a promising biomarker for ALS. Amyotroph Lateral Scler Frontotemporal Degener 14(2):146–149. doi:10.3109/21678421.2012.729596
Menke RA, Gray E, Lu CH, Kuhle J, Talbot K, Malaspina A, Turner MR (2015) CSF neurofilament light chain reflects corticospinal tract degeneration in ALS. Ann Clin Transl Neurol 2(7):748–755. doi:10.1002/acn3.212
Puentes F, Top** J, Kuhle J, van der Star BJ, Douiri A, Giovannoni G, Baker D, Amor S et al (2014) Immune reactivity to neurofilament proteins in the clinical staging of amyotrophic lateral sclerosis. J Neurol Neurosurg Psychiatry 85(3):274–278. doi:10.1136/jnnp-2013-305494
Budini M, Romano V, Quadri Z, Buratti E, Baralle FE (2015) TDP-43 loss of cellular function through aggregation requires additional structural determinants beyond its C-terminal Q/N prion-like domain. Hum Mol Genet 24(1):9–20. doi:10.1093/hmg/ddu415
Fang YS, Tsai KJ, Chang YJ, Kao P, Woods R, Kuo PH, Wu CC, Liao JY et al (2014) Full-length TDP-43 forms toxic amyloid oligomers that are present in frontotemporal lobar dementia-TDP patients. Nat Commun 5:4824. doi:10.1038/ncomms5824
Smith BN, Ticozzi N, Fallini C, Gkazi AS, Topp S, Kenna KP, Scotter EL, Kost J et al (2014) Exome-wide rare variant analysis identifies TUBA4A mutations associated with familial ALS. Neuron 84(2):324–331. doi:10.1016/j.neuron.2014.09.027
Acknowledgements
We acknowledge the financial support of an ALS Young Investigator Research Scholarship by the Deutsche Gesellschaft für Muskelkranke (DGM) (to M.O.) and EU funds (EFRE; SAB, 100111005; to M.H.).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Oberstadt, M., Claßen, J., Arendt, T. et al. TDP-43 and Cytoskeletal Proteins in ALS. Mol Neurobiol 55, 3143–3151 (2018). https://doi.org/10.1007/s12035-017-0543-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-017-0543-1